Personal genomics: ACTN3
Introduction to ACTN3 comparative genomics
The alpha actinin gene ACTN3 is a coding gene on human chromosome 11, quite interesting in its own right but best known as ground zero in the debate over frivolity and unexpected consequences of personal genomics. This gene first of all needs careful and exhaustive re-annotation before considering this controversy because its existing peer-reviewed scientific literature (some 22 papers) is a mixture of pre-genomic era obsolescence and gross factual errors on matters such as expression (falsely stated as specific to skeletal muscle).
Some unfortunate historic terminology needs to be explained. Actinins were erroneously thought similar to actin in early studies of myofibrillar components; instead they are homologically unrelated proteins that happen to bind actin. These 'actinins' were then improperly divided into 3 classes (alpha, beta, gamma) before it became known that their respective gene families were wholly unrelated (not homologous). For example, 'beta' actinins refer to heterodimers functioning as actin barbed-end capping proteins in skeletal muscle; they are comprised of the distinct gene families CAPZA and CAPZB themselves non-homologous and thus further misnamed. 'Gamma' actinins refer to yet other unrelated genes.
In this article, actinin shall mean alpha actinin, ie a protein encoded by one of the four paralogous genes ACTN1-ACTN4. Gene names are used for both gene or gene product (as this is always clear from context). Genus and species are indicated with standard 6-letter code (eg ACTN3_homSap). Care still must be taken with published articles and genBank entries that may fail to specify the alpha actinin under consideration despite a 2003 article calling for adherence to HGNC international nomenclature standards followed here.
The comparative genomics situation is further confused by high sequence conservation within the ACTN gene family, by paralog loss in some clades, by possible independent duplication events, and by pre-duplication parental genes only in early deuterostomes. It is not easy to assign transcripts or genomic fragments to correct orthology class by methods such as best reciprocal Blast, especially when the query itself is a fragment (eg third spectrin domain of ACTN3). Many genBank entries are unlabeled, mislabeled, or ambiguously labeled as to correct ortholog family.
However a reliable actinin classifier can be built by requiring flanking gene synteny, diagnostic signature residues and indels in building the reference sequence seed collection that focus on signature regions in which ortholog classes differ significantly from each other. For example ACTN2/3 share a five-residue deletion in exon 19 relative to (ancestral-length) ACTN1/4.
The human ortholog of ACTN3 is unusual (but not unique) in having a fairly abundant null allele, R577x, meaning the arginine at position 577 of the 901 residue protein has been replaced by a stop codon. Worldwide 18% of the human population is homozygous 577x 577x. It is very unlikely that a functional truncated protein can be produced because even if the promoter region is still functional, the mRNA would be degraded by nonsense-mediated decay (with no possibility of selenocysteine substitution) and the stable quaternary dimer necessary for function cannot form (as explained below).
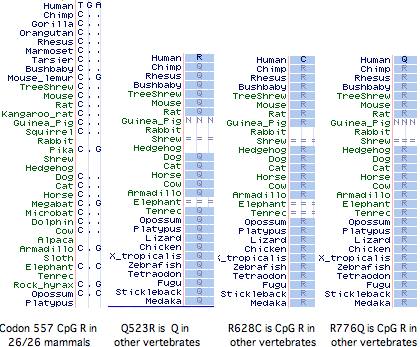
It has not been established whether the 577x was the initial inactivating mutation, as 3 additional amino acid changes (Q523R, R628C, R776Q) have also accrued at otherwise invariant sites in this allele (ie, in the dna donor to the public human genome relative to genBank reference sequence NM_001104). The latter two substitutions are also CpG mutational hotspots (the entire mRNA has 131 such sites). It is not known whether these other changes became widespread before or after R577x nor whether they affect ACTN3 function. However it is not easy to inactivate a large structural protein comprised of independent modules by single substitutions.
Curiously Q523R, R577x, and R628C all occur in the third spectrin repeat despite this region constituting only 11% of the gene, yet not a single non-synonymous base change has occurred in the 2706 bp coding region. With the advent of HapMap and similar projects, the phenotypic associations of these changes, possible co-occurrence with wildtype R577, and the date(s) of 577x founder mutations could be resolved.
All mammals with assembled genomes encode a CpG hotspot at codon 577. This has transitioned to TpG in the human 577x allele but is not a polymorphic site in any other known mammal, though the search has been restricted to the individual animals used in genome projects (since transcripts rarely extend this far), plus 36 unrelated baboons and 33 chimpanzees all genotyping to ‘wild-type’ 577R. Thus there is no support for 577x as balanced polymorphism in any mammal other than human even though Z-line skeletal muscle structures may be very similar.
In regards to the supposed evolutionary advantage of various allele combinations (proxied by sprinting or endurance sports prowess), humans remain slow and weak relative to other mammals regardless of their codon 577 status. The fastest human sprinter cannot outrun a dog with heartworms, much less a rabbit or chubby grizzly bear. A wild male chimp -- without any training or drug enhancement -- has the strength and aggression to rip apart the fittest human cage fighter.
Complete loss of ACTN3 does not give rise to a disease state or even observable phenotype in humans, despite a fallacious initial association with dystrophinopathy. Double knockouts of the orthologous gene in mouse are quite viable but exhibit various measurable effects. Over evolutionary time, the gene has been lost by natural genomic deletion in chicken and finch without known impact, yet retained in lizard, snake, and frog and even doubled in zebrafish (but not other ray-finned fish), again without known effect.
This suggested to some that human ACTN3 had an inessential or inconsequential physiological role to begin with (or become so since divergence with chimps), with its loss readily compensatable by other genes (presumably the 80% paralog ACTN3 with which it forms a heterodimer). In this view, ACTN3 may be on its way out the door, to disappear over time as a biallelic pseudogene. Given that over 80 other human genes have been lost, some very conserved over long evolutionary spans] since chimp divergence, it would not be surprising to catch a loss-in-progress.
The primary hurdle to clear here is the extraordinary conservation (proteome: 90th percentile) over 450 million years of ACTN3 amino acid sequence. It appears that this gene arose by segmental duplication from ancestral ACTN2 after the divergence of chondrichthyes (where all matches are ambiguous with respect to ACTN2/3). Gene duplicates are not usually retained over such a time frame unless they have a distinct functional niche that provides selective protection from constantly accruing deleterious mutations ('use it or lose it').
It is quite possible that earlier loss of a companion gene essential to the functional chain of ACTN3 has triggered its subsequent degeneration. The skeletal myosin gene MYH16 is potentially an attractive candidate, though actinins do not bind myosins directly but rather via actin. If ACTN3 functionality overlapped with ACTN2 apart from a critical role involving MYH16, then the loss of the latter gene would leave ACTN2 with no role that could not be compensated for by ACTN3. MYH16 was also lost to an internal stop codon after a half-billion years of conservation following its separation from other myosins.
Another explanation is balanced polymorphism ('somewhat less is more') along the lines of allele proportions maintained in sickle-cell hemoglobins (heterozygosity can be selectively advantageous in malarial resistance). The idea here that human groups are benefited if some individuals have exceptional speed or endurance.
For ACTN3, frequencies of the three possible diploid states (R577/R577, R577/577x, 577x/R577x) vary by ethnic group and supposedly correlate -- imperfectly but predictively -- with athletic prowess. While single-locus genetic determinism is preposterous as determinative of a complex and vaguely specified phenotype, this has nonetheless gained traction in the popular mindset.
Correlations per se have a poor track record in establishing causality -- for example, a low IQ might also correlate quite strongly with interest and participation in sports. Perhaps 577x -- ACTN3 is in fact expressed in brain -- just contributes to low IQ.
