Vertebrate Gene Origins
The Origin of Vertebrate Genes
Introduction to gene families
Contemporary vertebrates genomes contain about 20,000 coding genes, not many more than pre-Bilatera and far fewer than was believed prior to the genomic era. These genes fall into perhaps 1500 gene families each with its own history of expansion and contraction, so sizes today vary from singletons and doubletons to families such as GPCR with over a thousand members. We wish to track each of these gene histories back to the last ancestral genome in which a parental gene was present without further homologs.
No single sweeping explanation of vertebrate gene family histories is possible because the numerous gene duplication mechanisms have had varying impacts on different gene families. However histories can be shared to a degree when unrelated gene family members are duplicated or deleted in a contiguous block. Shared history can also be less than full-length gene in the case of mobile domains, internal tandem expansions and multi-domain chimeric proteins, complexities not considered further here.
Because converging homology of sequences and folds is immensely implausible, gene families with multiple members clearly experienced a history of net gene duplication (gains outpaced losses). However singleton genes too may have a significant duplication history when gene loss of paralogs is unmasked. And for completeness, we should also consider ghost proteins, present today only as pseudogene debris or entirely gone and inferred from continuing presence in extant species descended from ancestral nodes (if it can be established these are not lineage-specific gene family expansions).
Gene loss processes are quite significant, with human a net loser in gene count over the last 310 myr, contrary to cultural narratives assuming 'progress' towards 'complexity' as a given. It's well-known that human sensory genes (olfaction and vision) have been decimated but the list hardly stops there. Once the full gene complement has been established at each ancestral node, the timing of each human gene loss can be established. This may amount to several thousand genes relative to the amniote ancestor.
Definition of gene family
The specification of gene family is quite subtle because any practical operational definition is limited by the sensitivity and potential arbitrariness of phylogenetic depth for coalescence cut-off. This is illustrated by a simple gene family, the sulfatases. The 17 paralogs in human, seemingly the final count because of completeness of that genome and definitive probes, cluster quite distinctly within all-vs-all blastp searches and are readily organized by sequence and exon pattern conservation into a conventional gene tree whose duplications can be timed relative to various ancestral divergences of the metazoan lineage relative to human.
Yet when 3D structures at PDB are consulted, it becomes undeniable that sulfatases and alkaline phosphatases are structurally homologous over their entire length despite a complete lack of convincing primary sequence alignability even by profile. Cases of very ancient fold convergence are known (eg TIM beta barrels) but here the fold is so complex, unconventional and isolated in fold space that coincidental arrival at a common structure can be dismissed. That conclusion is independently supported by colocalization of active sites and similarity of catalytic mechanism (hydrolysis of ester) despite lack of post-translational modification to formylglycine in any phosphatase.
Consequently, the 4 phosphatase paralogs (comprised of a tandem triple on chr2 and an isolated gene on chr1) bring this human gene family size to 21 if we allow gene convergence back to common ancestor of human with bacteria. Setting the cut off at Urmetazoan or at 30% identity blast cutoff would keep these gene families separate (and even slightly partition the sulfatase group). Future methods could conceivably justify coalescence with ever more subtly related gene families. Once a genome is sequenced, blastp criteria for gene family inclusion becomes available across it; however tertiary structure determination of entire proteomes is not yet in sight, meaning that fold definitions can only be applied unevenly.
The small gene family consisting of PRNP and PRND is also instructive. Although in parallel tandem position and clearly homologous by hand-curational criteria despite deletions of the 40 residue repeat in the latter, the relatively short length of 255aa, compositional simplicity N-terminally, and percent blastp identity in the mid-20's invariably cause pipeline clustering procedures to miss the family association.
This gene family also illustrates the folly of attempting to date duplications by their degree of divergence -- it's not particularly ancient (tetrapod) despite the loss of sequence similarity. PRND can only be traced back by all-out manual curation; it is cleanly deleted in a gapless chicken contig but fortunately a second node representative (lizard) retains it. The high GC content of non-synonymous codon positions leads to high CpG rates in synonymous 4x positions as well through the hotspot effect. Compare this to a histone that differs at one residue between human and garden pea; here synonymous 4x position change has been saturated for 500 myr.
SUMF1 and SUMF2 have sequences closer than many duplicates originating in mammalian time yet their coalescence predates divergence of human with algae. The spotty retention of SUMF2 -- gene loss is often homoplasic -- illustrates that accurate gene family histories require dense phylogenetic sampling.
