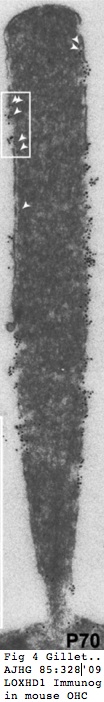
LOXHD1 SNPs
Introduction
LOXHD1, a large mis-annotated gene on chromosome 18 consisting solely of concatenated 120-residue PLAT domains (also seen in polycystin/lipoxygenase/alpha-toxin proteins), recently surfaced as a causative gene for presbycusis or age-progressive postlingual non-syndromic hearing loss type DFNB77. MYO3 (myosin IIIA) and DFNB59 (protein product pejvakin -- echo in Persian) are the only other genes to date with this autosomal recessive progressive phenotype.
There is no evidence suggesting that the three encoded proteins physically interact within auditory hair cells or could cause digenic disease (as perhapss seen in the Usher Syndrome complex). Perhaps with larger numbers of pedigrees, these conditions might exhibit syndromic effects (eg on vision or kidney function as well). LOXDH1 is specific to hair cell stereocilia within the cochlea but is also expressed in testis and placenta and other cell types (not including retina or RPE to date).
Effects are not seen during development but rather in mature stereocilia which degenerate over time. LOXDH1 binds to the inner plasma membrane of stereocilia, perhaps organizing protein binding there. As with other concatenated domain proteins such as CDC23 and USH2A, subtle dysfunction in one of many internally repeated domains evidently cannot be compensated for by the others. Here the mouse I1342N disrupts the hydrophobic pocket between two beta sheets of the tenth PLAT domain but the protein is still made and localized properly. The first morphological change is observed at day 21: fused stereocilia and membrane ruffling at the apical hair cell surface in the basal cochlear turn. Secondary effects arise later and include damage to spiral ganglion neurons.
The one known human allele R670* would eliminate all but the first five PLAT domains (of 16 total), possibly producing a normal but shortened protein in contrast to mouse, but might not produce protein at all due to nonsense-mediated decay. These individuals do not experience tinnitus, balance problems or vertigo, suggesting normal vestibular function even though LOXHD1 is expressed there too.
One difference between cochlear and vestibular hair cells is retention of the apical kinocilium in the latter. This is lost in developing cochlear hair cells as stereocilia mature. This correlates with peak LOXHD1 expression, suggesting to Grillet et al that LOXDHY might help stabilize the stereociliary bundle in the absence of kinocilium.
The kinocilium is a non-motile cilium-like structure present in the crista ampullaris of the semicircular ducts and the sensory maculae of the utricle and saccule but not in the cochlear duct organ of Corti (where they degenerate following maturation of the stereociliary bundle).
Since retinal receptor cells are modified cilia and LOXDH1 acts in hair cells after the kinocilium is gone, a syndromic effect is not expected based on kinocilium/photoreceptor homologization.
Phylogenetic occurence
The PLAT domain is quite ancient, readily tracing back to prokaryotes. However in other proteins it occurs only as single copy, often in conjunction with one or more catalytic domains. A common denominator of PLAT domain proteins is their association with lipid or membrane metabolism. This is perhaps mediated by extruded tryptophans which extend out from the standard beta barrel surface.
LOXHD1 itself has highly conserved orthologs throughout vertebrates, indeed deuterostomes (though the Branchiostoma gene model XM_002606291 contains other domains). These species all contain mechanosensory cells with ciliary F-actin based protrusions. No counterparts of the gene can be found in Ecdysozoa (strictly non-cilary mechanosensation) yet lophotrochozoa (Schistosoma, and perhaps Aplysia) can have full length counterparts, as can cnidrians (Hydra, Nematostella) though gene models here seem fragmentary.
This establishes gene loss in the ancestral ecdysozoan, rather than gene innovation in Bilatera. However the overall impression is phylogenetically spotty occurence outside of deuterostomes. It's unclear whether some other gene product can compensate for lost LOXHD1 or whether it does not play a central role.
Most surprisingly, full length orthologs of LOXHD1 with 16 PLAT domains and very similar intronation patterns are readily recoverable from early eukaryote genomes, notably Trichoplax and the choanflagellate Monosiga. This seems to rule out an obligatory association with F-actin-based mechanosensory organelles as the basal metazoan Trichoplax exhibits only 4 distinct cell types and the early animal Monosiga is unicellular. This rules out LOXDH1 as a metazoan innovation. Lateral gene transfer makes no sense in this context and is contradicted by the gene tree. The extraordinary conservation Monosiga to human -- surely in the 95th percentile of all genes -- suggests conservation of some fundamental intracellular role.
In the plant world, Chlamydomonas and Micromonas have proteins (XM_001703369, XM_002502455) with 20 PLAT domains and numerous WD40 domains (ie are not necessarily orthologous to animal pure poly-PLAT proteins). A curious note on the genbank entry states "identified by comparative genomics as being present only in organisms having motile cilia."
