Opsin evolution: transducins
See also: RBP3 (IRBP) | RPE65 | Usher: USH2A | Usher: CDH23 | LOXHD1 | Update Blog
Transducin-opsin co-evolution
Opsins have expanded (and contracted) considerably in deuterostomes. Those changes were accompanied by a mostly independent expansion in heterotrimeric G proteins that comprise an early step in converting the initial photoreception event into a cell signal. That signal can initiate with increased cyclic nucleotide or phosphoinositol and have many downstream regulatory sequalae.
The specific association between opsin orthology class and alpha subunit class of heterotrimeric G protein are gradually being worked out across cnidaria and bilatera, but with a steady stream of surprises. While the specific domains within opsins responsible for the Ga protein-protein interaction are well-understood from recent 3D structures and accompanying experiments, predicting the precise Ga specificity of a given opsin (or other GPCR) is very difficult at this time (the only online tool) just classifies to four gene families).
In theory, any opsin (or GPCR) in any given species could be threaded to a known 3D structure and computationally docked to each of the similarly modeled Ga subunits that occur in that species, the most favorable fit then being the prediction. That might not be feasible since the responsible opsin region is a cytoplasmic loop that is not really predicted from 7-transmembrane considerations.
The alternative, compiling extensive comparative genomics and seeking primary sequence correlations, requires extensive seeding from experimentally known binding pairs. In other words, this is not ab initio prediction so much as homology transfer. That could be quite useful however if it carried across opsin orthology classes to those lacking any experimental data (perhaps because standard model organisms lacked counterparts).
It has been argued that some GPCR classifying within opsins (such as peropsin, neuropsin, newropsin, and rgropsin) are non-signaling photoisomerases with roles in carotenoid or retinoid metabolism and recycling. That largely conflicts however with their observed comparative genomics -- if no Ga is bound, what selective pressure would conserve those binding motifs over billions of years of evolutionary branch length?
RGR is an especially interesting case because it appears to have signaling capacity (though unknown Ga) from tunicates to early placentals, yet lost crucial signaling residues and Ga binding domains in all boreoeutheres (meaning mouse cannot serve as model species). Here chicken or frog would provide an experimental system if the Ga cannot be reliably predicted.
Cnidarian photoreception, a very active area of research critical to understanding the origins of lensing vision, is also in a state of considerable confusion. Many of the purported opsins do not classify properly, species such as Nematostella and Hydra have far too many 'opsins' for their meagre photoreceptive structures, and the only reported signaling partner (in box jellyfish Carybdea rastonii) is of unexpected Gs class. Complete genomes of box jellyfish will prove necessary to establish both the opsin and Gz repertoires.
A perplexing issue within Ga gene family evolution arises from probable independent (parallel) expansions in different clades at different times. Compounding this complexity are separate expansions in the other two members of the heterotrimeric G protein. It is not straightforward to compare opsin-heterotrimer interaction from cnidaria, protostomes, and vertebrates -- the terms paralog and ortholog are woefully insufficient.
Earth history affects on gene expansion
The origin of vertebrate genes involves a complex sequence of gain and loss processes involving many thousands of events lineage-specific to greater or lesser extents. No single simple-minded scenario (such as 1R or 2R) or principle (eg parsimony, increasing complexity) could possibly account for the observed multiplicities of gene families in say human and their current ordering on chromosomes. For example human lineage has experienced a dramatic drop in opsin genes yet slow but uneven expansion in the three G protein subunits.
Evolution is a topic in one-off history, not accessible to resampling statistics. That history, once guided by unidirectional progress towards a manifest destiny of (human) perfection, is better understood as happenstance and adaptation to prevailing selective conditions which cannot anticipate future conditions and indeed often become maladaptive or of no utility.
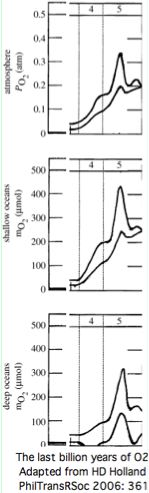
It's not even clear whether GPCR signaling, often taken as proxy for increasing sensory and multicellular communication complexity, has had any real trend in gene numbers since the Cambrian oxygenation of the oceans (which benefited multi-cellularity by enabling oxidative phosphorylation that permitted high-consumption tissues and systems).
Indeed atmospheric and surface water oxygenation peaked in the Carboniferous at nearly double today's level (supporting gigantic insects). Of course, early-diverging non-vertebrate lineages were not frozen in primitive ancestral condition but also could benefit from higher oxygen levels.
Genome sequencing projects surprisingly show a decreasing trend in gene count in later diverging deuterostomes. For example, sea urchin genome has 23,300 coding genes whereas humans have but 20,176 in the 11 Sept 2008 tally of consensus CDS and even this number seems inflated relative to the 17,052 distinct locus count by assignment of multiple CCDS IDs numbers to single genes.
Note genome sequencing here is very incomplete and assemblies are defective, adding many errors in coding gene annotation to those related to the intrinsic difficulty of gene-finding. Gene counts refer to contemporary organisms only roughly estimate actual ancestral counts at distant nodes.
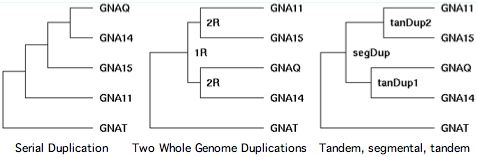
The processes that create new genes fall into four very distinct categories. The first involves single gene retro duplications that do not include the parental gene upstream promoters or untranscribed regulatory regions. The second, single gene tandem duplication (either inline or inverted), generally provides for initial transcription of the new copy from parent gene control regions. Small to large segmental translocations to a new chromosome bring with them -- at least for genes internal to the block -- the original transcription control apparatus though the chromatin milieu may be quite different.
Fourth, polyploidization (whole genome duplication) brings along a complete second system of interacting genes but along with it undesirable issues in gene copy number. This can already be seen in Down Syndrome, which is sometimes only partial aneuploidy involving the second smallest chromosome (271 or 1.3% of total genes) and involves multiple deleterious genes. The mammalian sex chromosome system, which has evolved relatively recently, also had to evolve compensatory mechanisms for gene copy number, notably random X inactivation and enhanced autosomal retrogene copies.
It would take a great many generations to lose or exapt all the deleterious genes genomewide expected from tetraploidization, raising the question of how the polyploidization event could ever become fixed in a population. Yet this process is common in grasses and is generally accepted in teleost fish (though ironically gene counts have not notably increased). Here it must be noted that the reported South American mouse tetraploidization has been disproven by chromosome staining and all five fish genome projects have been abandoned far from completion despite great numbers of gaps and contig misassembly and multiple use.
It is very difficult under these circumstances to distinguish whole genome duplication from extensive aneuploidy, robertsonian translocations, and numerous large segmental duplications. However the human genome does contain a significant number of unmistakable small and large paralogons with good retention of paralogs, illustrating that not all block duplications result in Down Syndrome copy number issues.
Paralogon is a neutral term preferred here that asserts regional homology but does not take a position on mechanistic origins (which can become quite muddied over the passage of time by subsequent overlaid inversions, partial translocations, gene insertions, and gene losses).
Retrogenes that arise from reverse transcription of mRNAs lose the introns (if any) of the parental gene, though they can subsequently acquire unrelated new introns. Single-exon genes, at 1832 genes the most frequent category at 9%, can be difficult to distinguish from their retrogenes and pseudogenes. Retrogenes, not at all uncommon, are difficult to distinguish from processed pseudogenes (which sometimes continue to be transcribed). The number of processed pseudogenes with significant alignment to a parental gene is very large, approximately equal to the number of genes.
Subunits of heterotrimeric G protein do not often give rise to either pseudogenes or retrogenes. The one notable exception is alpha subunit GNAZ. This gene appears to be a functioning processed retrogene with a single intron in novel location and phasing (meaning it could not have arisen from incomplete processing). The two events both date to lamprey stem. The gene is now on human chr22; the parent gene lies in the GNAI group with implications for its signaling mechanism.
The gene is exceedingly conserved, over 95% identity human to lamprey despite a billion years of branch length. Such genes cause confusion on Oxford grids (which ignore exon structure) because there is a fair chance of a coincidental non-orthologous high-scoring match in a given chromosomal comparison with with 16 paralogs. Obviously this gene did not arise by 1R or 2R and indeed itself remains single copy despite dating to the supposed whole genome duplication era pre-lamprey.
Tandem duplication, either inline or inverted, is a very common process. The descendent genes are often separated by translocation (which cuts down on gene conversion homogenization and favors retention). Translocation also occurs irrespective of tandem duplication, often in large blocks. Gibbon genome illustrates extremes of chromosomal joining and separation that have similar outcomes to translocation.
These circumstances make all-vs-all blast synteny (Oxford grids, dot plots) too coarse for determining gene histories. It would be better to first mask all single-exon genes and then to validate exon numbers between putative matches, as well as require more than matching of just some common domain. Further, if two regions are closely related, then their corresponding proteins should often be best reciprocal blast (not way down on the list).
GeneSorter at UCSC allows rapid curational distinction between large paralogons, regional segmental duplications, small block translocations and isolated retrogenes, even when core events have been overwritten by subsequent rearrangements, losses and gains. This is best illustrated by a concrete example.
Beginning with the +GNAQ +GNA14 tandem inline duplicate on human chromosome 9, we wish to establish its relation to the paralogous tandem duplicate +GNA11 +GNA15 genes on chromosome 19. First note that all four genes have 7 coding exons with homologous positions and identical phases and are unambiguously alignable over their entire lengths at high percent identity.