In other words, the plot of percent identity versus phylogenetic depth flattens so quickly that a given gene duplication cannot be dated at all reliably from its divergence -- neither quantitatively (absolute date) nor qualitatively (relative date with respect to speciation nodes). Gene pairs such as PRNP/PRND evolve at very different individual rates and those rates are quite non-uniform having different histories of accelerating and slowing. Consequently it is preposterous to assign events to supposed 1R and 2R whole genome duplications based on current divergence.
PRNP tracks back to teleost fish but again only using tools such as syntenic flanking genes and secondary structure can establish this because fish PRNP protein experienced an astonishing internal expansion amino-terminally. The gene didn't necessarily originate in teleosts but to date it has proven impossible to find it chondricthyhes or earlier diverging species because conserved residues have been swamped out by the expansion and assembled genomes are necessary to narrow down the search region.
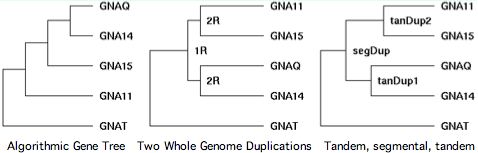
Chromosomal adjacency and broader synteny are thus important in accurately determining the gene tree. Sequence algorithms alone often get it wrong. That's seen quite dramatically in GPCR transducins, in which 10 genes occur in tandem pairs. Clearly these didn't come together by random rearrangement. Tandem pairs often separate rapidly in sequence because they must take on distinct roles to be supported by selection, yet this tendency can be offset by gene conversion homogenization.
The transducins also illustrate the fallacy of parsimony. Gene families are the result of a unique history, not statistics. That history is often but not always the most parsimonious -- Occam's razor cuts both ways. The third panel shows the correct history.
Gene family histories can also be exceedingly simple. Homogentisate 1,2-dioxygenase, HGD, is as simple as they get. It catalyzes an intermediate step in aromatic amino acid catabolism; without it, a toxic metabolite accumulates. This gene has not been lost from any eukaryotic lineage nor fixed a duplicate or processed pseudogene in 50 billion years of branch length. Sequence conservation is high; there is never ambiguity with blast searches.
Yet HGD illustrates another pitfall in defining gene families. The most recent human genome assembly (NCBI Build 36.1) has a left-over 500 kbp piece of DNA that did not fit into the regular assembly (for unknown reasons). This piece on chr3_random:18,988-73,308 would give rise to an artifactual paralog of HGD with automated procedures, in effect doubling gene family size. Such problems are rare in near-complete genomes such as human but all too common in lower coverage assemblies.
In summary, the non-chimeric proteome of a completed genome could be partitioned successfully today into gene families. However the resulting partition would be subject to subsequent lumping as more fold families become known and through manual curation. The former process could continue for decades because the fold class of many thousands of vertebrate proteins is still unknown -- indeed some 10,000 human proteins at SwissProt have never been the subject of any publication. Determining gene family tree topologies and dating duplications relative to speciations will require completion of 2x-6x genomes and more intensive phylogenetic sampling to understand lineage-specific gain and loss.
Bad gene names complicate the task
Six sources of gene terminology error makes these histories harder to explain:
- The term 'gene copy' is sometimes used for what are really deeply diverged paralogs with dimly related functions; gene duplication processes can make highly inexact and inequivalent copies right at the event (so they are never copies), subsequent evolutionary trajectories (other than gene conversion) only dilute the comparison further.
- The Greek prefix 'iso' is used consistently throughout science to mean 'same'; hence 'isoform' is highly inappropriate for products of distinct diverged genes or alternative splicings of a single gene (the vast majority of which are transcriptional artifacts to begin with). This usage perpetuates long-gone days of starch gel electrophoresis, an experimental technique that could only distinguish enzymatic forms if surface charge happened to differ.
- Two genes in two species are orthologous only if both genes are descended from a single gene in their last common ancestor. Gene function is completely irrelevant to the definition of orthology. Orthology cannot be defined (even in complete genomes) by reciprocal best-blast, a mere methodological option whose convenience is offset by many erroneous pairings. Orthology can be very difficult to establish even by hand-curation.
- Homologous genes within a given species are paralogs. Gene trees rapidly become very complicated even for moderately sized gene families and small numbers of species. Newly introduced terms such as in-paralogs and out-paralogs are utterly inadequate to describe this complexity. It is better to provide separate graphical species trees and gene trees coupled by the same time scale.