Phenotypic effects of ACTN1, ACTN2, and ACTN4 loss
Total absence of ACTN3, a well-conserved gene, does not result in genetic disease in either in human or double knockout mice . What about loss of its three paralogs?
ACTN4 is only member of the actinin gene family mapped to a human disease, focal segmental glomerulosclerosis, FSGS1. However no known mutation results in gene loss (which may be embryonic-lethal). Instead, all are autosomal dominant gain-of-function. Any of 6 distinct substitutions W59R, I149del, K255E, S262F, R310Q and V801 cause ACTN4 to bind too tightly (persistent switched-on mode) to the actin cytoskeleton in renal glomerular podocytes (visceral epithelial cells of kidneys involved in macromolecular filtration), resulting in a condition eventually progressing to renal failure. The mouse model for K255E is comparable.
Its tandem amino terminal calponin-homology CH domains crosslink actin filaments as regulated by Ca2+. These substitutions divert its usual localization away from actin stress fibers and focal adhesion points. As ACTN4 is also widely expressed in a variety of other cell types with diverse roles (motility, adhesion, endocytosis, tight junctions), this suggests pleiotropic affects beyond podocyte dysfunction.
ACTN4 forms a homodimer, meaning a compensation scenario as in ACTN2 replacement of ACTN3 is not feasible. The question is whether paralogous mutations in the calponin homology domains of other actinins have similar effects. That could be studied in knock-in mouse ACTN3 to help determine its normal functions. Sufficient conservation makes this feasible. The structure of the K255E protein shows that the calponin domains remain in compact configuration despite disruption of the CH2 bridge to W147 of CH1.
Less is known about disease alleles of ACTN2. A single report of dilated cardiomyopathy (CMD) seemingly attributes it to a heterozygous missense mutation at a conserved residue Q9R (which significantly precedes CH1). Again an autosomal dominant effect rather than loss, this mutation affected normal interaction with muscle LIM protein (MLP) and nuclear localization. The effect in ACTN3 of a mutation paralogous to Q9R is not known.
No disease allele of ACTN1 has ever been reported. This in itself is not surprising since only 1,500 human genes out of 20,000 (7.5%) have known disease alleles. However it could reflect essential functionality. ACTN1 does have a novel activity for an actinin, binding the metabotropic glutamate receptor type 5b, a GPCR protein rather than an ion channel, modulating receptor cell surface expression. An induced tyrosine phosphorylation mutation Y12F in pressure-induced adhesion response has been investigated. Despite oft-repeated claims to the contrary, ACTN1 is in fact expressed in skeletal muscle.
ACTN3 transcription is NOT restricted to skeletal muscle
In dozens of articles in low-impact factor journals and on at least one personal genomics advice page, it has been implied that ACTN3 is expressed exclusively in type 2 skeletal muscle fibers at the Z-line and so its absence or partial replacement must consequently relate to fast or slow twitch fiber performance. These copycat cites track back to a 1992 paper (see M86407 'skeletal muscle specific' 30-OCT-1994).
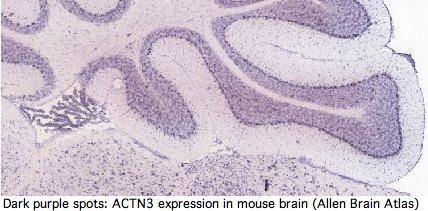
Yet GenBank and transcript-detecting chips contain numerous validated ACTN3 transcripts from other tissues, including heart muscle, pericardium, prostate, medulla oblongata, brain, salivary gland, skin, eye, hematopoietic stem cells, gastrointestinal tract, and testis. From its assembly, it can be seen that tree shrew genome contains a full length processed pseudogene -- which requires high expression of ACTN3 in germline tissue.
Actinins make cytoskeleton sense in all these cell types but none contain skeletal muscle fiber. It follows that ACTN3 has other functions unless the dominant modes of expression of this ancient conserved gene are gratuitous. Heart muscle obviously has a role in athletics but it is not of skeletal muscle type. The transcript database GeneHub-GEPIS shows heart with markedly higher expression of ACTN3 than skeletal muscle. Testis and hematopoietic cells too evidently play a role in athletic performance (why else would Tour de France cyclists load up on testosterone patches and erythropoietin shots?).
The interpretation of R577x has largely been based on the 'plausibility' of its expression in skeletal muscle explaining performance correlation. The argument largely vaporizes because ACTN3 is not expressed exclusively in skeletal muscle. It is just as easy to spin a tale of how a role in heart muscle or testes might account for the observations. Perhaps ACTN2 can compensate in skeletal muscle but not elsewhere. Thus the burden of proof requires demonstrating that ACTN3 roles in non-skeletal tissues do not have more explanatory power.
Thus ACTN3 is a good gene for illustrating the problems accompanying personal genomics testing. Some people will make significant life decisions based on test results regardless of disclaimers. If the accompanying explanation is shoddy -- and ACNT3 is commonly explained at the intellectual depth of an astrology column -- they cannot act in an informed way. At a bare minimum, ACTN3 needs a very detailed review of its published literature and comparative genomics. The widespread error in describing ACTN3 expression specificity is only the beginning of the serious omissions in this gene's existing annotation.
It is an immense public disservice to inform the public there is "a gene for college athletic scholarships" so their potentially elite 6-year old should be sequenced at R577 and channeled into proper coaching early on. In reality, hundreds of genes (not to mention environmental factors) are involved in vague phenotypes. A given gene may exhibit a statistically significant effect yet have very little predictive power.
ACTN3 transcription is NOT restricted to skeletal muscle: DN887432 Rabbit eye minus lens and cornea. AK313556 Homo sapiens cDNA pericardium DA827389 Homo sapiens cDNA clone PERIC2008976 pericardium BP318079 Homo sapiens cDNA Sugano cDNA library, pericardium AK125851 Homo sapiens cDNA testis AK303044 Homo sapiens cDNA clone TESTI4007778 testis GeneHub-GEPIS: human ACTN3 expressed in GI tract, heart, muscle, testis GeneHub-GEPIS: mouse ACTN3 expressed in bone, brain muscle, salivary gland, skin AK134757 Mus musculus adult male medulla oblongata cDNA, RIKEN full-length DT909761 Mus musculus cDNA Hematopoietic stem cells AF093775 Mus musculus alpha-actinin 3 (Actn3) mRNA, complete cds. BC111890 Mus musculus actinin alpha 3, mRNA (cDNA clone BC166600 Rattus norvegicus actinin alpha 3 mRNA Prostate
ACTN3-interacting gene products
Obviously a major actin organizing gene like the well-conserved and ancient gene ACTN3 cannot be totally lost from skeletal muscle Z-line and heart without in situ structural compensation by another protein. The lead candidate is ACTN2 because of similar expression and known heterodimer capacity, with ACTN1 and ACTN4 next in line. A more remote possibility is another protein similarly dimensioned as a dimeric actin-binding dumbbell, implying CH and spectrin domains. The human genome encodes 14 candidate genes based on spectrin domains but none of them has 4.
The compensating protein has another very stringent requirement, that it can approximately replicate ACTN3 interactions with myriad other structural and regulatory proteins without introducing inappropriate new ones. This implies two EF hand as the third and last of common chimeric protein domains.
Supposing that the compensatory protein is in fact ACTN2, its protein differs significantly from ACTN3 at xx residues out of 901. To the extent ACTN3 has co-evolved binding partners specifically adapted to its sequence, differences in efficacy may emerge upon substitution with ACTN2. These differences could in fact be responsible for the various sports correlations. In that case, polymorphisms in ACTN2 or these secondary genes associated linked with 577x could be major confounding issues (ie would need to be genotyped as well).
Other structural proteins known to bind at least one ACTN genes:
Protein Gene PubMed domain_bound actinin_bound fessilin 16450054 SP2 SP3 2 smooth muscle F-actin CH1 CH2 1,2,3,4 ...