Curious evolutionary origins of alpha subunit multiplicity
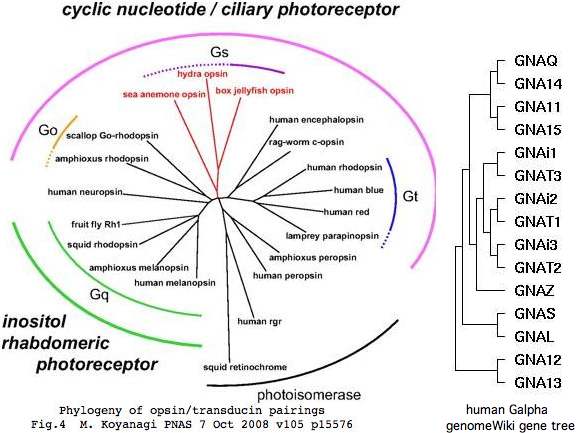
The human genome contains 16 paralogous alpha, 5 beta, and 12 gamma subunits of heterotrimeric guanine nucleotide binding protein (G protein) though not all combinations occur and few are specifically relevant to imaging opsins (namely GNAT1/GNB1/GNGT1 for rods and GNAT2/GNB3/GNGT2 for cones). Because opsins comprise only ~1% of total GPCR served and non-imaging early diverging species such sea urchin already have vast repertoires of GPCR -- 979 of them -- and complex multiplicities of heterotrimeric G protein subunits (below), Darwin's question on independent origins of vision is quite muddled by pre-existence of various subsystems that were later exapted.
The primary issue under discussion here is expansion of ciliary and imaging opsin genes during an era when signaling partner components were also increasing by different and not fully coordinated genetic mechanisms. A G protein alpha subunit can serve other GPCR in addition to opsins and a given opsin does not necessarily signal via a dedicated alpha subunit. Beta and gamma subunits of heterotrimeric G protein have still different temporal expansion histories, again with implications for opsins, but that complexity is considered only tangentially here as only the alpha subunit binds directly to opsins.
Consequently we do not expect nor find a 1:1 mapping over time as these gene families expanded by separate sequences of events, even though these proteins were manifestly co-evolving. Still, we would like to understand ancestral and contemporary opsin signaling because photoreception in isolation accomplishes nothing. That signaling can be described in part by its downstream small molecule and membrane channel components.
Cone and rod opsins have dedicated alpha subunits called Gt transducins that, like so much in vertebrate vision, are genes already established prior to lamprey divergence in its long and short photoreceptors. The situation is the same for the two gamma inhibitory subunits of cGMP phosphodiesterase PDE6 family ultimately activated by transducin but the alpha catalytic subunit appears to have not yet duplicated in lamprey.
For brevity, 'dating an event' is shorthand here for thoroughly examining paralog number and syntenic relations in relevant genome browsers (and ancillary data at GenBank) and taking simplest scenario compatible with the data, typically a short sequence of common genetic events such as tandem duplication and divergence. Dating is not quantitatively chronological but rather relative to consecutive divergence nodes of the deuterostome phylogenetic tree. Note hagfish and early chordate topology remain slightly equivocal. Lamprey contigs assemblies are often too short to hold complete genes much less reveal syntenic relations.
As future assemblies of certain incomplete but critical genomes (such as lamprey and shark) improve and as established knowledge of ancestral genetic events grows, these working hypotheses can be sharpened in their details or confidence, or even replaced. However no improvement can be expected today from pseudo-objective theories of maximal parsimony or likelihood that at best bury dubious curational assumptions in software code and at worst underperform common sense.
Curiously, the 16 alpha subunit paralogs in human include 5 deeply conserved tandem pairs on five separate chromosomes, for example cone transducin GNAT2 and GNAI3. That suggests some combination of multiple local tandem gene duplications coupled with segmental, whole chromosomal, or even whole genome duplication of pairs, as considered early on for 9 phototransduction gene classes. Note to minimize coincidental synteny, it is imperative to establish that gene relationships are ancestral by comparative genomics.
Tandem gene pairs conserved in Ga evolution (human coordinates) +GNAT1 chr3:50204047 +GNAi2 chr3:50248651* tail to head +GNAi3 chr1:109892709 -GNAT2 chr1:109947412 tail to tail +GNAi1 chr7:79602076 +GNAT3 chr7:79925923 tail to head +GNAQ chr9:79525011 +GNA14 chr9:79228368 tail to head +GNA11 chr19:3045400 +GNA15 chr19:3087191 tail to head GNAO1 chr16:54782752 GNAZ chr22:21742669 non-tandem GNA13 chr17:60437295 GNA12 chr7:2734267 non-tandem GNAL chr18:11679265 GNAS chr20:56861431 non-tandem *intervening gene +SNAT3 (SLC38A3) by local inversion
With gene order otherwise so scrambled by inversion and translocation, perhaps some functional constraint has kept these tandem pairs together (as with the LWS opsin locus control region). Yet upstream GNAT2 regulation does not seem physically or functionally appropriate to GNAI3. The five tandem pairs do not exhibit consistent strand orientations.
It is very implausible that these genes arose elsewhere and were brought together by chromosomal rearrangement. Consequently one member of the pair must be parental to the other. This relationship must trump gene trees that emerge from alignment tools (which can be thrown off by a rapidly evolving gene). If one member of a tandem pair retains ancestral function, the other may be rapidly pushed away in sequence space to develop a selective niche, meaning an excessive rate of divergence and consequent misclassification.
Four other alpha subunits (GNAL, GNAS, GNA12, and GNA13) are so distantly diverged that they have utility here only as basal outgroups. They appear to already have been established in placazoan and been immune to subsequent expansion and contraction.
The alpha subunit GNAZ is a functioning processed retrogene with one intron in novel location and phasing (meaning it could not have arisen from incomplete processing). The two events both date to lamprey stem. The gene is now on human chr22; the parent gene lies in the GNAI group with implications for its signaling mechanism. The gene is exceedingly conserved, over 95% identity human to lamprey despite a billion years of branch length. This could cause confusion on Oxford grids (which ignores exon structure) because with 16 paralogs there is a fair chance of a coincidental non-orthologous high-scoring match in a given chromosomal comparison, yet this gene obviously did not arise by 1R or 2R and indeed itself remains single copy despite dating to the supposed whole genome duplication era pre-lamprey.
Structure/function of signaling: the Gα interaction with opsins
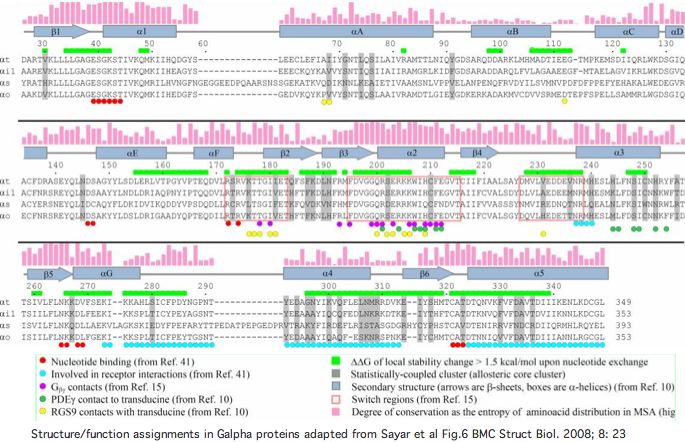
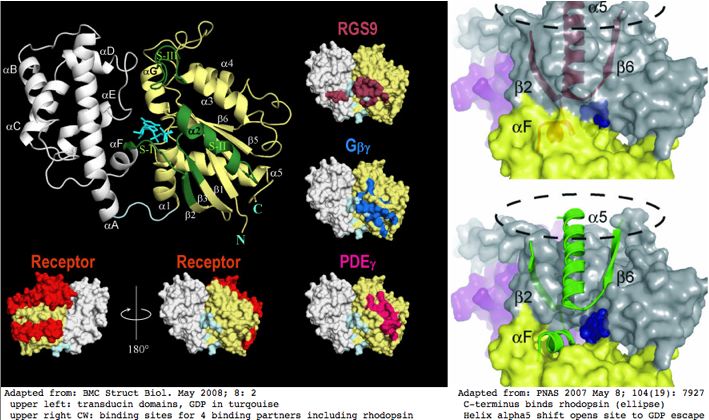
A great deal is known about structure/function relationships in Gα subunits. This helps in understanding why particular conserved residues are observed in linear sequence alignments. That information is summarized in the graphics below.