Within deuterostomes, the large length of the gene does not sit well with gappy, incomplete and sometimes chaotic assemblies. Chicken, lamprey, amphioxus, tunicate and sea urchin are among the many that do not yield satisfactory full length gene models, in part because transcript support is spotty. However these species do contain long poly-PLAT proteins no doubt orthologous to human LOXHD1. Some of these unreliable sequences are provided in the reference sequence compilation below. Ironically, the obscure Monosiga and Trichoplax have better assemblies and more accurate gene models presumably because their genomes are much smaller.
Thus within vertebrates it is only feasible to compare selected PLAT motifs that fall with an exon or two. Here there is a risk of using best-blast because many cross-ove matches will occur to other PLAT domains within the same protein, so some additional signature must be used.
3D Structure
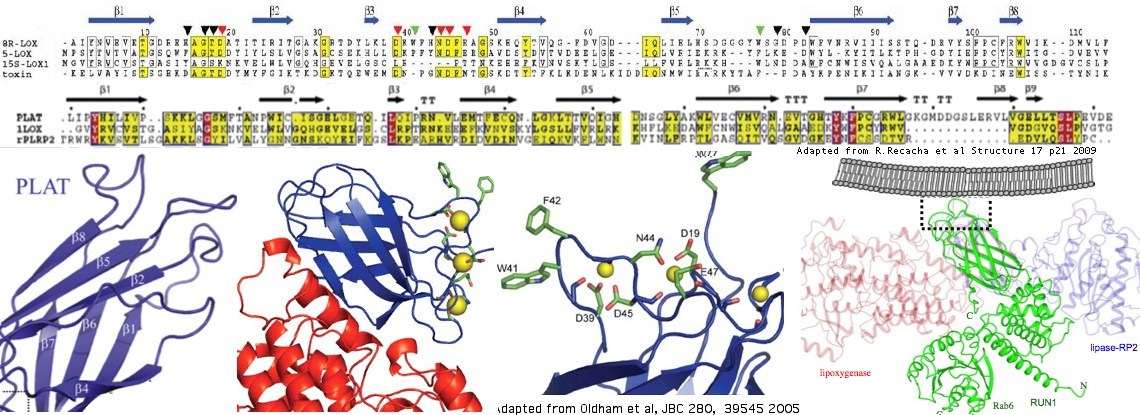
The structure of three proteins containing single PLAT domains have been determined. The best blastp of LOXHD1 to these is a coral lipoxygenase/allene oxidase natural fusion protein with PDB accession 3DY5. Lower quality but still useful matches occur with RAB6 (accession 3CWZ) and rabbit reticulocyte lipoxygenase (accession 2P0MA). The basic structure of a PLAT domain consists of a sandwich of two 2-stranded beta sheets (beta barrel) with certain conserved hydrophobic residues reaching out to the membrane. The lipoxygenases are calcium-binding via extended pairs of acid residues but neither these nor the hydrophobic residues are conserved in LOXHD1.
The independently folding PLAT domain is sometimes denoted as LH2 or C2 depending on the structural publication and domain tool (eg Pfam or SMART). The secondary structure is so basic that convergent evolution (non-homologous origins) must be considered, as with TIM proteins. Here however significant blastp matches at the primary sequence level cannot have originated repeatedly by chance given the vast combinatorial space available to amino acid sequences. Further, all PLAT/LH2/C2 domains share characteristic conserved residues. Thus these domains are in fact homologous (descended from a single source) and so multiplicity of domain names is inappropriate. PLAT is used here because it is an acronym of all protein classes containing the domain.
C2 is unacceptably short and unspecific nomenclature (does not work in PubMed or google searches). The name implies calcium binding yet some domains in this class function independently of calcium. It's not clear whether calcium binding is an ancestral feature or derived. If ancestral, it's not clear whether binding has been lost once or multiple times in different wings of the domain tree. LOXHD1, which tracks back to the earliest unicellular animals, lacks capacity for this binding in all 16 of its domains in all extant descendant clades.
The 12 other human proteins with significant blastp matching to the PLAT domain of LOXHD1 are shown below along with flanking domain context. After adjusting for gene expansions (paralogy), only 3 classes occur: polycystins, lipoxygenases, and RAB6-interacting. These all contain a single PLAT domain.
The 12 other human proteins with significant blastp matching to the PLAT domain of LOXHD1 are shown below along with flanking domain context. After adjusting for gene expansions (paralogy), only 3 classes occur: polycystins, lipoxygenases, and RAB6-interacting. These all contain a single PLAT domain. The final column shows its best-blastp to LOXHD1; no strong pattern emerges though perhaps PLAT:09 may conserve the most ancestral features and PLAT:15 the least. (The first two columns at bottom shows the blastp score of a given human PLAT domain averaged over the 12 other genes; the second two columns show the average match of other genes to human PLAT domains.)