(to be continued)
Phenotypic effects of ACTN1, ACTN2, and ACTN4 mutations
Total absence of ACTN3, a well-conserved gene, does not result in genetic disease in either in human or double knockout mice . What about loss of its three paralogs?
ACTN4 is only member of the actinin gene family mapped to a human disease, focal segmental glomerulosclerosis, FSGS1. However no known mutation results in gene loss (which may be embryonic-lethal). Instead, all are autosomal dominant gain-of-function. Any of 6 distinct substitutions W59R, I149del, K255E, S262F, R310Q and V801 cause ACTN4 to bind too tightly (persistent switched-on mode) to the actin cytoskeleton in renal glomerular podocytes (visceral epithelial cells of kidneys involved in macromolecular filtration), resulting in a condition eventually progressing to renal failure. The mouse model for K255E is comparable.
Its tandem amino terminal calponin-homology CH domains crosslink actin filaments as regulated by Ca2+. These substitutions divert its usual localization away from actin stress fibers and focal adhesion points. As ACTN4 is also widely expressed in a variety of other cell types with diverse roles (motility, adhesion, endocytosis, tight junctions), this suggests pleiotropic affects beyond podocyte dysfunction.
ACTN4 forms a homodimer, meaning a compensation scenario as in ACTN2 replacement of ACTN3 is not feasible. The question is whether paralogous mutations in the calponin homology domains of other actinins have similar effects. That could be studied in knock-in mouse ACTN3 to help determine its normal functions. Sufficient conservation makes this feasible. The structure of the K255E protein shows that the calponin domains remain in compact configuration despite disruption of the CH2 bridge to W147 of CH1.
Less is known about disease alleles of ACTN2. A single report of dilated cardiomyopathy (CMD) seemingly attributes it to a heterozygous missense mutation at a conserved residue Q9R (which significantly precedes CH1). Again an autosomal dominant effect rather than loss, this mutation affected normal interaction with muscle LIM protein (MLP) and nuclear localization.
The effect in ACTN3 of a mutation paralogous to Q9R cannot be asked because alignability to ACTN2 first begins at position 25 EYMEQE... ACTN1 and ACTN4 also lack a counterpart to this glutamine. However Q9 is strictly conserved in ACTN2 of mammals and amniotes along with the rest of exon 1, implying a functional basis.
No disease allele of ACTN1 has ever been reported. This in itself is not surprising since only 1,500 human genes out of 20,000 (7.5%) have known disease alleles. However it could reflect essential functionality. ACTN1 does have a novel activity for an actinin, binding the metabotropic glutamate receptor type 5b, a GPCR protein rather than an ion channel, modulating receptor cell surface expression. An induced tyrosine phosphorylation mutation Y12F in pressure-induced adhesion response has been investigated. Despite oft-repeated claims to the contrary, ACTN1 is in fact expressed in skeletal muscle.
In summary, gene loss in the 3 human paralogs of ACTN3 has never been observed. The known mutations are all autosomal dominant gain of function by stable in situ protein. This is consistent with essentiality of these genes.
K577R as ACTN3 phyloSNP
An alignment of exon 15 (which comprises about half of the third spectrin domain) shows that codon 577 was ancestrally K (lysine) in early vertebrates, a residue persisting without exception to the present day in all extant vertebrates diverging before platypus. In the mammalian stem, K was replaced by R (arginine) and that residue persisted in mammals (26/26 species). This is the definition of phyloSNP -- a clade-defining synapomorphy with persistent ancestral state whose conservation both before and after implies structural and/or functional significance in both the clade and its complementary tree but different roles. Evidently residue 577 has long been significant in the third spectrin domain of ACTN3, implying about any change in humans is disadvantageous.
The non-mammalian sequences below had to be individually established as orthologs of the human ACTN3 gene using retained flanking synteny of neighboring genes because the UCSC comparative genomics track misaligns paralogs here. The retained synteny is not always two-sided and in some cases inversions have resulted in loss of immediate adjacency.
R577x and co-evolution of actinin spectrin repeats
The four alpha actinin paralogs in human all have the same domain structure, namely 2 calponin-homology domains (that bind actin) followed by 4 spectrin rod domains followed by 2 EF hands that ancestrally bound calcium. These domains and their spacers comprise almost the entirety of the 901 amino acids of ACTN3. All three domain types are quite common in human proteins as various permutations, associations, and repeat numbers.
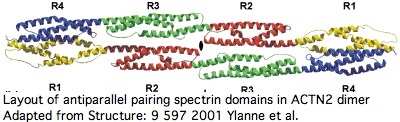
Actinins are a classic chimeric protein. The roles of each domain types are fairly well-known and excellent 3D domain structures are available; as an approximation, these domain folds independently so the global structure can be approximated. Actinins form anti-parallel dimers via their spectrin domains making the overall quaternary shape a dumbbell.
Dimerization (ie two actin binding sites) is obviously essential to the actin cross-linking functionality of actinins. ACTN3 and ACTN2 are known to form heterodimers in vivo. This suggests that ACTN2 could somewhat correct for reduced expression of ACTN3 in the R577x heterozygote and complete lack in 577x 577x.
Human genes encoding spectrin repeats: ACTN1 4 actinin alpha 1 ACTN2 4 actinin alpha 2 ACTN3 4 actinin alpha 3 ACTN4 4 actinin alpha 4 DMD 24 dystrophin DRP2 2 dystrophin related protein DST 29 dystonin isoform 1 PLEC1 7 plectin 1 isoform 6 SPTA1 22 spectrin alpha erythrocytic 1 SPTAN1 23 spectrin alpha non-erythrocytic 1 SPTB 17 spectrin beta isoform a SPTBN1 17 spectrin beta non-erythrocytic 1 SPTBN2 17 spectrin beta non-erythrocytic 2 SPTBN4 18 spectrin beta non-erythrocytic 4 SPTBN5 31 spectrin beta non-erythrocytic 5 SYNE1 31 spectrin repeat containing nuclear envelope 1 SYNE2 12 spectrin repeat containing nuclear envelope 2 UTRN 10 utrophin
In many domain-shuffled proteins (eg titin), the number of internal repeats is both evolutionarily unstable (due to the intrinsic propensity for mis-aligned recombination or replication slippage) and subject to domain-skipping alternative splicing, seemingly without drastic end-functional consequences. Such chimeric proteins are notoriously difficult to align homologically (ie by evolutionary descent) because quite different histories of contraction and expansion can result in same domain copy position.
Here it must be noted that splice boundaries in domain-shuffled proteins do not necessarily allow for neat gain or loss of extra domains by non-homologous recombination or exon skipping either because reading frame would not be preserved (leading to premature stop codons) or because fragmentary domains would be introduced.
The four human actinins have 21 coding exons but intron positions interrupt structural domains arbitrarily with incompatible reading frame phase. This suggests the ancestral actinin gene with completed internal expansions arose before the main era of intron gain, whereas expansions to vertebrate paralogs took place much later, yet not so recently that any flanking gene synteny nor any sequence alignability across spectrin domains could be retained.
When actinins from early eukaryotes are carefully considered (but not their intronation), it emerges that SP1 is likely the ancestral spectrin domain. Note one such domain still suffices for dimerization. An internal duplication prior to fungal divergence gave rise to SP4 and a protein with two spectrin repeats. SP2 and SP3 apparently arose from a second intragenic duplication, giving rise to the four spectin repeats seen in all extant bilateran actinins.