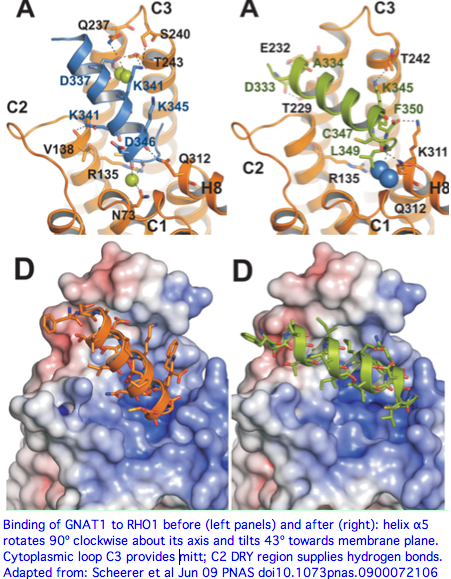
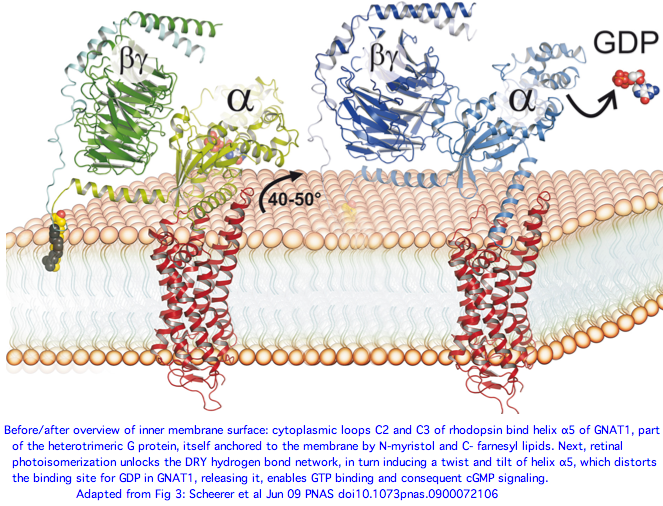
A June 2009 article in PNAS provides an excellent account of the molecular details of GPCR signaling. Here the terminal alpha helix 5 of transducin (GNAT1 for vertebrate rods) provides the key connection between photo-induced conformational changes in rhodopsin (RHO1 below) and the initiation of cGMP signaling. The cytoplasmic loops C2 and C3 of rhodopsin provide the binding pocket for the GNAT1 helix.
When rhodopsin adsorbs a photon, that triggers the isomerization of cis-retinal to all-trans and its release from the Schiff base lysine. That induces a conformational shift in the DRY region of loop C2, causing the already-bound helix of GNAT1 to rotate 90º about its helical axis and further tilt 43º in the mitt binding pocket of loop C3. This levers open the GDP binding pocket of GNAT1 causing GDP release. That site is then available for GTP and the beginning of the cGMP increase that leads to membrane hyperpolarization, which amounts to the rod neuron sending a pulse towards the brain indicating light has been perceived by the rod cell in question.
From the comparative genomics standpoint, not all observed sequence conservation is satisfactorily explained. First, many of the interactions at the binding pocket involve hydrogen bonds to the main peptide chain, not to the individual side chains. Yet the main chain is completely generic except at prolines so the proposed signaling mechanism offers no explanation why the side chains themselves -- which could provide specificity but apparently don't -- are conserved. Hydrophobic interactions are also important to the binding and its shift but these too cannot explain conserved polar or charged residues. Thus much of the conservation must be attributed to selection for other functionalities.
Since humans have 16 paralogs of GNAT1 transducin and many hundreds of non-opsin GPCR to interact with, it's of interest to compare the evolution of GNAT1 helix 5 across same-species paralogs (little conservation) versus its conservation within same-species orthologs (extreme conservation). Observe that GNAT2 helix 5 is identical to GNAT1, implying any specificity must reside in the C2 and C3 loops (which however are very similar in cone and rod opsins) or be effectuated by cell-type specific expression of the two transducins (ie GNAT1 could work in cone cells but it is not expressed there; vice versa for GNAT2 in rods).
Similarly, it will prove very difficult to distinguish melanopsin signaling via GNAQ from that of GNA11 because they have an identical fifth helix. There is shockingly little variation of these helices within vertebrates (human to lamprey). Indeed fifth helix sequence conservation was largely fixed in stone back not later than cnidarian divergence. It follows that tools attempting to predict signaling partners from GPCR sequence alone could be improved by including the cognate fifth helix in their HMM training set, as the protein pair is co-evolving.
Note the GNAT1 family has a complicated history of duplication that may be in part vertebrate-specific. It becomes important to work out separate histories in protostomes and cnidaria if opsin signaling there is to be understood. A point of confusion here is terms such as "Gq" or "Gs" signaling really refer to the downstream signaling chemistry without really addressing lineage-specific gene duplication and subfunctionalization issues of GNAQ or GNAS.
For example, arthropods have two major classes of melanopsins (ultraviolet and long wavelength sensitive) in addition to the RH7 class which has lost its C3 mitt. Has GNAQ experienced a gene duplication and subfunctionalization similar to GNAT1 and GNAT2 in ciliary opsins? Here the counterparts might be ocelli vs compound eyes.
On average, a given vertebrate Galpha must help around 60 different GPCR genes signal. GNAT1 is an exception, apparently be confined to rod rhodopsin and GNAT2 is also apparently restricted to a few cone opsins, while the breadth of GNAQ signaling beyond melanopsins is not so clear. Olfaction is at the other extreme with hundreds of genes signaling specifically through GNAT3. This leaves 13 Galpha coupling to the remaining 600 GPCR.
Consequently, the evolution of the fifth helix of Galpha genes with multiple partners is greatly constrained -- the slightest change can have multiple unintended consequences in GPCR signaling in diverse systems. This doesn't mean they evolve slower than fifth helices of other Galpha genes because the latter may be already highly optimized in an important process (eg vision). However faster evolution is expected and observed at most residue positions in the C2 and C3 loops of individual opsins. This variation -- to the extent the sole selective pressure is signaling -- describes the sensitivity of an unvarying fifth Galpha helix to changes in its shiftable pocket. That can take many forms in various opsin lineages without compromising their signaling (or improving it either).
(to be continued)
Selected alpha subunit reference sequences