PKD1L1 GPS-TM-LH2-TM chr07:47780815 polycystin-1L1 pdb:3CWZ PLAT:14 36% PKD1 GPS-TM-LH2-TM chr16: 2078712 polycystin 1 pdb:3CWZ PLAT:01 38% PKD1L2 GPS-TM-LH2-TM chr16:79691985 polycystin 1-like 2 signal pdb:3CWZ PLAT:11 35% PKDREJ GPS-TM-LH2-TM chr22:45030224 receptor egg jelly signal pdb:3CWZ PLAT:11 32% DENND5A RUN-LH2-RUN chr11: 9116949 RAB6 interacting 1 pdb:3CWZ PLAT:09 42% DENND5B RUN-LH2-RUN chr12:31426424 RAB6 interacting 2 pdb:3CWZ PLAT:09 37% ALOX5 NH2-LH2-lipox chr10:45189635 arachidonate 5-lipoxygenase pdb:2P0M PLAT:01 37% ALOX15 NH2-LH2-lipox chr17: 4480963 arachidonate 15-lipoxygenase pdb:2P0M PLAT:03 26% ALOX12 NH2-LH2-lipox chr17: 6840108 arachidonate 12-lipoxygenase pdb:2P0M PLAT:09 30% ALOX15B NH2-LH2-lipox chr17: 7888129 arachidonate 15-lipoxygenase pdb:2P0M PLAT:02 34% ALOX12B NH2-LH2-lipox chr17: 7916679 arachidonate 12R-lipoxygenase pdb:2P0M PLAT:09 29% ALOXE3 NH2-LH2-lipox chr17: 7939943 arachidonate lipoxygenase 3 pdb:2P0M PLAT:16 32% best PLAT best Gene PLAT:09 128 PKD1L2 152 PLAT:16 123 PKD1L1 142 PLAT:02 123 PKD1 138 PLAT:14 121 PKDREJ 125 PLAT:08 121 DENND5A 108 PLAT:01 115 DENND5B 105 PLAT:11 112 ALOX5 108 PLAT: 6 108 ALOXE3 82 PLAT:05 108 ALOX12B 77 PLAT:03 105 ALOX12 72 PLAT:04 99 ALOX15 72 PLAT:13 92 ALOX15B 70 PLAT:07 92 PLAT:10 88 PLAT:12 83 PLAT:15 60
Below all PLAT domains found in human proteins are aligned with the 16 domains of human LOXHD1. The coral lipoxygenase is included because it provides the template for beta strand assignment and locating the problematic calcium binding and hydrophobic extended residues. The lower alignment is just with human LOXHD1 domains and uses an optimized seventh PLAT domain (the KE expansion is removed).
Individual PLAT domains can be quite conserved. That is illustrated by a difference alignment of the fifth PLAT domain in 40 vertebrates (human to lamprey) below. Note columns where human sequence represents a unique change -- such as H->R in column 5 -- may be minor alleles or disease variants rather than the predominate residue in the overall human population.
01.hg18_6 MARYHVTVCTGELEGAGTDANVYLCLFGDVGDTGERLLYNCRNNTDLFEKGNADEFTIESVTMRNVRRVRIRHDGKGSGSGWYLDRVLVREEGQPESDNVEFPCLRWLDKDKDDGQLVRELLPSDSSATLK 02.panTro ....R.............................................................................................................................. 03.gorGor S..R...........................................................................--------------------------..................G...... 04.ponAbe ....R..............T.....................................................................................-....................N.... 05.rheMac ....R.........................................................................................................................N.... 06.calJac ....R...........................................................K.............C...............................................N.... 07.micMur ....R...........................................................M.............................................................N.... 08.otoGar ....R. ............K.................