Vertebrate spectrins exhibit a remarkable rare genomic event most helpful in establishing the relative timing of events: exon 19 in all ACTN2 and ACTN3 orthologs is shorter by 5 amino acids than its counterpart in ACTN1 and ACTN4 (which have ancestral length). This loss did not arise from a conventional dna deletion but rather from development of an earlier internal splice donor of the same reading phase (the intron in all cases remains of 00 type). A shorter exon 19 is not a alternative splice option in extant vertebrates (no GT donor available).
Exons 19 and 20 compared across vertebrate actinins: ACTN1_homSap 0 DHSGTLGPEEFKACLISLGYDIGNDPQ 00 GEAEFARIMSIVDPNRLG... ACTN4_homSap 0 DHGGALGPEEFKACLISLGYDVENDRQ 00 GEAEFNRIMSLVDPNHSG... ACTN1_petMar 0 DHSGKLGAEEFKACLISLGYDVGNDQQ 00 GEAEFARIMTVVDPNNTG... ACTN1_braFlo 0 DGSGRLEPNEFKSCLISIGYNIQEGDK 00 ACTN2_homSap 0 RKNGLMDHEDFRACLISMGYDL 00 GEAEFARIMTLVDPNGQG... ACTN3_homSap 0 KRNGMMEPDDFRACLISMGYDL 00 GEVEFARIMTMVDPNAAG...
The loss of these residues did not have a profound structural impact as it only removed the linker separating the two EF hands. More subtly, the event may have caused loss of CA+2 sensitivity in the EF hands; indeed that may have been the basis for its fixation.
Because the lamprey genome and transcript collection as of December 2008 lack actinins of ACTN2/3 class but contain actinins clustering with ACTN1/4 (and not exhibiting the exon 19 loss) whereas the next node chondrichthyes possess actinins of both types, it follows (from parsimony) that ACTN2, which is more fundamental than ACTN3, arose post-lamprey from a segmental duplication preserving the 21 exons but then acquired the shortened exon 19 (perhaps as part of sub-functionalization).
Subsequent to that, ACTN3 arose as a segmental duplication of ACTN2 and so had the same exon 19 structure. The key point here is rare events do not occur twice (rare-squared). Precise internode timing awaits a better assembly of Callorhynchus milii genome because sequence fragments do not clearly distinguish lineage-specific gene family expansions from fundamental clade expansions. All four actinin types clearly arose prior to teleost fish appearance -- actinin innovation had ceased 300 million years prior to mammal emergence.
Thus the history of actinin gene duplication conflicts with 2R (two rounds of whole genome duplication), illustrating the folly of assigning 4-merous gene families to 2R without fully annotating them.
The number of repeats determine the length of spacer, total binding strength, and access to other binding proteins such as fessilin. The domain composition of ACTN3 is shown relative to its exons below, followed by a comparison of ACTN gene family chromosomal contexts.
>ACTN3_homSap Homo sapiens (human) 21 exons 901aa NM_001104 Q523R R577x R628C R776Q CH1 Ch2 SP1 SP2 SP3 SP4 EF1 EF2 0 MMMVMQPEGLGAGEGRFAGGGGGGEYMEQEEDWDRDLLLDPAWEKQQRK 0 0 TFTAWCNSHLRKAGTQIENIEEDFRNGLKLMLLLEVIS 1 2 GERLPRPDKGKMRFHKIANVNKALDFIASKGVKLVSIGAE 1 2 EIVDGNLKMTLGMIWTIILRFAIQDISVE 1 2 ETSAKEGLLLWCQRKTAPYRNVNVQNFHTS 2 1 WKDGLALCALIHRHRPDLIDYAKLR 0 0 KDDPIGNLNTAFEVAEKYLDIPKMLDAE 1 2 DIVNTPKPDEKAIMTYVSCFYHAFAGAEQ 0 0 AETAANRICKVLAVNQENEKLMEEYEKLASE 0 0 LLEWIRRTVPWLENRVGEPSMSAMQRKLEDFRDYRRLHKPPRIQEKCQLEINFNTLQTKLRLSHRPAFMPSEGKLVS 0 0 DIANAWRGLEQVEKGYEDWLLSEIRRLQRLQHLAEKFRQKASLHEAWTR 1 2 GKEEMLSQRDYDSALLQEVRALLRRHEAFESDLAAHQDRVEHIAALAQELN 2 1 ELDYHEAASVNSRCQAICDQWDNLGTLTQKRRDALE 0 0 RMEKLLETIDQLQLEFARRAAPFNNWLDGAVEDLQDVWLVHSVEETQ 0 0 SLLTAHDQFKATLPEADRERGAIMGIQGEIQKICQTYGLRPCSTNPYITLSPQDINTKWDM 0 0 VRKLVPSRDQTLQEELARQQVNERLRRQFAAQANAIGPWIQAKVE 0 0 EVGRLAAGLAGSLEEQMAGLRQQEQNIINYKTNIDRLEGDHQLLQESLVFDNKHTVYSME 0 0 HIRVGWEQLLTSIARTINEVENQVLTRDAKGLSQEQLNEFRASFNHFDR 0 0 KRNGMMEPDDFRACLISMGYDL 0 0 GEVEFARIMTMVDPNAAGVVTFQAFIDFMTRETAETDTTEQVVASFKILAGDK 0 0 NYITPEELRRELPAKQAEYCIRRMVPYKGSGAPAGALDYVAFSSALYGESDL* 0
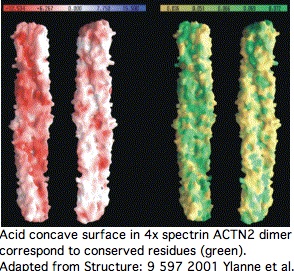
Proteins containing spectrin domains apparently all form dimers joined along the spectrin interface. This forces a two-fold symmetry axis per standard theories of protein oligomericity (fully supported by crystallographic studies), meaning anti-parallel complmentarity of numbered spectrin domains.
In other words, SP1 of one actinin monomer binds to SP4 of the other and SP2 to SP3 (ie 1234 to 4321). That has an additional twist for ACTN2 and ACTN3 in that they form a significant level of heterodimers in vivo (aligning 1234 to 4'3'2'1'). Similar considerations apply to all 18 human proteins containing spectrin domains.
This affects interpretation of R577x because it lies within SP3 of ACTN3 -- even if a transcript were still made that evaded nonsense-mediated decay, no stable dimer would be expected to form from non co-evolved SP1 and SP2.
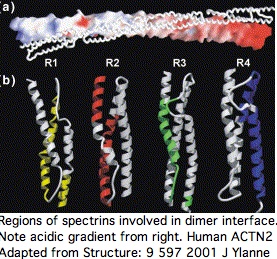
This structure induces important paired co-evolutionary constraints. For example, SP3 cannot fix a mutational change that might be acceptable in terms of its standalone fold unless this change is also compatible with its binding surface with SP2, yet a near-neutral slightly adverse change could be followed by compensatory change in SP2 (respectively SP2'), not the immediate homolog.
Sequence alignability however is primarily found across species in comparing orthologous and even paralogous (ie same-numbered) spectrin domains so the co-evolutionary effect is subtle. A single spectrin domain is itself a twisted 3-helix bundle and certain residues appear critical to that, notably an internal tryptophan at position 17 and penultimate leucine. The four spectrin domains are essentially unalignable to each other even co-registering to these invariant residues without recourse to 3D structural superpositioning. The indels giving rise to the slight length differences cannot be positioned. Their origin has not been dated.