>GNAT2_homSap Homo sapiens (human) Gt cone 8 exons chr1:109,952,320 tandem GNAi3 0 MGSGASAEDKELAKRSKELEKKLQEDADKEAKTVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHQDGYSPEECLEFKAIIYGNVLQSILAIIRAMTTLGIDYAEPSCA 0 0 DDGRQLNNLADSIEEGTMPPELVEVIRRLWKDGGVQACFERAAEYQLNDSASY 2 1 YLNQLERITDPEYLPSEQDVLRSRVKTTGIIETKFSVKDLNFR 2 1 MFDVGGQRSERKKWIHCFEGVTCIIFCAALSAYDMVLVEDDEV 0 0 NRMHESLHLFNSICNHKFFAATSIVLFLNKKDLFEEKIKKVHLSICFPEYD 1 2 GNNSYDDAGNYIKSQFLDLNMRKDVKEIYSHMTCATDTQNVKFVFDAVTDIIIKENLKDCGLF* 0 >GNAI3_homSap Homo sapiens (human) Gi 8 exons chr1:109,916,342 tandem GNAT2 stimulatory K channels 0 MGCTLSAEDKAAVERSKMIDRNLREDGEKAAKEVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGYSEDECKQYKVVVYSNTIQSIIAIIRAMGRLKIDFGEAARA 0 0 DDARQLFVLAGSAEEGVMTPELAGVIKRLWRDGGVQACFSRSREYQLNDSASY 2 1 YLNDLDRISQSNYIPTQQDVLRTRVKTTGIVETHFTFKDLYFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEM 0 0 NRMHESMKLFDSICNNKWFTETSIILFLNKKDLFEEKIKRSPLTICYPEYT 1 2 GSNTYEEAAAYIQCQFEDLNRRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKECGLY* 0 >GNAT1_homSap Homo sapiens (human) Gt rod 8 exons chr3:50,206,500 tandem GNAi2 intervening +SLC38A3 inversion 0 MGAGASAEEKHSRELEKKLKEDAEKDARTVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHQDGYSLEECLEFIAIIYGNTLQSILAIVRAMTTLNIQYGDSARQ 0 0 DDARKLMHMADTIEEGTMPKEMSDIIQRLWKDSGIQACFERASEYQLNDSAGYY 2 1 LSDLERLVTPGYVPTEQDVLRSRVKTTGIIETQFSFKDLNFR 2 1 MFDVGGQRSERKKWIHCFEGVTCIIFIAALSAYDMVLVEDDEV 0 0 NRMHESLHLFNSICNHRYFATTSIVLFLNKKDVFFEKIKKAHLSICFPDYD 1 2 GPNTYEDAGNYIKVQFLELNMRRDVKEIYSHMTCATDTQNVKFVFDAVTDIIIKENLKDCGLF* 0 >GNAI2_homSap Homo sapiens (human) Gi 8 exons chr3:50,260,220 tandem GNAT1 beta-adrenergic cAMP-inhibiting response 0 MGCTVSAEDKAAAERSKMIDKNLREDGEKAAREVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGYSEEECRQYRAVVYSNTIQSIMAIVKAMGNLQIDFADPSRA 0 0 DDARQLFALSCTAEEQGVLPDDLSGVIRRLWADHGVQACFGRSREYQLNDSAAY 2 1 YLNDLERIAQSDYIPTQQDVLRTRVKTTGIVETHFTFKDLHFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLAEDEEM 0 0 NRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKITHSPLTICFPEYT 1 2 GANKYDEAASYIQSKFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAT3_homSap Homo sapiens (human) Gt 8 exons chr7:79,925,923 tandem GNAi1 0 MGSGISSESKESAKRSKELEKKLQEDAERDARTVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHKNGYSEQECMEFKAVIYSNTLQSILAIVKAMTTLGIDYVNPRSA 0 0 EDQRQLYAMANTLEDGGMTPQLAEVIKRLWRDPGIQACFERASEY 2 1 QLNDSAAYYLNDLDRITASGYVPNEQDVLHSRVKTTGIIETQFSFKDLHFR 2 1 MFDVGGQRSERKKWIHCFEGVTCIIFCAALSAYDMVLVEDEEV 0 0 NRMHESLHLFNSICNHKYFSTTSIVLFLNKKDIFQEKVTKVHLSICFPEYT 1 2 GPNTFEDAGNYIKNQFLDLNLKKEDKEIYSHMTCATDTQNVKFVFDAVTDIIIKENLKDCGLF* 0 >GNAI1_homSap Homo sapiens (human) Gi 8 exons chr7:79,644,368 tandem GNAT3 beta-adrenergic cAMP-inhibiting response 0 MGCTLSAEDKAAVERSKMIDRNLREDGEKAAREVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEAGYSEEECKQYKAVVYSNTIQSIIAIIRAMGRLKIDFGDSARA 0 0 DDARQLFVLAGAAEEGFMTAELAGVIKRLWKDSGVQACFNRSREYQLNDSAAY 2 1 YLNDLDRIAQPNYIPTQQDVLRTRVKTTGIVETHFTFKDLHFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEM 0 0 NRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKKSPLTICYPEYA 1 2 GSNTYEEAAAYIQCQFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAO1_homSap Homo sapiens (human) Go 8 exons chr16:54,861,182 not in tandem 0 MGCTLSAEERAALERSKAIEKNLKEDGISAAKDVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGFSGEDVKQYKPVVYSNTIQSLAAIVRAMDTLGIEYGDKERK 0 0 ADAKMVCDVVSRMEDTEPFSAELLSAMMRLWGDSGIQECFNRSREYQLNDSAKY 2 1 YLDSLDRIGAADYQPTEQDILRTRVKTTGIVETHFTFKNLHFR 2 1 LFDVGGQRSERKKWIHCFEDVTAIIFCVALSGYDQVLHEDETT 0 0 NRMHESLKLFDSICNNKWFTDTSIILFLNKKDIFEEKIKKSPLTICFPEYT 1 2 GPSAFTEAVAYIQAQYESKNKSAHKEIYTHVTCATDTNNIQFVFDAVTDVIIAKNLRGCGLY* 0 >GNAZ_homSap Homo sapiens (human) Gi 2 exons chr22:21,769,945 chicken/fish/Callo/lamp/no ciona/no branch/no urch too not tandem pertussis-insensitive balance cochlear dopamine serotonin 0 MGCRQSSEEKEAARRSRRIDRHLRSESQRQRREIKLLLLGTSNSGKSTIVKQMKIIHSGGFNLEACKEYKPLIIYNAIDSLTRIIRALAALRIDFHNPDRAYDAVQLFALTGPAESKGEI TPELLGVMRRLWADPGAQACFSRSSEYHLEDNAAYYLNDLERIAAADYIPTVEDILRSRDMTTGIVENKFTFKELTFKMVDVGGQRSERKKWIHCFEGVTAIIFCVELSGYDLKLYEDNQT 0 0 SRMAESLRLFDSICNNNWFINTSLILFLNKKDLLAEKIRRIPLTICFPEYKGQNTYEEAAVYIQRQFEDLNRNKETKEIYSHFTCATDTSNIQFVFDAVTDVIIQNNLKYIGLC* 0 >GNAQ_homSap Homo sapiens (human) Gq 7 exons --tandem to GNA14 chr9:79,680,511 phospholipase C-beta melanopsin signaling 0 MTLESIMACCLSEEAKEARRINDEIERQLRRDKRDARRELKLLLL 1 2 GTGESGKSTFIKQMRIIHGSGYSDEDKRGFTKLVYQNIFTAMQAMIRAMDTLKIPYKYEHNKA 2 1 HAQLVREVDVEKVSAFENPYVDAIKSLWNDPGIQECYDRRREYQLSDSTKY 2 1 YLNDLDRVADPAYLPTQQDVLRVRVPTTGIIEYPFDLQSVIFR 2 1 MVDVGGQRSERRKWIHCFENVTSIMFLVALSEYDQVLVESDNE 0 0 NRMEESKALFRTIITYPWFQNSSVILFLNKKDLLEEKIMYSHLVDYFPEYD 1 2 GPQRDAQAAREFILKMFVDLNPDSDKIIYSHFTCATDTENIRFVFAAVKDTILQLNLKEYNLV* 0 >GNA14_homSap Homo sapiens (human) Gq 7 exons --tandem to GNAQ chr9:79,340,705 phospholipase C-beta delta opioid receptors 0 MAGCCCLSAEEKESQRISAEIERQLRRDKKDARRELKLLLL 1 2 GTGESGKSTFIKQMRIIHGSGYSDEDRKGFTKLVYQNIFTAMQAMIRAMDTLRIQYVCEQNKE 2 1 NAQIIREVEVDKVSMLSREQVEAIKQLWQDPGIQECYDRRREYQLSDSAKY 2 1 YLTDIDRIATPSFVPTQQDVLRVRVPTTGIIEYPFDLENIIFR 2 1 MVDVGGQRSERRKWIHCFESVTSIIFLVALSEYDQVLAECDNE 0 0 NRMEESKALFKTIITYPWFLNSSVILFLNKKDLLEEKIMYSHLISYFPEYT 1 2 GPKQDVRAARDFILKLYQDQNPDKEKVIYSHFTCATDTDNIRFVFAAVKDTILQLNLREFNLV* 0 >GNA11_homSap Homo sapiens (human) Gq 7 exons ++tandem to GNA15 chr19:3,058,931phospholipase C-beta ubiquitous 0 MTLESMMACCLSDEVKESKRINAEIEKQLRRDKRDARRELKLLLL 1 2 GTGESGKSTFIKQMRIIHGAGYSEEDKRGFTKLVYQNIFTAMQAMIRAMETLKILYKYEQNKA 2 1 