-..........H.P.V.......T....................N.... 09.tupBel ...R...........................................................K............A................................................N.... 10.mm9_6_ ....R...........................................................K.....V.......................................................N.... 11.rn4_6_ ....R...........................................................K.....V........................................Q................... 12.dipOrd ....R.....................Y..................E..................K.....V.......S.S.............................................N.... 13.cavPor ....R.......F.............Y...................................L.K.............................................................N.... 14.speTri .V..R.....................Y......-.....-------------............K............P......E...L.....................................N.... 15.oryCun ..........S.............K..........................................................TG.N.... 16.ochPri ....R...........................................................K.............S....V..........................................N.... 17.vicPac ...R.......V.............................I.....................K.................F.E.........................................N.... 18.turTru ....R...........................................V...............K.....V...........F.......................---------...........N.... 19.bosTau ....R.....................Y.....................................K.................F.E....................-....................N.... 20.equCab ....R...........................................................K............A....F.E.........................................N.... 21.felCat ....R...........................................................K............G....F.E.........................................N.... 22.canFam ....R...........................................................K............G....F.E.........................................N.... 23.myoLuc ...HR......D.R......S.....Y............T.K..DEM.Q...............K........T.V.G....F..................Y........................N.... 24.pteVam ...R.........................................M.................K..........R.G....F.E.........................................N.... 25.eriEur ....R.....................Y.....................................K............G....F.A..............M..........................N.... 26.loxAfr ..Y.......-..M.....................A..L.K............G......E.........................................N.... 27.proCap ....R.....................Y..A.......M...........R.........A..L.K............A......E..............M...---...................NN.... 28.echTel ....R.....................Y..........MF.....V..............A..L.K......M.....A......