>SP1 KLMEEYEKLASELLEWIRRTVPWLENRVGEPSMSAMQRKLEDFRDYRRLHKPPRIQEKCQLEINFNTLQTKLRLSHRPAFMPSEGKLVSIANAWRGLEQVEKGYEDWLLS >SP2 HLAEKFRQKASLHEAWTRGKEEMLSQRDYDSALLQEVRALLRRHEAFESDLAAHQDRVEHIAALAQELNELDYHEAASVNSRCQAICDQWDNLGTLTQKRRDALE >SP3 QLQLEFARRAAPFNNWLDGAVEDLQDVWLVHSVEETQLLTAHDQFKATLPEADRERGAIMGIQGEIQKICQTYGLRPCSTNPYITLSPQDINTKWDMVRKLVPSRDQTLQ >SP4 RLRRQFAAQANAIGPWIQAKVEEVGRLAAGLAGSLEEQMAGLRQQEQNIINYKTNIDRLEGDHQLLQESLVFDNKHTVYSMEHIRVGWEQLLTSIARTINEVENQVLT
To the extent ACTN2 and ACTN3 form both homo- and heterodimers in a phenotypically significant manner, there are both internal and external co-evolutionary constraints on each spectrin binding patch residue. The default evolutionary hypothesis of ACTN2 gene duplication creating ACTN3, raising gene dosage and introducing heterodimers initially of identical sequence. Over time, the gene sequences diverged with some retention of significantly different but still co-evolutionarily constrained heterodimers, accompanied by a degree of regulatory divergence in developmental timing and cellular location of expression.
At the domain scale, should a spectrin repeat be lost in one allele through deletion or alternative splicing, say the third, that creates the problem of aligning 124 to 4321 from the remaining wildtype allele (all actinins lie on autosomal genes) or more subtly 124 to 421 which mis-pairs SP2 to itself whereas it was evolutionary adapted to compliment SP3.
Thus it comes as no surprise to find all vertebrate actinins retain four spectrin subunits over tens of billions of years of branch time (see reference sequences). The apparent exception -- loss of SP4 in ACTN4 in human -- is due to annotation error at SwissProt and pipeline databases, the domain is in fact present though diverged from canonical sequence. It is better to use SuperFamily and Pfam here.
Actinin reference sequence collection and classifier tool
To understand the loss of ACTN3 in its phylogenetic context, it becomes essential to develop and maintain a large set of curated sequences for its entire gene family. The seed set of reliably assigned sequences can then be used as a classifying tool utilizing uBlast to recruit and properly label additional sequences and sequence fragments.
The final sampling density should be in proportion to the rate of evolutionary change of the feature under investigation. Thus many placental mammals might be valuable if the question is recurrence of 577x (and polymorphisms at other sites). For complete loss of gene, the time frame must include amniotes (ie chicken and finch). For gene duplication history (and subsequent sub- and neo-functionalization and their possible association with morphological or physiological change), tunicate, amphioxus, lamprey, and sharks are needed. For internal domain history, the collection must go back to fungi, and for intronation history still earlier. More ambitiously, the other 20 human proteins with spectrin domains could be included.
The sequence set below is only a start on an actinin classifier. A full-blown classifier (which is needed for every human gene family) requires a large-scale community effort. An excellent start has been made by Virel and Backman who also carefully determined spectrin repeat coordinates.
The sequences below are specialized to deuterostomes with genome projects with introns shown for human actinins.
>ACTN3_homSap Homo sapiens (human) 21 exons 901aa NM_001104 Q523R R577x R628C R776Q 0 MMMVMQPEGLGAGEGRFAGGGGGGEYMEQEEDWDRDLLLDPAWEKQQRK 0 0 TFTAWCNSHLRKAGTQIENIEEDFRNGLKLMLLLEVIS 1 2 GERLPRPDKGKMRFHKIANVNKALDFIASKGVKLVSIGAE 1 2 EIVDGNLKMTLGMIWTIILRFAIQDISVE 1 2 ETSAKEGLLLWCQRKTAPYRNVNVQNFHTS 2 1 WKDGLALCALIHRHRPDLIDYAKLR 0 0 KDDPIGNLNTAFEVAEKYLDIPKMLDAE 1 2 DIVNTPKPDEKAIMTYVSCFYHAFAGAEQ 0 0 AETAANRICKVLAVNQENEKLMEEYEKLASE 0 0 LLEWIRRTVPWLENRVGEPSMSAMQRKLEDFRDYRRLHKPPRIQEKCQLEINFNTLQTKLRLSHRPAFMPSEGKLVS 0 0 DIANAWRGLEQVEKGYEDWLLSEIRRLQRLQHLAEKFRQKASLHEAWTR 1 2 GKEEMLSQRDYDSALLQEVRALLRRHEAFESDLAAHQDRVEHIAALAQELN 2 1 ELDYHEAASVNSRCQAICDQWDNLGTLTQKRRDALE 