NALLIREVDVEKVTTFEHQYVSAIKTLWEDPGIQECYDRRREYQLSDSAKY 2 1 YLTDVDRIATLGYLPTQQDVLRVRVPTTGIIEYPFDLENIIFR 2 1 MVDVGGQRSERRKWIHCFENVTSIMFLVALSEYDQVLVESDNE 0 0 NRMEESKALFRTIITYPWFQNSSVILFLNKKDLLEDKILYSHLVDYFPEFD 1 2 GPQRDAQAAREFILKMFVDLNPDSDKIIYSHFTCATDTENIRFVFAAVKDTILQLNLKEYNLV* 0 >GNA15_homSap Homo sapiens (human) Gq 7 exons ++tandem to GNA11 chr19 3,100,978 phospholipase C-beta hematopoietic cells 6x faster 0 MARSLTWRCCPWCLTEDEKAAARVDQEINRILLEQKKQDRGELKLLLL 1 2 GPGESGKSTFIKQMRIIHGAGYSEEERKGFRPLVYQNIFVSMRAMIEAMERLQIPFSRPESKHH 2 1 ASLVMSQDPYKVTTFEKRYAAAMQWLWRDAGIRAYYERRREFHLLDSAVY 2 1 YLSHLERITEEGYVPTAQDVLRSRMPTTGINEYCFSVQKTNLR 2 1 IVDVGGQKSERKKWIHCFENVIALIYLASLSEYDQCLEENNQE 0 0 NRMKESLALFGTILELPWFKSTSVILFLNKTDILEEKIPTSHLATYFPSFQ 1 2 GPKQDAEAAKRFILDMYTRMYTGCVDGPEGSKKGARSRRLFSHYTCATDTQNIRKVFKDVRDSVLARYLDEINLL* 0 >GNAL_homSap Homo sapiens (human) Gs 12 exons chr18:11,679,824 imprinted dopamine receptors D1 and D5 Golf alpha 0 MGCLGGNSKTTEDQGVDEKERREANKKIEKQLQKERLAYKATHRQTHRLLLL 1 2 GAGESGKSTIVKQMRILHVNGFNPE 2 1 EKKQKILDIRKNVKDAIV 0 0 TIVSAMSTIIPPVPLANPENQFRSDYIKSIAPITDFEYSQ 0 0 EFFDHVKKLWDDEGVKACFERSNEYQLIDCAQY 2 1 FLERIDSVSLVDYTPTDQ 00 DLLRCRVLTSGIFETRFQVDKVNFH 2 1 MFDVGGQRDERRKWIQCFN 1 2 DVTAIIYVAACSSYNMVIREDNNTNRLRESLDLFESIWNNR 2 1 WLRTISIILFLNKQDMLAEKVLAGKSKIEDYFPEYANYTVPED 1 2 ATPDAGEDPKVTRAKFFIRDLFL 0 0 RISTATGDGKHYCYPHFTCAVDTENIRRVFNDCRDIIQRMHLKQYELL* 0 >GNAS_homSap Homo sapiens (human) Gs 13 exons complex imprinted expression 0 MRKEALEKRAQKRAEKKRSKLIDKQLQDEKMGYMCTHRLLLL 1 2 GAGESGKSTIVKQMRILHVNGFNGE 2 1 EKATKVQDIKNNLKEAIETIV 0 0 AAMSNLVPPVELANPENQFRVDYILSVMNVPDFDFPP 0 0 EFYEHAKALWEDEGVRACYERSNEYQLIDCAQY 2 1 FLDKIDVIKQADYVPSDQ 00 DLLRCRVLTSGIFETKFQVDKVNFH 2 1 MFDVGGQRDERRKWIQCFN 1 2 DVTAIIFVVASSSYNMVIREDNQTNRLQEALNLFKSIWNNR 2 1 WLRTISVILFLNKQDLLAEKVLAGKSKIEDYFPEFARYTTPED 1 2 ATPEPGEDPRVTRAKYFIRDEFLRIST 0 0 ASGDGRHYCYPHFTCAVDTENIRRVFNDCRDIIQRMHLRQYELL* 0 >GNA12_homSap Homo sapiens (human) G12 4 exons chr7:2,792,376 MDCK cell tight junction 0 MSGVVRTLSRCLLPAEAGGARERRAGSGARDAEREARRRSRDIDALLARERRAVRRLVKILLLGAGESGKSTFLKQMRIIHGREFDQKALLEFRDTIFDNILK 0 0 GSRVLVDARDKLGIPWQYSENEKHGMFLMAFENKAGLPVEPATFQLYVPALSALWRDSGIREAFSRRSEFQL 0 0 GESVKYFLDNLDRIGQL 0 0 NYFPSKQDILLARKATKGIVEHDFVIKKIPFKMVDVGGQRSQRQKWFQCFDGITSILFMVSSSEYDQVLMEDRRTNRLVESMNIFETIVNNKL FFNVSIILFLNKMDLLVEKVKTVSIKKHFPDFRGDPHRLEDVQRYLVQCFDRKRRNRSKPLFHHFTTAIDTENVRFVFHAVKDTILQENLKDIMLQ* 0 >GNA13_homSap Homo sapiens (human) G12 4 exons chr17:60,460,255 peculiar phase shift verified exon 1 0 MADFLPSRSVLSVCFPGCLLTSGEAEQQRKSKEIDKCLSREKTYVKRLVKILLLGAGESGKSTFLKQMRIIHGQDFDQRAREEFRPTIYSNVIK 1 2 GMRVLVDAREKLHIPWGDNSNQQHGDKMMSFDTRAPMAAQGMVETRVFLQYLPAIRALWADSGIQNAYDRRREFQL 0 0 GESVKYFLDNLDKLGEP 0 0 DYIPSQQDILLARRPTKGIHEYDFEIKNVPFKMVDVGGQRSERKRWFECFDSVTSILFLVSSSEFDQVLMEDRLTNRLTESLNIFETIVNNRVFSNVSIILFLNKTDLLEEKVQIVS IKDYFLEFEGDPHCLRDVQKFLVECFRNKRRDQQQKPLYHHFTTAINTENIRLVFRDVKDTILHDNLKQLMLQ* 0 >GNAT2_galgal Gallus gallus (chicken) cone-type transducin alpha AF200339 missing in genome 90% 0 MGSGASAEDKEMAKRSKELEKKLQEDADKEAKTVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHQDGYTPEECMEFKAVIYGNILQSILAIIRAMSTLGIDYAESGRA 0 0 DDGRQLFNLADSIEEGTMPPELVDCIKKLWKDGGVAGVFDRAAEYQLNDSAAY 2 1 YLNQLDRITAPGYLPNEQDVLRSRVKTTGIIETKFSVKDLNFR 2 1 MFDVGGQRSERKKWIHCFEGVTCIIFCGALSAYDMVLVEDDEV 0 0 NRMHESLHLFNSICNHKFFAATSIILFLNKKDLFEEKIKKVHLSICFPDYD 1 2 GPNTFEDAGNYIKTQFLDLNMRKDVKEIYSHMTCATDTQNVKFVFDAVTDVIIKENLKDCGLF* 0 >GNAI3_galgal Gallus gallus (chicken) NP_989580 0 MGCTLSAEERAALERSKAIEKNLKEDGISAAKDVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGYSEEECKQYKVVVYSNTIQSIIAIIRAMGRLKIDFGEVARA 0 0 DDARQLFVLAGSAEEGVMTAELAGVIKRLWRDAGVQACFSRSREYQLNDSASY 2 1 YLNDLDRISQPTYIPTQQDVLRTRVKTTGIVETHFTFKDLYFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEM 0 0 NRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKKSPLTICYPEYT 1 2 GSNTYEEAAAYIQCQFEDLNRRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKECGLY* 0 >GNAT1_galgal Gallus gallus (chicken) rod-type transducin alpha AF200338 missing in genome 96% 0 MGAGASAEEKHSRELEKKLKEDAEKDARTVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHQDGYSLEECLEFIAIIYSNTLQSMLAIVRAMTTLNIQYGDSARQD 0 0 DARKLLHLSDTIEEGTMPKEMSDIIGRLWKDAGIQACFDRASEYQLNDSAGY 2 1 YLSDLERLVTPGYVPTEQDVLRSRVKTTGIIETQFSFKDLNFR 2 1 MFDVGGQRSERKKWIHCFEGVTCIIFIAALSAYDMVLVEDDEV 0 0 NRMHESLHLFNSICNHRYFATTSIVLFLNKKDVFLEKIKKAHLSICFPDYD 1 2 GPNTYDDAGNYIKLQFLELNMRRDVKEIYSHMTCATDTENVKFVFDAVTDIIIKENLKDCGLF* 0 >GNAI2_galgal Gallus gallus (chicken) NM_205402 95% 0 MGCTVSAEDKAAAERSRMIDRNLREDGEKAAREVKLLLL 1 2 GAGESGKSTIVKQMKIIHEDGYSEEECRQYKAVVYSNTIQSIMAIIKAMGNLQIDFGDSSRAD 0 0 DARQLFALACTAEEQGIMPEDLANVIRRLWADHGVQACFNRSREYQLNDSAAY 2 1 YLNDLERIARADYIPTQQDVLRTRVKTTGIVETHFTFKDLHFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLAEDEEM 0 0 NRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIVHSPLTICFPEYT 1 2 GANKYDEAAGYIQSKFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAT3_galgal Gallus gallus (chicken) 81% +GNAT3-GNAI1 tandem 0 MGGGASSESKESARRSRELEKKLQEDAEREARTVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHKDGFTYQERMEFRPIIYSNTVQSILSIVKAMTKLGISYENPARI 0 0 EDERKLCDMETNLDDSNMSSELVELIKQLWKDGGIQACFARASEYELNDSAAy 2 1 YLNDLDRLAMPDYVPSEQDVLHSRVKTTGIIETQFSFKDLNFR 2 1 MFDVGGQRSERKKWIHCFEGVTCIIFCAALSAYDMVLVEDKEV 0 0 NRMHESLQLFNSICNHRCFATTSIVLFLNKKDLFQEKIAKVHLNICFPEYN 1 2 