- The lack of awareness or refusal of authors to comply with agreed-upon international conventions for gene nomenclature (HUGO) causes many downstream problems. Gene names in the genomic era need to stay clear of subscripts, superscripts, mixed Roman and Greek letters, single or double letters, unreadable italics, lower case denoting mouse, easily confused i's and 1's (or o's and 0's), lab jokes, contrived acronyms, and naming by soon-forgotten experiment, tissue expression, end phenotype, or common disease symptom.
- Doubled nomenclature is a very poor idea: proteins should be named after their genes, not given separate unrelated acronyms. These practice hits bottom in hemoglobin terminology which uses greek letters, superscripts, accent marks, unrelated symbols for ancestral genes etc, all completely independent of gene names.
It's all been tried before and it doesn't work. There's a reason why Linnaean genus-species nomenclature took over for the species tree -- it's anchored in evolution. Gene trees too can have evolution-based nomenclature, say 3-4 letters to denote gene family, numbered paralogs after that (hopefully by sequence relatedness), and suffixed with a species code adequate for comparative genomics (as in SUMF2_homSap).
This creates a unique hierarchical identifier that narrows Google, PubMed, and GenBank searches. It clusters members of gene families by name and extends to the coming many-genome era. With complete genomes, we are in a position to create stable nomenclature, imperfect perhaps but much improved. Gene terminology need not be immutable (Linnaean taxonomy certainly is not), we'd settle for a good approximation.
Browser centers can provide global update cycles for gene names and maintain synonymy lookup tables. Probably nine vertebrate gene families in ten could be named forever stably today. Official gene sets now at UCSC, NCBI and Ensembl set gene names as random unmemorable alphanumeric strings such as uc003emt.1 (that serve as indexing fields in immense relational databases). We see from opsins that strings for a gene family are entirely unrelated: uc003emt.1 uc003vnt.2 uc004fjz.2 uc001hza.1 uc003hzv.1.
Since the geneSorter resource at UCSC already carries the all-vs-all blastp data needed for a quick gene tree as well as the requisite associated column for HUGO name, the uc and version number could be moved to the rear (.uc1, .uc2, ...) and the 6 free characters be used for a gene tree-driven nomenclature that incorporates HUGO names to the extent these are available and unambiguous.
The nomenclatural problem has significant practical impacts in synteny. Relationships that should jump out from name along are instead obscured. For example, flawed gene names mask the extent of human paralogons on chr15 and chr9 -- gene names of close human paralogs often bear no resemblance (eg PRUNE and ATCAY). Comparative genomics in other mammals brings in still other names (more commonly no name) and uninformative transcript or RefSeq strings. It's not a sustainable practice.
Vertebrate gene numbers have not increased over time
The number of coding human genes is still not known (Oct 2008) to within 1500-gene accuracy even seven years after sequencing. In fact, no serious effort to enumerate human genes is even underway for lack of funding and unwillingness to curate. Official compilations miss ordinary enzymes like COMT2 (transcribed only in special cells of inner ear) yet still include gravely decayed pseudogenes (that still have spliced transcripts) and non-genes (supported by transcript noise but no comparative genomics even to chimp). Beyond that, weak gene models may guess at initial methionines, lack stop codons and propose impossible alternate splices missing structurally essential core exons, active sites and any phylogenetic support.
Despite these limitations, it's worth compiling gene counts from original articles describing new genome assemblies and as subsequently amended. Being gene counts for contemporary species, they don't directly provide ancestral gene counts at divergence nodes. Yet since no trend whatsoever relates depth of divergence or simplicity of body plan with diminished gene count, ancestral counts may not be so different.
The scientific literature contains some incredulous assertions in the scientific literature about gene counts. If salmon truly experienced four rounds of whole genome duplications since cephalochordate, that implies (neglecting gene loss between rounds and gene gain in Branchiostoma) 350,400 coding genes. Yet the immense collection of salmon cDNA shows no indication whatsoever of any such growth in gene family size. On the contrary, salmon gene count seems very similar to every other vertebrate, as do all of the supposed 3R teleost fish. Their genomes show roughly 20,000 genes, nowhere near the 175,200 expected. How could mature gene-finding algorithms miss 155,200 coding genes given strong known paralogs?
Large-scale gene loss between rounds of duplication reduces the paradox somewhat, yet duplication followed shortly by loss of half the genes seem largely irrelevant to contemporary gene histories, like counting trees falling in the forest that got back on the stump.
Fish genomes have an astronomical numbers of gaps as of October 2008 assemblies. Unbridged gaps in particular imply a great many chromosomal misplacements. It is exceedingly premature to consider global syntenic relationships in a situation with 23,322 unbridged gaps because no known method can distinguish whole genome duplication from the steady accrual of ordinary segmental and re-translocated tandem duplications.
To the contrary, genome sequencing projects show if anything a decreasing trend in gene count in more recently emerging deuterostomes. For example, sea urchin genome has 23,300 coding genes whereas humans have but 20,176 in the 11 Sept 2008 tally of consensus CDS and even this number seems to be inflated over the 17,052 distinct locus count by assignment of multiple CCDS IDs numbers to single genes.
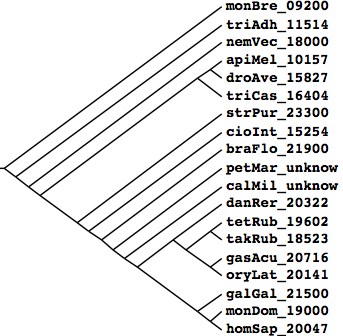
Tree generated at Phylodendron by the following Newick string from peer-reviewed literature compilation: (monBre_09200,(triAdh_11514,(nemVec_18000,(((apiMel_10157,droAve_15827),triCas_16404), (strPur_23300,(cioInt_15254,(braFlo_21900,(petMar_unknow,(calMil_unknow,((danRer_20322,((tetRub_19602,takRub_18523), (gasAcu_20716,oryLat_20141))),(galGal_21500,(monDom_19000,homSap_20047)))))))))))); Species Assembly Gaps Unbridged #Genes Organism homSap Mar 06 387 373 20,047 Homo sapiens (human) monDom Jan 06 72,803 5,336 19,000 Monodelphis domesticus (opossum) galGal May 06 78,478 17,335 21,500 Gallus gallus (chicken) anoCar Feb 07 43,237 ND ND Anolis carolinensis danRer Jul 07 49,727 11,857 20,322 Danio rerio (zebrafish) tetNig Feb 04 25,763 23,322 19,602 Tetraodon nigroviridis (pufferfish) takRub Oct 04 7,213 7,213 18,523 Takifugu rubripes (fugu) gasAcu Feb 06 16,945 1,913 20,716 Gasterosteus aculeatus (stickleback) oryLat Apr 06 134,426 8,131 20,141 Oryzias latipes (medaka) calMil Dec 06 ND ND ND Callorhinchus milii (eshark) petMar Mar 07 202,409 ND unpub Petromyzon marinus (lamprey) braFlo Mar 06 94,815 3,031 21,900 Branchiostoma floridae (amphioxus) cioInt Mar 05 22,521 ND 15,254 Ciona intestinals (tunicate)* strPur Sep 06 80,391 ND 23,300 Strongylocentrotus purpuratus (urchin) monBre_09,200 Monosiga brevicollis triAdh_11,514 Trichoplax adhaerens nemVec_18,000 Nematostella vectensis apiMel_10,157 Apis melifera triCas_16,404 Tribolium castenatum droAve_15,827 Drosophila 12_species strPur_23,300 Strongylocentrotus purpuratus cioInt_15,254 Ciona intestinalis* braFlo_21,900 Branchiostoma floridae petMar_unknow Petromyzon marinus calMil_unknow Callorhinchus milii danRer_20,322 Danio rerio tetRub_19,602 Tetraodon nigroviridis takRub_18,523 Takifugu rubripes gasAcu_20,716 Gasterosteus aculeatus oryLat_20,141 Oryzias latipes galGal_21,500 Gallus gallus monDom_19,000 Monodelphis domestica homSap_20,047 Homo sapiens * recent major update
Gene dosage issues
Gene duplication has the initial effect of increasing gene dosage in mammals. That might seem fairly innocuous in view of various homeostatic feedback mechanisms that might adjust expression and activity to whatever level is optimal. As duplicates are assimilated over time (sub- and neo-functionalization) the extra copies are then important sources of adaptive innovation and complexification.