-...LKK.........K..G.G....................N...R 29.choHof .V..R.....................Y.........W.........M....S...Y...A..L.K.K.I.VG.....GN...F.E..............M....N...AR.Q.....I......G.N.... 30.monDom ..Q.R..L...DI.....N.QAFV.....A......M.......MEICQ..........A..V.K......G.....K....F.VK..................N..F...Q..............NSR.. 31.ornAna IVK.R...V..D.N......R.FI..I.........I.........T.........FV.A..LKQ......G.....GS.....AK.I.........EA.....Y.....NE....I....V.AGE.PL.. 32.galGal VIK.R......MVS.S......FV..I..Q....D.V..K.I..VNK.........F..A..LKQ......G.....GS.....AK.I.........EA.....Y.....NE....I....V.AGE.PL.. 33.anoCar IK.R...H..NVS.S....H.FA..I..Q....D.V.Q.SV.TVNK....S....I..A.NLKQ......G.....GS.....GK.I...D.....EAQ....Y.....NE....II...V.AGR.SL.E 34.xenTro ...F.IE.K..MNK.......D.I..A..LKQ.K....G.....G.C...M.K.V...D.K..TEAI....N..F.RNE...HII...V.AGD.QN.R 35.tetNig LK.RI.I...NVG.G....S.F.NII..L......SMITSK..VNK.....H...L....CLGR.S...VG...R.G.C..F..K.T.........SAI....F....HNE....I....V.AGDGRLF 36.fr2_6_ LIK.R..I...NVS.G....S.F.NVI..L.......MIMSK..VNK.....H...L....SLGQ.....VG...R.G.C..F..K.M.........SAI....F....HNE....I....V.A 37.gasAcu LIK.R..I...NVS.S....S.F.NVI..L.......MFMSK..VNK.....H...L....SLGQ.....VG...R.G.C..F..K.M...D.....LAT....F....RNE....I....V.AGDGRLF 38.oryLat IK.R..I...TVS.S....S.F.NVI..L.......M.MSK..VNK.....H...L....WLGQ.....VG...R.G.C..F..K.M.........AAI....N....QNE....II...V.AGEG 39.danRer VK.R...Y..DVS.S....H.F...I..L....D.S.I..K..INK..R......I..A.SIGP......G...R.G.C.....K.TI.....A..QA.....S..F.RNE....I....Q.FADGRLY 40.petMar VK.L..IH..KHG.G......FV..Y.EQ....D.M..TS...VNK..R..V...V..C..LKR.T....G...R.GS...F..KIM.K-D.RRQCEE.....N....D.E....I....VAGGCQMLK Consensus ...yrvt..tg...g.gtda.v..cl.gd.gdtger.ly.crnn...fekgna...t..s.t$rk.r...!r...k.ggs..y$d.!lvre#.qpes.nv...c.r.l.k#k....l!...l.sgsnatlk
Reference sequences (intronated)
The sequences below are broken into individual exons with reading phase (codon overhang) indicated by 012. Alternating colors show odd and even numbered PLAT domains. The 7th domain (which contains the charged KE expansion loop in mammals) is shown in red.
>LOXHD1_homSap_full 41 exons|2273 aa|chr18|span 179,559 bp|no signal peptide|no conserved cysteines|16 PLAT domains|EIW --> WNW mouse deafness 0 MMPQKKRRRKKDIDFLALYEAELLNYASEDDEGELEHEYYKAR 1 2 VYEVVTATGDVRGAGTDANVFITLFGENGLSPKLQLTSK 2 1 SKSAFEKGNVDVFRVRTNNVGLIYKVR 2 1 IEHDNTGLNASWYLDHVIVTDMKRPHLRYYFNCNNWLSKVEGDRQWCRDLLASFNPMDMPR 1 2 GNKYEVKVYTGDVIGAGTDADVFINIFGEYGDT 1 2 GERRLENEKDNFEKGAEDRFILDAPDLGQLMKINVGHNNKGGSAGWFLSQ 0 0 IVIEDIGNKRKYDFPLNRWLALDEDDGKIQRDILVGGAETT 1 2 AITYIVTVFTGDVRGAGTKSKIYLVMYGARGNKNSGKIFLEGGVFDRGRTDIFHIELAVLLSPLSRVSVGHGNVGVNRGWFCEK 0 0 VVILCPFTGIQQTFPCSNWLDEKKADGLIERQLYEMVSLRKKRLK 1 2 KFPWSLWVWTTDLKKAGTNSPIFIQIYGQKGRTDEILLNPNNKWFKPGIIEKFR 0 0 IELPDLGRFYKIRVWHDKRSSGSGWHLER 0 0 MTLMNTLNKDKYNFNCNRWLDANEDDNEIVREMTAEGPTVRRIMG 1 2 MARYHVTVCTGELEGAGTDANVYLCLFGDVGDTGERLLYNCRNNTDLFEKGN 0 0 ADEFTIESVTMRNVRRVRIRHDGKGSGSGWYLDRVLVREEGQPESDNVEFPCLR 2 1 WLDKDKDDGQLVRELLPSDSSATLK 