GLNTFEDAGNYIKKQFLDLNIRKEDKEIYCHLTCATDTQNVKFVFDAVTDIIIKENLKDCGLF* 0 >GNAI1_galgal Gallus gallus (chicken) NM_205403 98% +GNAT3-GNAI1 tandem 0 MGCTLSAEDKAAVERSKMIDRNLREDGEKAAREVKLLLL 1 2 GAGESGKSTIVKQMKIIHEAGYSEEECKQYKAVVYSNTIQSIIAIIRAMGRLKIDFGDPTRAD 0 0 DARQLFVLAGAAEEGFMTADVAGVIKRLWKDSGVQACFNRSREY 2 1 QLNDSAAYYLNDLDRIAQTSYIPTQQDVLRTRVKTTGIVETHFTFKDLHFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEM 0 0 NRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKRSPLTICYPEYA 1 2 GSNTYEEAAAYIQCQFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAQ_galgal Gallus gallus (chicken) NP_001026598 98% -GNA14-GNAQ tandem chrZ 0 MTLESIMACCLSEEAKEARRINDEIERQLRRDKRDARRELKLLLL 1 2 GTGESGKSTFIKQMRIIHGSGYSDEDKRGFTKLVYQNIFTAMQAMIRAMDTLKIPYKYEHNKA 0 0 HAQLVREVDVEKVSTFENPYVDAIRSLWNDPGIQECYDRRREYQLSDSTKY 2 1 YLNDLDRIADSTYLPTQQDVLRVRVPTTGIIEYPFDLQSVIFR 2 1 MVDVGGQRSERRKWIHCFENVTSIMFLVALSEYDQVLVESDNE 0 0 NRMEESKALFRTIITYPWFQNSSVILFLNKKDLLEEKIMYSHLVDYFPEYD 1 2 GPQRDAQAAREFILKMFVDLNPDSDKIIYSHFTCATDTENIRFVFAAVKDTILQLNLKEYNLV* 0 >GNA14_galgal Gallus gallus (chicken) -GNA14-GNAQ tandem chrZ 0 MAGRCLSADEKESQRISAEIERQLRRDKRDARRELKLLLL 1 2 GTGESGKSTFIKQMRIIHGSGYTEEDRKGFTKLVYRNIFTAMQAMIRAMDILKIQYASEENEV 2 1 NAQMIRRVEVDKVTALERKQVEAIKNLWDDPGIQECYDRRREYQLSDSAY 2 1 YLTNIDRIAMPSFVPTQQDILRVRVPTTGIIEYPFDLENVIFR 2 1 MVDVGGQRSERRKWIHCFESVTSIIFLVALSEYDQVLAECDNE 0 0 NQMKESKALFKTIITYPWFLNSSVILFLNKKDLLEEKIMYSHLTSYFPEYT 1 2 GPKQDVKAAGDFILKLYQDQNPDKQKVIYSHFTCATDTENIRFVFAAVKDTILQLNLREFNLV* 0 >GNA11_galgal Gallus gallus (chicken) 7 exons AF364328 97% -GNA11 no tandem 0 MTLESMMACCLSDEVKESKRINAEIEKQLRRDKRDARRELKLLLL 1 2 GTGESGKSTFIKQMRIIHGSGYSEEDKKGFTKLVYQNIFTAMQSMIRAMETLKILYKYEQNKA 0 0 NAVLIREVDVEKVMTFEQPYVSAIKTLWNDPGIQECYDRRREYQLSDSAKY 2 1 YLSDVDRIATPGYLPTQQDVLRVRVPTTGIIEYPFDLENIIFR 2 1 MVDVGGQRSERRKWIHCFENVTSIMFLVALSEYDQVLVESDNE 0 0 NRMEESKALFRTIITYPWFQNSSVILFLNKKDLLEDKILYSHLVDYFPEFD 1 2 GPQRDAQAAREFILKMFVDLNPDSDKIIYSHFTCATDTENIRFVFAAVKDTILQLNLKEYNLV* 0 >GNAQ7a_calMil Callorhinchus milii TGESGKSTFIKQMRIIHGSGYTDEDKRGFTKLVYQNIFTAVQAMIRAMDTLKIQYKYDYNKV >GNA147b_calMil Callorhinchus milii TGESGKSTFIKQMRIIHGDGYSDEDRKCFTKLVYQNIFTAMQAMIKAMDTLRIQYKNGQN >GNAI18a_calMil Callorhinchus milii 8th exon GESGKSTFIKQMR 21 RIIHEDGYSEEECKQYKAVVYSNTIQSIIAIIRAMGRLKIDF >8b_calMil Callorhinchus milii 8th exon GESGKSTIVKQMK >GNAT2term1_calMil Callorhinchus milii no GNAT3, no GNAi3 no evidence of tandems GNNSFDDAGLYIKMQFLDLNMRKDVKEIYSHLTCATDTENVKFVFDAVTDIIIKENLKDCGLF* >GNAT1_calMil Callorhinchus milii GPNTYEDAGNYIKLQFLELNMRKDVKEIYAHMTCATDTKNVKFVFDAVTDIIIKENLKECGLF* >GNAI1_calMil Callorhinchus milii GSNTYEEAAAYIQCQFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAI2term4_calMil Callorhinchus milii GANKYDEAAAYIQTKFEDLNKRKDTKEIYTHFTCATDTKHVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAZ_calMil Callorhinchus milii AAVX01066028 (97%) exon2 0 SRMAESLRLFDSICNNNWFINTSLILFLNKKDLLAEKIKRIPLTVCFPEYKGQNTYEEAAVYIQRQFEDLNRNKETKEIYSHFTCATDTSNIQFVFDAVTDVIIQNNLKYIGLC* 0 >GNAO1term6_calMil Callorhinchus milii GPNSYEDAAAYIQAQFESKN RSPNKEIYCHLTCATDTNNIQVVFDAVTDIIIANNLRGCGLY* 0 >GNAZ_calMil Callorhinchus milii 0 SRMAESLRLFDSICNNNWFINTSLILFLNKKDLLAEKIKRIPLTVCFPEYKGQNTYEEAAVYIQRQFEDLNRNKETKEIYSHFTCATDTSNIQFVFDAVTDVIIQNNLKYIGLC* 0 >GNAT1_petMar Petromyzon marinus (lamprey) EU571208 short photoreceptor transducin-alpha subunit rod MGSGASAEDKDQAKHSKELEKKLAEDAEKDARTVKLLLLGAGESGKSTIVKQMKIIHQSGYSIEECMEFIAIIYSNTLQSILAIVRAMGTLSIDFGDSARMD DARQLQNLADSIDEGTMPQELYLIIKRLWTDSGIQVCFDRASEYQLNDSAEYYLTDIDRLVQPGYLPTEQDVLRSRVKTTGIIETQFSFKDLHFRMFDVGGQRSERKKWIHCFEGV TCIIFCAALSAYDMVLVEDDEVNRMHESLHLFNSICNHRYFNATSIVLFLNKKDLFEVKVKKAHLSICFPDYDGPNTYDDAGNFIKLQFLDLNMRKESKEIYSHMTCATDTKNVKFVFDAVTDIIIKENLKDCGLF >GNAT2_petMar Petromyzon marinus (lamprey) EU571207 long photoreceptor transducin-alpha subunit contig4334 cone short intron still 8 exons MGSGASAEDKESAKHSKELEKKLAEDAEKEARTVKLLLLGAGESGKSTIVKQMKIIHKNGYSEAECLEFKAIIYSNTLQSILAIVRAMETFSIDYGDPARAA DGRQLFNLADSLEEGSMPNELSAIIIRLWKDTGVQASFDRASEYQLNDSASYYLNDLDRLMNPSYLPNEQDVLRSRVKTTGIIEDSFCFKDLQFRMFDVGGQRSERKKWIHCFEGV TCIIFCGALSAYDMVLVEDDEVNRMHESLHLFNSICNHRYFNDTSIVLFLNKKDLFEEKVKKVHLNICFPDYDGPNTFDDAGAYIKNQFLDLNLRKEAKEIYSHLTCATDTQNVKFVFDAVTDIIIKNNLKDCGLF >GNAI1_petMar Petromyzon marinus (lamprey) MGCTLSTEDKAAVERSRMIDRNLREDGEKASREVKLLLLGASHT GAGESGKSTIVKQMK IIHEAGYTEEECKQYKAVVYSNTIQSVIAIIRAMGNLRIDFGDAGRA DDARQLFVLAGSAEDGLMTPELAQVIKRLWADPGVQACFRRAREYQLNDSAA YLNDLERISQPSYVPTQQDVLRTRVKTTGIVETHFTFKDLHFK MFDVGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLAEDEEM NRMHESMKLFDSICNNKWFIETSIILFLNKKDLFEEKVIRSPLTICYPEYTGS AGGNTYEEAAAYIQTQFENLNKRKESKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAI2_petMar Petromyzon marinus (lamprey) frag GAGESGKSTIVKQMK 21 IIHEDGYSEDECKQYTAVVFSNAIQSIIAIIRAMGKLKIDFGDVSRA EDARQLFVLAGVAEE-GVMTPDLSEVIKRLWSDSGVQACFRRSREYQLNDSAA YLNDLERISNLSYIPTQQDVLRTRVKTTGIVETHFTFKDLHFK MFDVGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLAEDEE NRMHESMKLFDSICNNKWFTETSIILFLNKKDLFEEKINKSPLFICFAEYFG >GNAZ_petMar Petromyzon marinus (lamprey) frag exon1 RAYDAVQLFALTGPAESKGEISPELLAIMRRLWCDPGVQLCFGRSSEYHLEDNAAYYLGDLERIAAPGYVPTVEDILRSRDMTTGIVENRFTFKELTFKMVDVGGQRSERKKWIHCFEGVTAIIFCVELSGYDLKLYEDNLT 0 >GNAI_eptBur Eptatretus burgeri (hagfish) BJ646870 frag leukocytes 78% chicken intermediate MGCTISNAEEREAAEQSRNIDRGLHQDHVRSLREIKLLLLGAGESGKSTIVKQMKIIHDT