However gene dosage diseases (eg Patau and Down syndromes) emerge just from partial trisomy of small chromosomes, suggesting feedback mechanisms cannot always compensate for increased gene dosage. (Note many karyotypic abnormalities in humans have no phenotypic sequalae.) The evolution of sex chromosomes in theran mammals evidently raised gene dosage issues because we observe extensive dose compensation mechanisms fixed for chrX genes such as processed retrogenes on autosomals and random X inactivation.
Although the mammalian genome is peppered with small and large paralogons (segmental duplications of multiple coding genes) of all age classes, there may still be strong constraints on paralogon possibilities. We may only see the ones that offered both immediate and long-term selective advantages not offset by gene dosage ills. Many older paralogons are today a mix of retained duplicates and singletons where a gene has dropped out from one paralogon. The best explanation of this phenomenon (pre-adaptation arising from multi-functionality) has been supplied by Piatigorsky.
Whole genome duplication is potentially more benign vis-a-vis gene dosage that other modes of gene duplication because the entire scattered genomic regulatory apparatus is also duplicated, possibly facilitating homeostasis. Yet experimentally induced tetraploidy in mouse is invariably lethal in early embryonic development. Reports of tetraploidy in other rodents are demonstrably false (see below).
If one believes internet twitter, two rounds of whole genome duplication occurred in the vertebrate lineage. Here it is worth noting the highly preliminary genome assemblies of lamprey and elephantshark, the critical two nodes beyond cephalochordate and urochordate. These assemblies completely omit many expected genes in their trace archive coverage; assembled contigs are typically of sub-gene length and contain a small fraction of expected exons. It is very difficult to assembly whole genes from several independent contigs since this could easily chimerize paralogs. As the next node, teleost fish, is greatly confused by large-scale paralogons, the entire discussion seems premature.
We have to wonder how non-tetraploid Branchiostoma could have more coding genes that supposedly octaploid fish. It could equally be argued that whole genome duplication is a mechanism to reduce gene count.
Large scale duplications (a lot of gene dosage coming on board suddenly) is very difficult to disentangle from gradual incremental gene family gain and loss arising from segmental and tandem duplication, given the extent of scrambling from inversions, fusions, and fissions. For example, humans have 6 pericentric inversions and a chromosomal fusion just relative to chimp. A lot of macro-variation is observed just one human to the other.
Small scale duplicon-driven inversions and duplications, often at homoplasic hotspots, go into the evolutionary hopper at nearly every childbirth. This greatly raises the odds that some will be fixed as near-neutral. Over vast evolutionary times, this could produce the many paleo-paralogons we observe today in the mammalian genome rather than paleo-polyploidy.
Tetraploid mammal faces oblivion
Polyploidy is quite important for understanding the evolutionary potential of the human genome. It has long been believed that polyploidy played less of a role in animal evolution than plants, the explanation (White, 1973) being chromosomal sex determination and obligatory cross-fertilization in addition to gene dosage issues. Bisexual polyploidy seemed to have stopped with the 12 known species of polyploid amphibians, with no polyploid amniote (much less mammal) ever documented.
If so, whole genome duplication has played no role in mammalian evolution over the last 400 million years and is not even an available mechanism for gene family expansion for in this lineage -- whole genome duplication is at best a paleo-process.
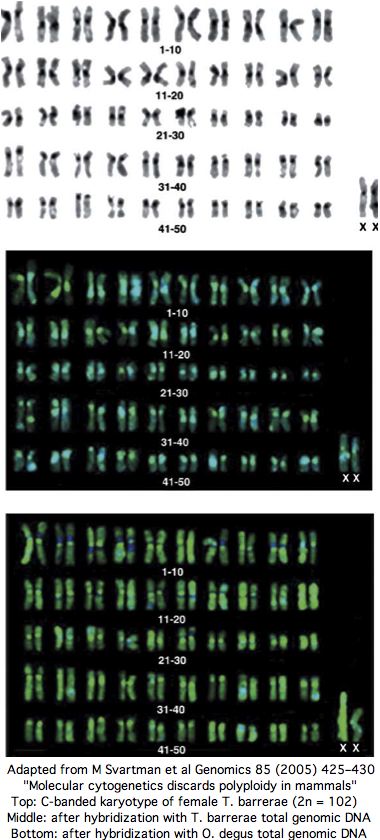
Consequently it came as a great surprise when Nature published a paper in 1999 asserting tetraploidy in a hystricognath rodent, Tympanoctomys barrerae. Four followup papers from the same group appeared over the next ten years in J Evol Biol, Biol Research and Genomics, culminating in an August 2008 paper in Mammalian Genome reporting tetraploidy in a second genus Pipanacoctomys.