0 0 RMEKLLETIDQLQLEFARRAAPFNNWLDGAVEDLQDVWLVHSVEETQ 0 0 SLLTAHDQFKATLPEADRERGAIMGIQGEIQKICQTYGLRPCSTNPYITLSPQDINTKWDM 0 0 VRKLVPSRDQTLQEELARQQVNERLRRQFAAQANAIGPWIQAKVE 0 0 EVGRLAAGLAGSLEEQMAGLRQQEQNIINYKTNIDRLEGDHQLLQESLVFDNKHTVYSME 0 0 HIRVGWEQLLTSIARTINEVENQVLTRDAKGLSQEQLNEFRASFNHFDR 0 0 KRNGMMEPDDFRACLISMGYDL 0 0 GEVEFARIMTMVDPNAAGVVTFQAFIDFMTRETAETDTTEQVVASFKILAGDK 0 0 NYITPEELRRELPAKQAEYCIRRMVPYKGSGAPAGALDYVAFSSALYGESDL* 0 >ACTN2_homSap Homo sapiens (human) length=894 0 MNQIEPGVQYNYVYDEDEYMIQEEEWDRDLLLDPAWEKQQRK 0 0 TFTAWCNSHLRKAGTQIENIEEDFRNGLKLMLLLEVIS 1 2 GERLPKPDRGKMRFHKIANVNKALDYIASKGVKLVSIGAE 1 2 EIVDGNVKMTLGMIWTIILRFAIQDISVE 1 2 ETSAKEGLLLWCQRKTAPYRNVNIQNFHTS 2 1 WKDGLGLCALIHRHRPDLIDYSKLN 0 0 KDDPIGNINLAMEIAEKHLDIPKMLDAE 1 2 DIVNTPKPDERAIMTYVSCFYHAFAGAEQ 0 0 AETAANRICKVLAVNQENERLMEEYERLASE 0 0 LLEWIRRTIPWLENRTPEKTMQAMQKKLEDFRDYRRKHKPPKVQEKCQLEINFNTLQTKLRISNRPAFMPSEGKMVS 0 0 DIAGAWQRLEQAEKGYEEWLLNEIRRLERLEHLAEKFRQKASTHETWAY 1 2 GKEQILLQKDYESASLTEVRALLRKHEAFESDLAAHQDRVEQIAAIAQELN 2 1 ELDYHDAVNVNDRCQKICDQWDRLGTLTQKRREALE 0 0 RMEKLLETIDQLHLEFAKRAAPFNNWMEGAMEDLQDMFIVHSIEEIQ 0 0 SLITAHEQFKATLPEADGERQSIMAIQNEVEKVIQSYNIRISSSNPYSTVTMDELRTKWDK 0 0 VKQLVPIRDQSLQEELARQHANERLRRQFAAQANAIGPWIQNKME 0 0 EIARSSIQITGALEDQMNQLKQYEHNIINYKNNIDKLEGDHQLIQEALVFDNKHTNYTME 0 0 HIRVGWELLLTTIARTINEVETQILTRDAKGITQEQMNEFRASFNHFDR 0 0 RKNGLMDHEDFRACLISMGYDLGEAEFARIMTLVDPNGQGTVTFQSFIDFMTRETADTDTAEQVIASFRILASDK 0 0 PYILAEELRRELPPDQAQYCIKRMPAYSGPGSVPGALDYAAFSSALYGESDL* 0 >ACTN1_homSap Homo sapiens (human) length=892 MDHYDSQQTNDYMQPEEDWDRDLLLDPAWEKQQRKTFTAWCNSHLRKAGTQIENIEEDFR DGLKLMLLLEVISGERLAKPERGKMRVHKISNVNKALDFIASKGVKLVSIGAEEIVDGNV KMTLGMIWTIILRFAIQDISVEETSAKEGLLLWCQRKTAPYKNVNIQNFHISWKDGLGFC ALIHRHRPELIDYGKLRKDDPLTNLNTAFDVAEKYLDIPKMLDAEDIVGTARPDEKAIMT YVSSFYHAFSGAQKAETAANRICKVLAVNQENEQLMEDYEKLASDLLEWIRRTIPWLENR VPENTMHAMQQKLEDFRDYRRLHKPPKVQEKCQLEINFNTLQTKLRLSNRPAFMPSEGRM VSDINNAWGCLEQVEKGYEEWLLNEIRRLERLDHLAEKFRQKASIHEAWTDGKEAMLRQK DYETATLSEIKALLKKHEAFESDLAAHQDRVEQIAAIAQELNELDYYDSPSVNARCQKIC DQWDNLGALTQKRREALERTEKLLETIDQLYLEYAKRAAPFNNWMEGAMEDLQDTFIVHT IEEIQGLTTAHEQFKATLPDADKERLAILGIHNEVSKIVQTYHVNMAGTNPYTTITPQEI NGKWDHVRQLVPRRDQALTEEHARQQHNERLRKQFGAQANVIGPWIQTKMEEIGRISIEM HGTLEDQLSHLRQYEKSIVNYKPKIDQLEGDHQLIQEALIFDNKHTNYTMEHIRVGWEQL LTTIARTINEVENQILTRDAKGISQEQMNEFRASFNHFDRDHSGTLGPEEFKACLISLGY DIGNDPQGEAEFARIMSIVDPNRLGVVTFQAFIDFMSRETADTDTADQVMASFKILAGDK NYITMDELRRELPPDQAEYCIARMAPYTGPDSVPGALDYMSFSTALYGESDL* >ACTN4_homSap Homo sapiens (human) length=911 MVDYHAANQSYQYGPSSAGNGAGGGGSMGDYMAQEDDWDRDLLLDPAWEKQQRKTFTAWC NSHLRKAGTQIENIDEDFRDGLKLMLLLEVISGERLPKPERGKMRVHKINNVNKALDFIA SKGVKLVSIGAEEIVDGNAKMTLGMIWTIILRFAIQDISVEETSAKEGLLLWCQRKTAPY KNVNVQNFHISWKDGLAFNALIHRHRPELIEYDKLRKDDPVTNLNNAFEVAEKYLDIPKM LDAEDIVNTARPDEKAIMTYVSSFYHAFSGAQKAETAANRICKVLAVNQENEHLMEDYEK LASDLLEWIRRTIPWLEDRVPQKTIQEMQQKLEDFRDYRRVHKPPKVQEKCQLEINFNTL QTKLRLSNRPAFMPSEGKMVSDINNGWQHLEQAEKGYEEWLLNEIRRLERLDHLAEKFRQ KASIHEAWTDGKEAMLKHRDYETATLSDIKALIRKHEAFESDLAAHQDRVEQIAAIAQEL NELDYYDSHNVNTRCQKICDQWDALGSLTHSRREALEKTEKQLEAIDQLHLEYAKRAAPF NNWMESAMEDLQDMFIVHTIEEIEGLISAHDQFKSTLPDADREREAILAIHKEAQRIAES NHIKLSGSNPYTTVTPQIINSKWEKVQQLVPKRDHALLEEQSKQQSNEHLRRQFASQANV VGPWIQTKMEEIGRISIEMNGTLEDQLSHLKQYERSIVDYKPNLDLLEQQHQLIQEALIF DNKHTNYTMEHIRVGWEQLLTTIARTINEVENQILTRDAKGISQEQMQEFRASFNHFDKD HGGALGPEEFKACLISLGYDVENDRQGEAEFNRIMSLVDPNHSGLVTFQAFIDFMSRETT DTDTADQVIASFKVLAGDKNFITAEELRRELPPDQAEYCIARMAPYQGPDAVPGALDYKS FSTALYGESDL* >ACTN2_galGal Gallus gallus MNSMNQIETNMQYTYNYEEDEYMTQEEEWDRDLLLDPAWEKQQR KTFTAWCNSHLRKAGTQIENIEEDFRNGLKLMLLLEVISGERLPKPDRGKMRFHKIAN VNKALDYIASKGVKLVSIGAEEIVDGNVKMTLGMIWTIILRFAIQDISVEETSAKEGL LLWCQRKTAPYRNVNIQNFHLSWKDGLGLCALIHRHRPDLIDYSKLNKDDPIGNINLA MEIAEKHLDIPKMLDAEDIVNTPKPDERAIMTYVSCFYHAFAGAEQAETAANRICKVL AVNQENERLMEEYERLASELLEWIRRTIPWLENRTPEKTMQAMQKKLEDFRDYRRKHK PPKVQEKCQLEINFNTLQTKLRISNRPAFMPSEGKMVSDIAGAWQRLEQAEKGYEEWL LNEIRRLERLEHLAEKFRQKASTHEQWAYGKEQILLQKDYESASLTEVRAMLRKHEAF ESDLAAHQDRVEQIAAIAQELNELDYHDAASVNDRCQKICDQWDSLGTLTQKRREALE RTEKLLETIDQLHLEFAKRAAPFNNWMEGAMEDLQDMFIVHSIEEIQSLISAHDQFKA