1 2 NFRYHISLKTGDVSGASTDSRVYIKLYGDKSDTIKQVLLVSDNNLKDYFERGRVDEFTLETLNIGN 0 0 INRLVIGHDSTGMHASWFLGSVQIRVPRQGKQYTFPANRWLDKNQADGRLEVELYPSEVVEIQK 1 2 LVHYEVEIWTGDVGGAGTSARVYMQIYGEKGKTEVLFLSSRSKVFERASKDTFQ 0 0 LEAADVGEVYKLRLGHTGEGFGPSWFVDTVWLRHLVVREVDLTPEEEARKKKEKDKLRQLLKKERLKAKLQRKKKKRKGSDEEDEGEEEESSSSEESSSEEEEMEEEEEEEEFGPGMQEVIEQHKFEAHRWLARGKEDNELVVELVPAGKPGPE 1 2 RNTYEVQVVTGNVPKAGTDANVYLTIYGEEYGDTGERPLKKSDKSNKFEQGQ 0 0 TDTFTIYAIDLGALTKIRIRHDNTGNRAGWFLDRIDITDMNNEIT 2 1 YYFPCQRWLAVEEDDGQLSRELLPVDESYVLPQSEEGRGGGDNNPLDNLALEQK 1 2 DKSTTFSVTIKTGVKKNAGTDANVFITLFGTQDDT 1 2 GMTLLKSSKTNSDKFERDSIEIFTVETLDLGDLWKVRLGHDNT 1 2 GKAPGWFVDWVEVDAPSLGKCMTFPCGRWLAKNEDDGSIIRDLFHAELQTRLYTP 1 2 FVPYEITLYTSDVFAAGTDANIFIIIYGCDAVCTQQKYLCTNKREQKQFFERKSASRFIVE 0 0 LEDVGEIIEKIRIGHNNTGMNPGWHCSHVDIRRLLPDKD 0 0 GAETLTFPCDRWLATSEDDKKTIRELVPYDIFTEKYMKDGSLRQVYKEVEEPLD 1 2 IVLYSVQIFTGNIPGAGTDAKVYITIYGDLGDTGERYLGKSENRTNKFERGT 0 0 ADTFIIEAADLGVIYKIKLRHDNSKWCADWYVEKVEIWNDTNEDEFLFLCGRWLSLKKEDGRLERLFYEK 0 0 EYTGDRSSNCSSPADFWEIALSSKMADVDISTVTGPMADYVQEGP 1 2 IIPYYVSVTTGKHKDAATDSRAFIFLIGEDDERSKRIWLDYPRGKRGFSRGSVEEFYVAGLDVGIIKKIE 0 0 LGHDGASPESCWLVEELCLAVPTQGTKYMLNCNCWLAKDRGDGITSRVFDLLDAMVVNIGVK 0 0 VLYEMTVWTGDVVGGGTDSNIFMTLYGINGSTEEMQLDKKKAR 2 1 FEREQNDTFIMEILDIAPFTKMRIRIDGLGSRPEWFLER 0 0 ILLKNMNTGDLTMFYYGDWLSQRKGKKTLVCEMCAVIDEEEMMEWTSYTVAVKTSDIL 1 2 GAGTDANVFIIIFGENGDSGTLALKQSANWNKFERNNTDTFNFPDMLSLGHLCKLRVWHDNK 1 2 GIFPGWHLSYVDVKDNSRDETFHFQCDCWLSKSEGDGQTVRDFACANNKICDELEET 1 2 TYEIVIETGNGGETRENVWLILEGRKNRSKEFLMENSSRQRAFRK 2 1 GTTDTFEFDSIYLGDIASLCVGHLAREDRFIPKRELAWHVKTITITEMEYGNV 2 1 YFFNCDCLIPLKRKRKYFKVFEVTKTTESFASKVQSLVPVKYEVIVTTGYEPGAGTDANVFVTIFGANGDTGKRELKQKMRNLFERGSTDRFFLETLELGELRKVRLEHDSSGYCSGWLVEKVEVTNTSTGVATIFNCGRWLDKKRGDGLTWRDLFPSV* 0 >Gallus gallus gappy genome,flawed XM_425221 15 PLAT domains MKETKKNKAEEEEEEEEEEEEENEEVVEDVVEEAVEEDGEDESQQKKSKKKSKSEDSEDDGEVKKKKKKKKKTKRKEYSSEEEDDYERRKKKKKKGKKSK EKKKKKGDKSKKGKKEDNFLELYEQELRDYHSDSSNATEDEYNKKKVYEVVTVTGDVRGAGTDANVFVTLFGEFGITPKTHLTSKSSTAFERSKTDVFRV KTNNVGQIKKIRIEHDNTGLNAGWFLDRVIVTDMNRPHLRFYFPCNNWLSKEDGDGLYVRDLIGSLNPMDVPKINKYVVRVFTGEVSGSGTDADVFINIF GEKGDTGVRKLDNDKDNFEKGAEDKFTLDAPNLGRLRKINIGHNNKGGSAGWFLAKVIIEDIGNKCVYQFPVGRWFALDEDDGKIQRDILVGGTEATGIV YNVAVVTGDIRGAGTNSKIHVILHGSKGLKNSGTIFLEGGEFERARTDLFNVEIASLLSPLSRVTIGHDNCGVSSGWYCEKVVVYCPFTGIEQTFPCGKW LDEDEGDGLIERELYEMVSLRQRRLKKNPWSLWIWTSDIKNAGTDATIFFQIYGDKGKSDEMKLDNNSDNFEAGQTDKFMIELPDLGTFYKLRIWHEKRN PFAGWHLDKVTLLKTLTKDKYSFNCGRWLDINEDDNEIVRELPAEGSLVTEVMPVIKYRVTVCTGMVSGSGTDANVFVCLIGDQGDTGDRVLYKCINNVN KFEKGNADEFFVEAVTLKQVRRVRIGHDGKGGSSGWYLAKVIVREEGQPESEAVEFPCYRWLDKNEDDGQIVRELVPAGESPLLKNVSYHISVKTGDIPG ASSDSKVFIKLYGEKADTSKETLLVSDNDLGNYFERGRVDEFTIDTMDIGKINRILIGHDNVGLRSGWFLASVQITVPVQGRQYMFPCNRWLDKDEADGR VEVEVYPSEILPIEKLINYEVSVVTGDVRAAGTNAKVFMQIYGETGKTELIILENRSNNFERGATDIFEVEAADVGKIYKIRIGHDGKGIGDGWFLESVT LKRLATKMDETDKKKKKKKKKSEEEEEEEETKVEEVMDVYTFVAHRWLAKDEGDKELVVELVPDGESELEENTYEVHVLTGSVWGSGTDANVFLSIYGIE RGDTGERQLKRSNNLNKFEKGQVDVFTIKAIDLGELKKLRIRHDNSGSSPSWFLERVEIVDLKESTTYYFPCQRWLAVEEDDGQIVRELVPVDEAFVKKD