GFSQEECKKYRAVVYSNTIQSMVAILHAMGKLHIDFGDPDRANDARQLFDLANMVEDCSF PTEICMVLAHLWADSGVQACFARSREYQLNDSAAYYLNDLERLGQEEYVPTQEDVL RTRVKTTGIVETHFTFKELHFKMFDVGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLA EDEEMNRMHESMKLFDSICNNKWFTETSIILFLNKKDLFEEKIERSPLTICFPEYTGRNT YDCASVFIRTQFENLNKRKDTKEIYSHFTCATDTENVQFVFDAVTDVIIKNNLKDCGLF >GNAI_cioInt Ciona intestinalis G protein alpha 8 exons not tandem PMID: 12426469 expressed in ocellus alt YLDSLDRLTEPRYVPTQQDVLRTRVKTTGIVEVDFNFKGLTFK 0 MGCTVSTDDKAANERSRAIDRNLRVDGDKQSREVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGYSEEECLQYKAVVYSNTLQSLITIVRAMGNLKIDFGSSDRA 0 0 DDARQLFSLAGSLEDGEMTQELGDCMKRMWGDKGVQVCFNRSREFQLNDSAQY 2 1 YLDSLDRLVASDYVPTEQDVLRSRVKTTGIVETQFEHKDLHFKMFD 2 1 VGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLAEDEEMNRM 0 0 HESMKLFDSICNNKWFTETSIILFLNKKDLFEVKILKSPLSICFPEYP 1 2 GQNTYAEAAAYIQLQFEDLNKRKDSKEIYTHFTCATDTTNIQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAQ_cioInt Ciona intestinalis G protein alpha 7 exons not tandem 0 MPLMTILANCCKSSDEIEAEKINGQIERELRRHKKDARRELKLLLL 1 2 GTGESGKSTFIKQMK IIHGAGYSDEDKRSFIKLVYQNIVTSIQNMSAAMQTLNLEYEIEENNE 2 1 HAEEIREVQVDKISSYDDFITNISYIECLWKDTGIQKCYDRRREYQLSDSTY 2 1 YYLSDLDRIKKPDFLPTQQDILRVRIPTTGIIEYPFDLDQIIFR 2 1 MVDVGGQRSERRKWIHCFENVTSIIFLVALSEYDQVLVEAGNE 0 0 NRMEESKALFRTIITYPWFDGSSVILFLNKKDLLEEKIAYSDLADYFPQFD 1 2 GPPKNADAAREFILGMFVELNPNKDKIVYSHFTCATDTENIRFVFAAVRDTILQANLKEYNLV* 0 >GNAO1_braFlo Branchiostoma floridae (amphioxus) ABEP01019035 83% 8 exons 0 MGCTMSAEERAAIEKTKQIDKNLKEDGLVAAKDIKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGFTTDDMQQFKPVVYSNTIQSLTSILRAMEVLKVEYG 0 0 DAKMVFEVVQRMEDTEPFSPELLAAMKRLWTDKGVQECFSRANEYQLNDSAK 2 1 YLDDLDRLGADEYEPTEQDILRTRVKTTGIVETHFTFKNLNFR 2 1 LFDVGGQRSERKKWIHCFEDVTAIIFVAALSGYDLVLHEDETT 0 0 NRMHESLKLFDSICNNKWFTETSIILFLNKKDLFEEKITRSPLTMAFPEYT 1 2 PPGPNTYTEAAAYVQAQFESKNKSPNKEIYTHMTCATDTSNIQFVFDAVTDVIIANNLRGCGLY* 0 >GNAI1_braFlo Branchiostoma floridae (amphioxus) ABEP01040635 BW845279 mrna 91% still has 8th exon 0 MGCAISAEDKAAAERSKMIDKNLRADGEKAAREVKLLLL 1 2 GAGESGKSTIVKQMK 21 IIHEDGYSEEECMQYKAVVYSNTIQSLIAIIRAMGTLKIDFG 0 0 DDARQLFALASTAEEGEMTPELAGIMKRLWADGGVQACFGRSREYQLNDSASR 2 1 YLNSLDRLAAGGYVPTQQDVLRTRVKTTGIVETHFTFKDLHFK 2 1 MFDVGGQRSERKKWIHCFEGVTAIIFCVALSAYDLVLAEDEETVGR 0 0 NRMHESMKLFDSICNNKWFTETSIILFLNKKDLFEEKITKSPLTICYPEYT 1 2 GGSNTYEEAAAYIQMQFEDLNKRKETKEIYTHFTCATDTNNIQFVFDAVTDVIIKNNLKDCGLF* 0 >GNAQ_braFlo Branchiostoma floridae (amphioxus) ABEP01058052 ABEP01054441 frag very high percent id 7 exons 1 MNKMACCLSEEAKEQKRINQEIEKQLRKDKRDARRELKLLLL 1 2 TGESGKSTFIKQMRIIHGAGYSDEDRRGYTKLVYQNIFMAMHSMIRAMDTLKIAYKNKENE 0 0 SVSTFEKEYVEAIQSLWEDAGIQECYDRRREYQLTDSAKY 2 1 YLSDLERIAQPDYLPTEQDVLRVRVPTTGIIEYPFDLDNVIFR 2 1 MVDVGGQRSERRKWIHCFENVTSIMFLVALSEYDQVLVESDNE 0 0 NRMEESKALFRTIITYPWFQNSSVILFLNKKDLLEEKIMYSHLVDYFPEFD 1 2 GPQRDAQAAREFILKMFVDLNPDSDKIIYSHFTCATDTENIRFVFAAVKDTILQLNLKEYNLV* 0 >GNAS_braFlo Branchiostoma floridae (amphioxus) FE588508 mrna 73% KILHQNSFDEQERRQKIADIKKNIRDAIITITGAMSTLTPPVPLADHTLQARVDY IQDVATQPEFSYPPEFYEHTELLWKDGGVQACYERSNEYQLIDCAQYFLDRVHVVKQPDY EPTDQDILRCRVLTSGIFETKFEVNDVKFHMFDVGGQRDERRKWIQCFNDVTAIIFVVACSSYNMVLREDPSQNRLREALDLFKSIWNNRWLRTISVILFLNKQDLLKQKV >GNA12_braFlo Branchiostoma floridae (amphioxus) BW845279 ABEP01001798 74% EYIPSKQDVLYARKATKGIVEHEFDIKGIPFLMVDVGGQRSQRQKWFQCFESVTSILFLVSSSEFDQVLMEDRKTNRLVESLNIFETIVNNKTFTEVSIILFLNKTDLLQDKVTYVSIKE YFPEFPEMSDPHN-LTDVQNFILNLF-DAKRRERNKPLFHHFTTAVDTENIKFVFHAVKDTILQDNLKQLML >GNA13_braFlo Branchiostoma floridae (amphioxus) BW845279 ABEP01001790 63% QDIEQRQRSKQIDKMLAKEKVHLRRQVKILLLGAGESGKSTFLKQMRIIHGKDFDVEALKEYRPTVYNNIVKGMKVLVDAQRKLGIKMKEPSNELYCDQVMKFEGTIKIDTALF LEYCPAIRALWSDAGIQEAWDRRREFQLVRNSSSYNLEYIPSKQDVLYARKATKGIVEHEFDIKGIPFLMVDVGGQRSQRQKWFQCFESVTSILFLVSSSEFDQVLMEDRKTNRLVESLNIFET IVNNKTFTEVSIILFLNKTDLLQDKVTYVSIKEYFPEFPEMSDPHNLTDVQNFILNLFDAKRRERNKPLFHHFTTAVDTENIKFVFHAVKDTILQDNLKQLML >GNAQ_strPur Strongylocentrotus purpuratus NM_001001475 PUBMED 15003628 MACCLSEEAKEQKRINQEIEKQLRKDKRDARRELKLLLLGTGESGKSTFIKQMRIIHGAGYTEEDRKTFTKLVYQNIFMAINAMIRAMDTLKIAYGDPTNEKKAQEVRLIDHETVTVFHEPYIGYVDCIWNDSGIQECYDRRREYQLTDSAKYYLSDLKR ISDSNYIPTEQDVLRVRVPTTGIIEYPFDLDSIIFRMVDVGGQRSERRKWIHCFENVTSIMFLVALSEYDQLLVESDSENRMEESKALFRTIITYPWFQNSSVILFLNKKDLLEEK IMHSHLVDYFPEFDGPSRDATAAREFILKMFVELNPDSDKIIYSHFTCATDTENIRFVFAAVKDTILQLNLKEYNLV >GNAI_strPur Strongylocentrotus purpuratus NM_001001475 PUBMED 15003628 still has short 8th exon MGCATSAEDKAAAERSKMIDRNLRLEGEKAAREVKLLLLGAGESGKSTIVKQMKIIHEEGYSEEDCRQYKPVVYSNTIQSMIAIIRAMGSLKIDFGDTERAD DARQLFALAGQAEEGELSTELAAVMKRLWADSGVQACFSRSREYQLNDSASYYLNALDRLSAPGYIPTQQDVLRTRVKTTGIVETHFTFKELHFKMFDVGGQRSERKKWIHCFEGV TAIIFCVALSAYDLVLAEDEEMNRMHESMKLFDSICNNKWFTETSIILFLNKKDLFEEKIQKSPLTICFPEYTGSNTYEEAAAYIQMQFEDLNKRKDQKEIYTHFTCATDTNNIQFVFDAVTDVIIKNNLKDCGLF >GNAO1_strPur Strongylocentrotus purpuratus genomic approx MGCAMSSEERESQERSKQIDKNLKEDGLQAARDVKLLLLG AGESGKSTIVKQMKIIHEEGFTAEDSKVYRPVVYSNLLQSMVSMLRAREKFETPFGEEEREDAQLVYDTVSKLQDSAPYSPSLTAAIQRLWTDSGLLEIFNRAREYQLNDSAK FLDNLDRIGSPDYLPNEQDILRTRVKTTGIVETHFTFKNLHFRFHLITCRLFDVGGQRSERKKWIHCFEDVTAIIFCVALSGYDQRLLEDDVTNRMQESLKLFDSI CNNKWFTDTSIILFLNKKDLFEEKIQKSPLTICFQEYTGANEYLPAAGYIQLQFEALNKSTNKEIYTHMTCATDTTNIQFVFDAVTDTIIANNLRGCGLY >GNAS_strPur Strongylocentrotus purpuratus NM_001001475 PUBMED 15003628 MGCFGNGLSSEEKDEEKKRKEANKKIEKQLQKDKQIYRATHRLLLLGAGESGKSTIVKQMRILHVDGFSPDERKKKIEDIRRNIRDAIITITGAMSTLSPPI