TLPEADGERQAILSIQNEVEKVIQSYSMRISASNPYSTVTVEEIRTKWEKVKQLVPQR DQSLQEELARQHANERLRRQFAAQANVIGPWIQTKMEEIARSSIEMTGPLEDQMNQLK QYEQNIINYKHNIDKLEGDHQLIQEALVFDNKHTNYTMEHIRVGWELLLTTIARTINE VETQILTRDAKGITQEQMNDFRASFNHFDRRKNGLMDHDDFRACLISMGYDLGEAEFA RIMSLVDPNGQGTVTFQSFIDFMTRETADTDTAEQVIASFRILASDKPYILADELRRE LPPEQAQYCIKRMPQYTGPGSVPGALDYTSFSSALYGESDL* >ACTN1_galGal Gallus gallus MDHHYDPQQTNDYMQPEEDWDRDLLLDPAWEKQQRKTFTAWCNS HLRKAGTQIENIEEDFRDGLKLMLLLEVISGERLAKPERGKMRVHKISNVNKALDFIA SKGVKLVSIGAEEIVDGNVKMTLGMIWTIILRFAIQDISVEETSAKEGLLLWYQRKTA PYKNVNIQNFHISWKDGLGFCALIHRHRPELIDYGKLRKDDPLTNLNTAFDVAEKYLD IPKMLDAEDIVGTARPDEKAIMTYVSSFYHAFSGAQKAETAANRICKVLAVNQENEQL MEDYEKLASDLLEWIRRTIPWLENRAPENTMQAMQQKLEDFRDYRRLHKPPKVQEKCQ LEINFNTLQTKLRLSNRPAFMPSEGKMVSDINNAWGGLEQAEKGYEEWLLNEIRRLER LDHLAEKFRQKASIHESWTDGKEAMLQQKDYETATLSEIKALLKKHEAFESDLAAHQD RVEQIAAIAQELNELDYYDSPSVNARCQKICDQWDNLGALTQKRREALERTEKLLETI DQLYLEYAKRAAPFNNWMEGAMEDLQDTFIVHTIEEIQGLTTAHEQFKATLPDADKER QAILGIHNEVSKIVQTYHVNMAGTNPYTTITPQEINGKWEHVRQLVPRRDQALMEEHA RQQQNERLRKQFGAQANVIGPWIQTKMEEIGRISIEMHGTLEDQLNHLRQYEKSIVNY KPKIDQLEGDHQQIQEALIFDNKHTNYTMEHIRVGWEQLLTTIARTINEVENQILTRD AKGISQEQMNEFRASFNHFDRKKTGMMDCEDFRACLISMGYNMGEAEFARIMSIVDPN RMGVVTFQAFIDFMSRETADTDTADQVMASFKILAGDKNYITVDELRRELPPDQAEYC IARMAPYNGRDAVPGALDYMSFSTALYGESDL* >ACTN4_galGal Gallus gallus MVDYHSAGQPYPYGGNGPGPNGDYMAQEDDWDRDLLLDPAWEKQ QRKTFTAWCNSHLRKAGTQIENIDEDFRDGLKLMLLLEVISGERLPKPERGKMRVHKI NNVNKALDFIASKGVNVVSIGAEEIVDGNAKMTLGMIWTIILRFAIQDISVEETSAKE GLLLWCQRKTAPYKNVNVQNFHISWKDGLAFNALIHRHRPELIEYDKLRKDDPVTNLN NAFEVAEKYLDIPKMLDAEDIVNTARPDEKAIMTYVSSFYHAFSGAQKAETAANRICK VLAVNQENEHLMEDYEKLASDLLEWIRRTIPWLEDRSPQKTIQEMQQKLEDFRDYRRV HKPPKVQEKCQLEINFNTLQTKLRLSNRPAFMPSEGRMVSDINTGWQHLEQAEKGYEE WLLNEIRRLEPLDHLAEKFRQKASIHEAWTEGKEAMLKQKDYETATLSDIKALIRKHE AFESDLAAHQDRVEQIAAIAQELNELDYYDSPSVNARCQKICDQWDVLGSLTHSRREA LEKTEKQLETIDELHLEYAKRAAPFNNWMESAMEDLQDMFIVHTIEEIEGLIAAHDQF KATLPDADREREAILGIQREAQRIADLHSIKLSGNNPYTSVTPQVINSKWERVQQLVP TRDRALQDEQSRQQCNERLRRQFAGQANIVGPWMQTKMEEIGRISIEMHGTLEDQLQH LKHYEQSIVDYKPNLELLEHEHQLVEEALIFDNKHTNYTMEHIRVGWEQLLTTIARTI NEVENQILTRDAKGISQEQMQEFRASFNHFDKDHCGALGPEEFKACLISLGYDVENDR QGDAEFNRIMSLVDPNGSGSVTFQAFIDFMSRETTDTDTADQVIASFKVLAGDKNYIT AEELRRELPPEQAEYCIARMAPYRGPDAAPGALDYKSFSTALYGESDL* >ACTN3_echOce Echis ocellatus (snake) fragment not covering codon 577 EYMAQEEDWDRDLLLDPAWEKQQRKTFTAWCNSHLRKAGTQIENIEEDFRNGLKLMLLLE VISGERLPKPDKGKMRFHKIANVNKALDFIASKGVKLVSIGAEEIVDGNLKMTLGMIWTI ILRFAIQDISVEETSAKEGLLLWCQRKTAPYRNVNVQNFHISWKDGLALCALIHRHRPDL DYAKLRKDDPIGNLNTAFEVAEKYLDIPKMLDAEDIVNTPKP >ACTN3_anoCar Anolis carolinensis (lixard) 19 of 22 exons scaffold_1178:54057-80055 weak synteny ZDHHC24 MTTQIETHVQYNHSYMTTEDYMAQEEDWDRDLLLDPAWEKQQRK ETSAKEGLLLWCQRKTAPYRNVNVQNFHIR WKDGLALCALIHRHRPDLIDYAKLRK DDPIGNLNTAFEVAEKYLDIPKMLDAE DIVNTPKPDEKAIMTYVSCFYHAFAGAEQ AETAANRICKVLAVNQENEKLMQEYEKLASE LLEWIRRTIPWLENRVPEKTMSAMQRKLEDFRDYRRVHKPPRVQEKCQLEINFNTLQTKLRLSNRPAFMPSEGKMVS DIANAWKGLEQVEKGYEEWLLTEIRRLERLDHLAEKFKQKATLHENWTR GKEDMLTQKDYESASLYEIRALMRKHEAFESDLAAHQDRVEQIAAIAQELn ELDYHDAASVNSRCQAICDQWDTLGTLTQKRRDSLE RVEKLLETIDQLYLEFAKRAAPFNNWMDGAIEDLQDMFIVHSIEEIQ SLITAHEQFKATLPEADKERMAILGIQNEIQKIAQTYGIKLSGINPYTNLSHLDIANKWDT VKQLVPHRDQTLQEELARQQANERLRRQFAAQANIIGPWVQTKME EIGHISVDISGSLEDQMNHLKQHEQNIINYKSNIDKLEGDHQLIQEALVFDNKHTNYTME HIRVGWEQLLTTIARTINEIENQILTRDAKGISQEQMNEFRASFNHFDR KRNGMMDPDDFRACLISMGYD GEVEFARIMTLVDPNNTGVVTFQAFIDFMTRETAETDTAEQVMASFKILASDK SYITIEELRRELPPEQAEYCISRMTKYTGADAVSGALDYVSFSSALYGESDL* >ACTN3_xenTro NP_001135513 length=896 solid right synteny homSap MTTIETHIQYSNSYNMTEEDYMVQEEDWDRDLLLDPAWEKQQRKTFTAWC NSHLRKAGTQIENIEEDFRNGLKLMLLLEVISGERLPKPDKGKMRFHKIA NVNKALDFIASKGVKLVSIGAEEIVDGNLKMTLGMIWTIILRFAIQDISV EETSAKEGLLLWCQRKTAPYRNVNVQNFHISWKDGLALCALIHRHRPDLI DYSKLRKDDPIGNLNTAFDVAEKYLDIPKMLDAEDIVNTPKPDEKAIMTY VSCFYHAFAGAEQAETAANRICKVLAVNQENEKLMEEYEKLASELLEWIR RTIPWLENRTPEKNMGAMQKKLEDFRDYRRVHKPPRVQEKCQLEINFNTL QTKLRLSNRPAFMPSEGKMVSDIANAWKGLEQVEKGYEEWLLLEIRRLER LEHLAEKFKQKASLHESWTRGKEELLTQRDYESASLMEIRALVRKHEAFE SDLAAHQDRVEQIAAIAQELNELDYHDAASVNSRCQAICDQWDNLGTLTQ KRRDALERVEKLLETIDQLYLEFAKRAAPFNNWMDGAVEDLQDMFIVHSI EEIQSLITAHEQFKATLPEADKEKMSILGIQAEIQKIAQTYGIKLSGINP YTNLTHLDISNKWETVKQLVPHRDQTLQEELARQQANERLRRQFAAQANV IGPWIQSKMEEIGRISVDISASLEDQMNHLKQYEQNIISYKSNIDKLEGD HQLIQESLIFDNKHTNYSMEHIRVGWEQLLTTIARTINEVENQILTRDAK GISQEQMNEFRASFNHFDRKRNGMMDPDDFRACLISMGYDLGEVEFARIM TLVDPNNTGVVTFQAFIDFMTRETAETDTAEQVMASFKILASDKSYITIE ELRRELPPEQAEYCITRMTKYTGADAISGALDYMSFSSALYGESDL* >ACTN2_xenTro NP_001005053 length=894 solid bilateral synteny homSap MTQAVTNETYNYCYDEEEYMNQEEEWDRDMLLDPAWEKQQRKTFTAWCNS HLRKAGTMIENIEEDFRNGLKLMLLLEVISGERLPKPDRGKMRFHKIANV NKALDFIASKGVKLVSIGAEEIVDGNVKMTLGMIWTIILRFAIQDISVEE TSAKEGLLLWCQRKTAPYRNVNIQNFHTSWKDGLGLCALIHRHRPDLIDY SKLNKDDAVGNINLAMDVAEKYLDIPKMLDAEDIVSTAKPDERAIMTYVS CFYHAFAGAEQAETAANRICKVLAVNQENERMMEEYERLASELLEWIRRT IPWLENRTSEKTMQAMLKKLEDFRDYRRKHKPPRVQEKCQLEINFNTLQT KLRISNRPAFMPSEGKMVSDIANAWQRLEQGEKGYEEWLLTELRRLERVE HLAEKFRQKASLHESWTAGKEQLLLQKDYETASLTEVRALLRKHEAFESE LAARQDRVEQIAAIAQELNELDYYDAARINERCQKICDQWDRLGTLTQKR REALERTEKLLETVDQLHLEFIKRAGPFNIWMEGAMEDLQDMFIVHNVEE IQKLITAHEQFKATLPEADSERQAILGIQNEVEKVIQSYNIKISSHNPYS SITLDEIRTKWEKVKQLVPIRDQTLQEELARQQNNERLRRQFAGQANVIG PWIQKQMEEIGRSCIDFTGTLEDQMNSLKMLEHNIVNYKHNVDKLEGDHQ MIQESLIFDNKHTNYSMEHIRVGWELMLTTVARTINEVETQILTRDAKGI TQEQINVFRSSFNHFDKKKNGLMEHDDFRACLISMGYDLGEAEFARIMAL VDPSGIGTISFQSFIDFMTRETAETDTSEQVIAAFRILAADKPYILPEEL RRELPPEQAQYCLSKMPTYTGPGAVPGALDFTCFSSALYGESDL* >ACTN3a_danRer Danio rerio (zebrafish) NP_571597 partial synteny MTAVESQVQYGSYMMTATEEYMIQEDDWDRDLFLDPAWEKQQRKTFTAWC NSHLRKAGTQIENIEEDFRNGLKLMLLLEAISGERLPKPDKGKMRFHKIA NVNKALDFISSKGVKLVSIGAEEIVDGNLKMTLGMIWTIILRFAIQDISV EETSAKEGLLLWCQRKTAPYRNVNVQNFHISWKDGLALCALIHRHRPDLI DYSKLRKDDPIGNLNTAFDIAEKFLDIPKMLDADDIVNTPKPDEKAIMTY VSCFYHAFAGAEQAETAANRICKVLAVNQENEKLMEEYEKLASELLEWIQ RTIPWLENRVAEQTMHAMQQKLEDFRDYRRVHKPPKVQEKCQLEINFNTL QTKLRLSNRPAFMPSEGKMVSDIADAWKGLEQVEKGYEEWLLTEIRRLER LDHLAEKFRQKSTLHQSWTTGKEELLSQKDYETASLMEIRALMRKHEAFE SDLAAHQDRVEQIAAIAQELNELDFYDAATINAQCQGICDQWDNLGTLTQ KRRESLERVEKLWETIDQLYLEFAKRAAPFNNWMDGAMEDLQDMFIVHSI EEIQSLITAHDQFKATLPEADKERMAIMGIHSEVLKIAQTYGIKIVSENP YTVLSHQNIVNKWEAVKQLVPMRDQMLQEEVARQQSNERLRRQFAAQANI IGPWIQTKMEEISHVSIDIAGSLEEQMSNLKTYEQNIINYKSNIDKLEGD HQLSQEGLIFDNKHTNYTMEHVRVGWEQLLTTIARTINEVENQILTRDAK GISQEQLNEFRASFNHFDRKRNGMMDPDDFRACLISMGYDLGEVEFARIM TLVDPNNTGVVTFQAFIDFMTRETAETDTAEQVMASFKILASDKAYITVE ELRRELPPEQAEYCISRMTKYMGGDAPPGALDYISFSSALYGESDL* >ACTN3b_danRer Danio rerio (zebrafish) NP_991107 length=890 partial synteny MQYNNTYMMSTTQDYMIQEDDWDRDLLLDPAWEKQQRKTFTAWCNSHLRK AGTQIENIEEDFRNGLKLMLLLEVISGERLPKPDKGKMRFHKIANVNKAL DFICSKGVKLVSIGAEEIVDGNVKMTLGMIWTIILRFAIQDISVEETSAK EGLLLWCQRKTAPYRNVNVQNFHISWKDGLALCALIHRHRPDLIDYSKLR KDDPIGNLNTAFEVAEKYLDIPKMLDAEDIVNTPKPDEKAIMTYVSCFYH AFAGAEQAETAANRICKVLAVNQENERLMEEYEKLASELLEWIRRTIPWL ENRAAEQTMRAMQQKLEDFRDYRRVHKPPRVQEKCQLEINFNTLQTKLRL SNRPAFMPSEGKMVSDIANAWKGLEQVEKGYEEWLLTEIRRLERLDHLAE KFKQKCSLHESWTTGKEHLLSQKDYETASLMEIRALMRKHEAFESDLAAH QDRVEQIAAIAQELNELDYHDAATVNARCQGICDQWDNLGTLTQKRRDSL ERVEKLWETIDQLYLEFAKRAAPFNNWMDGAIEDLQDMFIVHSIEEIQSL ITAHDQFKATLPEADKERMAVMGIQNEIVKIAQTYGIKLVGVNPYSVLSP QDITNKWEAVKQLVPLRDQMLQEEVARQQANERLRRSFAAQANIIGPWIQ TKMEEIGHVSVDIAGSLEEQMNNLKQYEQNIINYKSNIDKLEGDHQLIQE ALIFDNKHTNYTMEHIRVGWEQLLTTIARTINEVENQILTRDAKGISQEQ LNEFRASFNHFDRKRNGMMDPDDFRACLISMGYDLGEVEFARIMTLVDPN NTGVVTFQAFIDFMTRETAETDTAEQVMASFKILASDKPYITVEELRREL PPEQAEYCISRMTKYVGPEGALGALDYISFSSALYGESDL* >ACTN1_leuEri Leucoraja erinacea (ray) est ambiguous fragment TFTAWCNSHLRKADTQIESIEEDFRDGLKLMLLLEVISGERLAKPERGKMRVHKISNVNK ALDFIAGKGVKLVSIGAEEIVDGNAKMTLGMIWTIILRFAIQDISVEQTSAKEGLLLWCQ RKTAPYKNVNIQNFHISWKDGLGFCALIHRHRPELIDYAKLRKDDPYTNLNTAFDVAEKY LDIPKMLDAEDIVGTARPDEKAIMTYVSSFYHAFSGAQ >ACTN23_squAca Squalus acanthias (shark) est ambiguous fragment EYMLQEEDWDRDLLLDPAWEKQQRKTFTAWCNSHLRKAGTQIENIEDDFRNGLKLMLLLEVISGERLPKPDQGKMRFHKIANVNKALDYISRKGVKLVSI GAEEIVDGNVKMTLGMIWTIILRFAIQDISVEETSAKEGLLLWCQRKTAPYKNVNVQNFHTSWKDGL >ACTN23_calMil Callorhynchus milii (elephant_shark) assembly ambiguous fragment GERLPKPDKGKMRFHKIANVNKALDYIASKGVKLVSIGAEEIVDGNVKMTLGMIWTIILRFAIQDISVEETSAKEGLLLWCQRKTAPYKNVTVQNFHT