SENDGQSLATLGLEQKAKSTTYIVKVKTGDKKNAGTDANVFITLYGSKDDTGIVSLKASKLNKNKFERGKIDEFTVESVDIGDLKKIKIGHDNAGNSNGW FLEWVEIDAPSLGQCLKFPCGRWLDKSEDDGAIERFIFPAELQTTEYIPFVPYEITVYTSDIFGAGTDADVFIVLYGSDGICTQQKSLCLNKREQRMYFE RNSVNQFIVELEDVGDIIEKIRIGHNGGGLNSGWHLDRVAIRRLLPNGKGSETITFPCERWLAKSEDDGEIIRELVPSDIFTEKLMKDGTLKQIEEEVED PLEVHTYKISVFTGDIYGAGTDANVFLNIYGDLGDTGERKLSKSETNFNKFERGQSYDGERRSLSESSVSLSSLDSSGNPKEKKSRLVKSAEEGLLIPYH ITVTTGTEYDSSTDSRVFIIIMGPQKVRTERLWLDLPEGKDEFADGSVEKFSVWGLDVGEIKKVEVGHDGATPESCWLMEELTIVVPTKGVMYNFVCKCW LARDKGDGLTSRILNILDADCVNVGIKILYEVTVVTGDIESGGTDAGIFMTVFGSNGNTEEMQLDKNGDRFERGQEDSFIMEIADIAPLRKMRIRTDAKG TRPDWFLERIVMRNLTNQEVATFTYGDWLSKVKNAKGSLVCEMPAMVNDEQMMEDTTYTIQVKTSDIGGAGTDANVSLILFGENGDSGTLALKESNKSNK FERNQMDEFNFPNMLSLGDLCKVRIWHDNKAYEIVTVTSNREDAETKENIWIILEGKLGRSKEFLMENSSKKRRFERRGSTDTFQFSSKNLGDIAAICVG HCPKDGKKSSAKADVYWHVKEIIITEMELCNKYFFRCNGKIPLRYKRRDYKVFECAKVIESFASKARSLVPVKYETIVVTGFEKGAGTDANVFITIFGLN GDSGKRALKQKFRNLFERGKTNRFYLETLDMGELKKVRIEHDNSGLAPGWLVERVEITNSATGVTTIFPCGKWLDENRGDGLTWRELFPRY* >LOXHD1_cioSav Ciona savigny approx gene model 15 PLAT domains 0 EVIVEDTKYDRTYTFPCDRWLSLYKEDCQLTRHLMAVSGRQS 1 KRTTFEITTVTGNGEKRGAGTDANVFLTMRGSKGVSPKLQLKSEYCSRRNTFERGQSDVFRVKSVNVGNLKRIRMEIDDTGFAPAWFLELQVVVVDLANPSQRIYFPCGQWLSDDEQLFRDL VGSSNPLAR 2 PKSVNAYVVHTFTGDVRGAGTDATVKITLFGDHGDSGQLVLDDSKNNFERARKDTFSLQCPHLGKIKKIRIGKGHDNGGVSPGWYLDKVVVDDTMMDCVYSFPCQRWFAK 3 DEDDGRITRELVAGVGEAGIPYQVQVFTGRFTSSCMGGRRVKSVQGKSILDGKFQRNRVDICDIESTTMLSPLDHIEIGHDNSGTGPGWYLEKVVITCPTNGCEQVFSCRKWLATDEGDGLISR QLYESKSLRK 4 QVVQSAWNCLIWTSDVRNAGTDANVFIQVYGENGKSDELPLDNETDNFETGQKDKFKINLPEIGCIYKLRVWHDDSNPFSGWHLDKIELEPIGGKKGDLYTFKCGKWLDTNEGDGEVIREIPST GPGVKR 5 PQPLVRYSVSVVTGDKRGAGTDANVSCCLYGRQGDTGNRPLTASKNNRNKFERGQLHLMLIIQTQHHRWLATDEDDGAIVRELTSEGSP 6 QLLNTSYHVAVKTGDIRGAGTDADVYIQIYGSDGDTGRLKLTRGENIRNKFERGRTDKFKLEATDIGRLERIYIGHSDSGIGAGWFLDGIEIDVPSSGMHYTFKANRWLASDEGDGKTEIELYPT QTR 7 QQDKLPYEITVATGDTLNAGTDAQVVLQLYGEDGKSEVMKLRNKTDNFERKAIDKFKIEAPDIGPITKIRIGHDGRGVGSGWFLDSVKIQRFVHKTSKRKKRRRGQNQSEVETYWFVCKRWFDKSEDDGQ 8 IIRELEYTVDVYTGDKYGAGTDSNVFLTLYGENGDSGERKLDKSVNINKFERNQVDSFKLKAADLSTLKKLKIRHDNSEIGSSWFVDRVEVKVKGETWVFPCQRWLSTKEDDGQIERTLVPV DATYQRLQQQKQGRKASIAVRDAIALET 9 KAQLTTYEVSVKTGDVWGSGTDANVYLTLFGEKDDSGINLKTSKTYRDKFERNHIDVFTVESADLGVLKKIKIGHDNAGLSPGWYLEYVEVDAASLGMKWRFPCGRWIAKDKDDGKLEKELHLAT DETVG 10 YAAKVPYEITVYTSEISGAGTDADVYVVLYGREGIATEQTSLCSSKDERKSRFNKKGIDTFVLELDDIGEVIEKLRIGHDGKGWGAGWHLDKVEVRRLLEGVKGSETSTFTCGRWLARDEDDGEIVRELVPT KIVKEQVTKSGKLKRSESKLD 11 DALKQQYKVHVFTGDVFRGGTDANVFITIFGENGDTGERKLLKSETNTDKFERGKEDIFKLDAVDLGKLFKVHIRHDNALIQPAWFLDRVEIEDTDTGSIYVFHCERWLAKNKEDGKIARSLYVV GYDGETSTRRSLATTKSGGSLGKKSILSDTIP 12 EGPAIPYTVKIETGSEEGSSCKAPAFIKIFGPSKKGTVSRRIPLEPYSGKFKRGTIETFNIEAIDVGSIKKIEIGHSSVDEEWFVESVMVNQPTLGKSYTFKCHKWFARSKDDGLVSRIFTTD DAEKLS 13 FKAKIPYEITLHTGNVDNAGTDCTITLSVYGSKGNSPDITVKKSEEDGVLLERGSIDTINVELDEIAPLKMVRVGHDGRGQRADWFCDKKTLSRELSASVGGRS 14 VLKKTNYKINVKTSNLRSAGTNANVSCIVFGTFGETGELKLKVSETHKNKFERDAMDTFTFQDKLSVGEPQKLRVWHDSKGFSSGWHLDHIEVCDESTQQKYMFPCGRWLADDEDDKQTIREL