QLAEPQNQFRLDYIQDVSSSPDFDYPEEFWDHTKHLWIDAGVQGCYDRSHEYQLIDSAQYFLDRVDTIRRPDYAPDLQDILRCRVLTSGIFETKFQVDKVNFHMFDVGGQRDERRK WIQCFNDVTAIIFVVACSSYNLVLREDPNQNRLRESLELFRSIWNNRWLRTISVILFLNKQDLLAEKVQAGRSKIEDYFSEYAMYTIPPDAATDTGEPEDVLRAKYFIRDEFLRISTASGDGRHYCYPHFTCAVDTENIRRVFDDCRDIIQRMHLRQYELL >GNA12_strPur Strongylocentrotus purpuratus NM_001001475 PUBMED 15003628 located on cytoplasmic vesicles MAGTLLTCCLTPTDKQALNHSKDIDKQLQRDKNYIRREVKVLLLGAGESGKSTFLKQMKIIHEQQFTDQEVKEFRNIIYGNIIKGMKVLADARDKLGIPWGD SGNEKHAEFVMSFNTQAAQLEPPLFVQYVQPCVELWKDSGIQSAFDRRREFQLADSVKYFLDEIDRVGRKDYIPSLTDILHSRKATKAFQEHVIDIRNVPFRFVDVGGQRSQRQKW FQCFESVTSILFLASSSEFDQVLMEDRITNRLLESCNIFDTIVNHKCFASISIILFLNKTDLLEEKIKHVSIKDYFPNFQGDPHSMNDVQNFILKMFDVRRRERGSKALFHYFTTAVDTNNIRYVFQAVRDTILQENLKRLMLQ >GNAI1_triAdh Trichoplax adhaerens (placazoa) XM_002115978 77% homSap 71% GNAi2 still 8 exons +GNAI2_triAdh +GNAI1_triAdh MGCAASAGDKVAAAKSKEIDKKIKSDAEKAAREVKLLLLGAGESGKSTIVKQMRIIHESGFSEEDRAQYKPVVFSNTMQSMAAIIRAMGVLRIEFGDKTS LVGDARRLFEIMDAPGVQEFTPEIVSLLKRLWSDHGVQQCFSRSREYQLNDSAPYYLNSIDRLGKPEYIPSEQDVLRTRVKTTGIVETHFTFKDLHFKMF DVGGQRSERKKWIHCFEGVTAIIFCVSLSAYDLVLAEDEEMNRMMESMKLFDSICNNKWFTETSIILFLNKKDLFQEKILKSPLTICFPEYTGANTYEEA SAYIQMKFEDLNKMKDQKEIYTHFTCATDTNNIQFVFDAVTDVIIKNNLKDCGLF* >GNAI2_triAdh Trichoplax adhaerens (placazoa) XM_002115977 70% homSap 56% GNAi3 MGCLVSKDERAAAERSKIIDKNLKASGDVSAKEVKLLLLGAGESGKSTIVKQMRIIHEKGYSEQDCVQYRPVVYNNTVQSLATIIRACGPLGIPFENPSL KDLSKEYFSMIERQGDSVELSKKLLTLMKTIWADNGIQESFKRSREYQLNDSAGYYLNDIDRLGTSNYIPTQQDVLRTRVKTTGIVETQFSFRDFRFKMV DVGGQRSERKKWIHCFEGVTAIIFCVSLSAYDLKLAEDEEMNRMVESMRLFDSICNNQFFEETSIILFLNKKDLFQQKIAVSPLTLCFPEYSGANNYQEA SSYIQTVFEDLNRKKESKEIYTHFTCATDTDNIQFVFDAVTDVIIKNNLKDCGLF* >GNAI3_triAdh Trichoplax adhaerens (placazoa) XM_002116075 60% homSap 61% GNAi1 MGITVSGEDKAAREKSTDIDKKIQNEKDKSLSEVKLLLLGAGESGKSTIAKQMRIIHESGYSDEDRQQYKSIIHCNAIYSLKAIIEAMKVLKIDISRSHT KIDAEDFLRLIYDSPDEVTPELKKIMKRLWNDPDVQKCFNRSREYQLMDSASYYLDDLDRLVQDSYLPSEQDILRARVKTSSIKETEFEYKGLEFKMIDV GGQRSERRKWIHCFENVTAVIFCAALSAYDLVLQEDYFTNRMKESLNLFDSVCNNQWFKKTSIILFLNKTDIFKEKIRKSPITTCFPEYNGTNSYEETTS YIQKKFISLNSNGKEKTIYSHFTCATDTENIVFVFAAVTDVILQKNIKEHGLLF* >GNAO1_triAdh Trichoplax adhaerens (placazoa) XM_002111534 53% homSap 51% GNAi1 MGCGSSTVDQKAVIANNQIEKDIREQELQAKKIIKLLLLGAAESGKSTIAKQLKIIHMEGFTKNDIEKAKPIIYSNIVHTFIQILQNMRPLKLEFNSEQR QADANQLFDIIGKMKDTDPYPPSVLKSMNALLADGGFQTTIKRGHEYHLHDSAEYFLKSLDRIGNDNYEPTEQDILRSRLRTTGVNQIEFEFKMLNFQVI DVGGQRSERRKWIHVFDSVTAIIFCVSLSCYDMTVYEDGNTNSMHESLKLFDWIVNNEFFKETSIILFLNKKDLFEEKIKSVSLTVCFPEYDGTKSYEDT SLFIQKQFIDRKQSSQKEIYCHLTCATDTQNISVVFDAVTDIVISNNLRNCGLL* >GNAQ_triAdh Trichoplax adhaerens (placazoa) XM_002116172 76% homSap 48% GNAi3 MACCLSDEAREQRRINREIEKELKKHKRDAKRELKLLLLGTGESGKSTFIKQMRIIHGKGYTDNDRAEFTQLVFQNIFTAIQALIKAMETLNITYEHQSN RQRVDVVRTVDPETVGSLSKEHVEAIDSIWNDSGVQECYDRRREYQLSDSAKYYLTDLHRLAEPNYLPTQQDILRVRAPTTGIIEYDFNLDTVMFRMVDV GGQRSERRKWIHCFENVTSIMFLVALSEYDQILAEADSQNRMEESKALFKTIITYPWFQNSSIILFLNKKDILEEKVQKSNIADYFPEYDGPPRDAQAGR EFILKMFVDLNPDSEKIIYSHFTCATDTENIRFVFAAVKDTILQFNLREYNLV* >GNAS_triAdh Trichoplax adhaerens (placazoa) XM_002116172 74% homSap 44% GNAQ MGCFGNQTEDSRLQKKENTRIERQLKKDKAAYRSTHRLLLLGAGESGKSTIVKQMRILHVDGFNEEEKRQKIADIKRNIRDSIVAIVTAMGTLTPPCTLANL NNQFRVDYITEIASADDFNYPPVFFEHTKELWKDQGVQQCYERSNEYQLIDCAKYFLDKIDVVKLPDYQPTDQDVLRCRVLTSGIVETRFQVERVNFHMFDVGGQRDERRKWIQCF NDVTAIIFVVACSSYNLVLREDPSQNRLKESLELFQTIWNNRWLKTISIILFLNKQDLLAEKVRAGRSKIEDYFSEFSRYTTPTDATTEPGDDENVKRAKYFIRDAFLRISTATGE GKHYCYPHFTCAVDTENIRRVFNDCRDIIQRMHLRQYELL* >GNA12_triAdh Trichoplax adhaerens (placazoa) XM_002116172 51% homSap 48% GNA13 MKRRNSKLIDKELSKEKKSRGRQIKILLLGAGESGKSTFLKQMRIIHGEEYSQKDLMEFKNLIYGNVVKNMRVLITARDSLGIKWANADYEDYAQELLAIDT KSTVFDYAAFMSYAGKVVDLWQDRAIQQTYDKRNLYQLSDSTYYFMDRMKSLMDKAYVPTKQDVLRSRKATTNIVELTLNINRVPFTFVDVGGQRSQRRKWLQCFEGVTSVLFLVS SCAYDQVLLEDNRTNRIVESCQIFDTIINNKFFAKVAIILFFNKTDILIEKVSLVSIKDYFPEFSRDPKKIEDVKHFLITMFEKVSNDQKRGLYHHFTTATDTENIKFVFNAVREM ILEENMSILMLQ* >GNA13_triAdh Trichoplax adhaerens (placazoa) XM_002109597 48% homSap 39% GNAQ MDTVLCFKANSERREQIRHSKIIDQEILQERTEYYKTIKILLLGASECGKSTFLKQMRILHGQDFDVQDLLEFRSIIYGNIIRIMKVLVTARRSFEIQWKDS SHQNYADQILNFNTKVNEIEPHEFVAVVDMIRELWLDEAIQETYRRRNEYILADSTKYFMDRLEVIGKEDYVPIRKDALRMRKATKTIVEFTTTINKIPFVFIDVGGQRSQRRKWL QCFESITAILFLAAASDYNQVSLEDRKTNRLLESLEIFGAIVNHELLAKASKILFLNKIDLLEERLTISNIKNFFSAFNGDENDLTTVKEFILQLFSNKMEANNDNDKSLYHHYTI ATDTENIKVVFRDVKQTILQERLGSLLLH* >GNAI_monBre Monosiga brevicollis 3 exons no short XM_001747738 0 MGICMSAEQKAQQARTAAVEAQLERDAQLASRTIKLLLL 1 2 GAGESGKSTLVKQMKIIHGDGFSNEELKSYKPTICDNLVHSMRAVLEAMGPLVIDIGDQVRPP 0 0 HAKVVLSYIELGTSGGLTPELTEALKALWADSGVQECFRRSNEYQLNDSAEYFFNNIDRIAQSNYLPTQEDVLRARVRTTGV IETTFRYKDLIYRMFDVGGQRSERRKWIHCFNDVTAVLFVAALSGYDMKLFEDQETNRIHESLTLFDAICNNSFFINTAIILFL NKTDLFSQKIARTPLKDYFPEYDGPPNNASEAKKFIAGMFKRLNKNPNKPVYEHFVCATETQNIRYVFDAVK* 0 >GNAQ_monBre Monosiga brevicollis no short XM_001745795 55% GNAQ_homSap 0 MPCGPPDETRRRSLAIDRQLRKERMSKQREYKILLL GTGESGKSTIIKQMRIIYGQGFNESDRLAYKPLVYRNIITSMKRMLDALDQLSLQLADSSLEEDAYDK LDVDVNTVDAIEPYYPLLKKLWNDNGIQQVFQRRNEYQLSDSTAYYYNRLDAVAAADYIPTVDDVLRSRQATTGIHEFEFDLDSVVFRMMDVGGQRSERRKWIHSFE 0 0 GVTSIIFIAACNEYDQVLAEDTNVNRMQESLALFGQIIQYHW 2 1 FANSSFILFLNKQDLLEEKVKTHPIKPFFPDYTGQE 0 0 GDYENIKKFIETMYRSRKPAGKDLYTHFTMATDTSNIQFVFNAVRSTLLRIHLKDYNLF* 0
See also: RBP3 (IRBP) | RPE65 | Usher: USH2A | Usher: CDH23 | LOXHD1 | Update Blog