Midway, a thorough experimental paper from karyotyping experts appeared in Genomics in 2005 entitled " Molecular cytogenetics discards polyploidy in mammals". As that journal never releases its articles online, the expense of reprints diminished the article's impact. Perhaps it contained some technical error since the tetraploidy authors and journal peer-reviewers seemed to ignore it subsequently. According to downstream google search, mammalian tetraploidy is a done deal.
Yet the senior author of the 2005 paper has published 86 previous papers over 25 years in mammalian karyotypic phylogenomic analysis, so can be assumed familiar with best contemporary experimental methods and their potential pitfalls. Contacted by email in Sept 2008, the authors confirm that nothing has changed their view that Tympanoctomys barrerae is an ordinary diploid with extensive repeats that simply don't respond to C-banding (nothing new there).
The experimental evidence for T. barrerae diploidy is overwhelming. No form of tetraploidy -- autotetraploidy or allotetraploidy (two-species hybrid) -- is consistent with the data. Every chromosome has two copies: that is diploidy. The experimental approaches includes painting by separated chromosomes of closely related diploid Octodon degus as well as human and mouse FISH, G-banding, C-banding, meiotic studies, nucleolus organizer region, whole genome karotyping and effects of amplified heterochromatin repeat size heteromorphism on hybridization, as well as fragmented but not duplicated ancestral syntenies.
Moving on from this particular non-debate, we could ask if some other tetraploid amniote might be discovered: yet thousands (admittedly not all) of extant species have been karyotyped. This represents over ten billion years of branch length years with no tetraploidy observed to date and none expected on theoretical grounds. Recent paleo-polyploidy is impossible in any of the 30 sequenced mammalian genomes on all-vs-all blastp grounds.
Chromosome painting should not be dismissed quite yet as buggy-whip technology in the genomic era (which is yet to complete a single deuterostome genome). The resolution may be low (4 mbp) but it has two very great advantages: truly complete genomes and lots of them. That plays out in a recent paper thrashing the bioinformatics of inversion as applied to misassembled chicken.
By karyotyping many genomes, it becomes possible to scan for symplesiomorphic species (such as Xenarthra) that retain a high proportion of ancestral gene order, rather than scrambled genomic species like gibbon that confuse more than inform reconstruction efforts.
Genomics has retained its traditional focus on boutique model organisms and rarely surveys for sensible species (Callorhinchus and Fugu being the exceptions). That needs to change because species with slowly evolving gene sequences are a far better guide to the origin of vertebrate genes.
To summarize, whole genome duplication never occurred during amniote evolution and was never an option. It is a paleo-process at best, irrelevant to recent vertebrate gene histories.
Determining paralogon histories
This section will consider the problem of determining boundaries of ancient paralogons and parental source in the face of subsequent gene loss and later expansions, as complicated by local and large-scale inversions and other chromosomal rearrangements. A major technical issue here is quality of blastp matches and ambiguity arising from multiple paralogs with highly uneven rates of evolution.
Evidently it is more favorable to consider more recently created paralogons than those too muddied by later events. However if too recent an event, then gain and loss processes will not have had time to be clearly played out. We don't expect the gene retention pattern to be the same when deeper clades are compared, nor that surviving genes will reside predominantly on the parental block.
Why some coding gene duplicates survive in paralogons but not others has no ready explanation. Genes at the boundaries may experience difficulties if their extended upstream regulatory apparatus was truncated by the duplication (or non-duplicative rearrangement event. It will be quite difficult to gain statistics on this because genes do not persist very long in detectably recognizable form.
Paralogons are a living laboratory in the sense they are created simultaneously, share a similar chromatin milieu and indeed inhabit the same nucleus of the same individual in the same species. Being subject to the same subsequent mutational processes, they serve as internal controls for the evolution of the others.
Here we know already from the mid-size paralogon on chr20q/chr8 that matched pairs of genes diverge at vastly different rates despite the common circumstances. This proves that it is impossible to infer the date of a gene duplication from the divergence of amino acid sequence because each matched pair would give a different date. It's not clear that discarding outliers or using median or average rates improve this situation.
The issue then is whether synonymous change at 4x codons is any better. Put another way, paralogons allow measurement of the real dispersion of these values, given that they should all be the same (up to some narrow statistical fluctuation).
(to be continued).