TCSNLDQ 15 AKEGGQYTATVRTSDKRDAAPNSDGWVILHGKMGKSRRMKLREGKLGRGKASTFKLDSNDLGALQKITLGLEGDNMGRWHVDDVVIENPSGGQFKFDCHAWVSNEDGTNTRGSNFDC TATKGRTISMS 16 SLLPVKYEVTVVTADEKGSGTDSNVSLTIYGTNGDSGKRPLKQRFRDLFERKQTDKFTLEILDLGDLTKVQIRLEHDDAGFNADWLCDNVTIRNPVSGQSWTFICREWIGLKKANQLFREVLPR >LOXHD1_cioInt Ciona intestinalis XM_002122635 defective assembly; frag gene model 16 PLAT domains EVIVEDTKYDRTYTFPCDRWLSLYKEDCQLTRHLMAVSGRQSKRTTFEITTVTGNF GEKRGAGTDANVFITLRGSKGVSPKLQLKSD 21 RHNTFERGNSDVFRLKSANVGNLKRVRVEIDENGFAPAWFLERVVVVDMSKPSQRIFFPCGQWLSDDEQLFRDLVGSTDPLARPKM NAYVVHVFTGDMRGAGTDAEVFITLYGGKKGQLSSGKIVLDGKFQRNRVDICDVESASMLSPLDHIEIGHDNSGSGPGWFLDKVVVTCPTNGCEQVFSCRKWFATDEGDGLTSRELYESKSLRKQVMQK SAWNCLIWTSDVRNAGTDANVFIQVYGENGKSDEIALDNETDNFETGQKDKFKINIPEVGRMYKLRVWHDDSNPFSGWHLDKIVLEPVGGKKGSSYTFNCQKWLDTNEGDGQIIREIPATGPGVKRPQPL VRYQVTVVTGDKRNAGTDANVFCCLYGRQGDTGNRPLEASKSNRNKFERGQSDEFTIEAVDLKSVNKVKIGHDDSGSGWHLKEIIVRDETNGESTVFPCDRWLATDEDDGAIVRELTSEGSPQLLNT TSYHVSVKTGDMRGAGTDADVYIQIYGSDGDGRLKLHQGENIRNKFERGRTDRFKLEATDIGRIERIYIGHNDSGVGAGWFLDGVEIDVPSSGMHYTFKSNRWLASDEGDGKTEVELYPTQTRQQDKLL PYEITVTTGDIMNGGTDAQVVVQLYGEDGKSEVIRLRNKTDNFERKAVDKFKIEAPDIGPITKIRIGHDGRGVGSGWFLDSVKIQRFVRKTSKRKKRRRESMVSNDQNEVETYWFVCRRWFDKSEDDGQIIRELVPTDESGKPIDEALQE LEYSVDVYTGDKYGAGTDSNVFITLYGENGDSGERKLDKSDNINKFERNQVDGFKLKAADLSNLKKLKIRHDNSELGSSWFVDRVEVKVKGQTWVFPCQRWLSTNEDDGQIERTLVPVDVATYQRLQQKQGSGRKASVAVRDAIALETKAQL TTYEVSVKTGDIRGSGTDANVYLVLFGENDDSGMINLKTSKTHRDKFERNHTDVFTVESADLGNLKKVKIGHDNAGLSPGWYLEYVEVDAPSLGMKWRFPSGRWIAKDKDDGKLEKELFLAMDETVGYAAK VPYEITVYTSEVSGAGTDADVYVVLYGRDGMSTEKTSLCPTKDERKSKFNKKSVDTFVLELDDVGEVIEKLRIGHDGKGWGAGWHLDKVEIRRLLEGAKGSETYSFTCGRWLARDEDDGEIVRELVPTQLVQEKVTKS QQYKVHVYTGDVFNAGTDANVYITIYGENGDTGERKLLKSETNTDKFERGKEDIFKIDAVDLGKLFKVHIRHDNALLNPSWNLNRVEVEDTDTGNRYVFHCERWLAKGKDDGKIVHVYTGDVF IPYTVKIETGSDEGSSSTASAFIKIYGPSKKGKVSRRITLDPPSGKFKKGTVESFNIDAIDVGTIKKIEIGHTGVDDEWFIESVMINQPTLGKSFTFKCHKWLARSKDDGLTTRIFTIDDAEKLTYKAK IPYEVTIHTGNVPNAGTDCLITMSVYGTKGNAPDITIKKSEEEGILLERGSVDVFNVEMEEIAPLKMIRISHDGRGQRQDWFCDKIELRNLNSGKVTVFPCDSWLSKKLGEKTLSRELSASVGGRSVLKK TTYKLNVKTSNMRSAGTNANVSCIVFGTFGDTGELRLKESETHKNKFERDSLDVFTYQDKLSIGQLQKLRVWHDSKGFSSGWHLEYIEVVDESTQQKFMFPCNRWLADDEDDKQTIRELTCSNLDQATEGHTV GQYTATITTGDKRDAASSCDGWIILHGKMGKSRRLKIRSGKMGRGKKSEVKLDAGDVGKMEAITLGLHDMGRWYVEDVTIETGNGGQYKFDCHAWVSNEDGTNTRGRTFECSATKGRTVSMSSLIP IKYEVTVVTADEKGAGTDANVKLTIYGENGDSGSRALKQRFRDLFERKQTDKFTLEILDLGELTKIRLEHDDGGFNADWLCDHVIVRNPVIGQSWTFVCREWIGTKKAGQLFREVMPRSTG* >Schistosoma mansoni (flatworm) XP_002576380 2301aa Bilatera|Platyhelminthes blast 0 MNLEDSTNDDYLKNSKSRFSFHRKEEKEHKGIICCGHNTCHLDRYNLISSVKPFTARPYTLYKVVLTTADVPGAGTSAQ 1 2 IYITLKGEWGSSTRQKLRKEKVPTRNLRFYFYPGSTNTFSVVSPDLGGLHSVFIE 0 0 HDSLRKSDSWLLESVQVFHPLTKKRYMFMCNHWFSLYKEDGLIARELFGIRSAKT 1 2 KYSIVTVTGDQEGSGTNSKVYITIYGRTGITPRIELSQENKSTGKDILCAPFGRGTSTKFIVKAPNVGAITNIRIKQDESGNEPHWFLERVVVTDMSYPQWTYYFHLSCWLSSKYGDGKSCRLVRGYREPTGTGV 1 2 ETEYKLTFYTSDQKDAGTTGEVYVKLYGEKDSSREIWVNSINQKNRRQPTSYHFTRGSTVEVYLPPCPQLGEITKLKVGHNRAGSSPSWFLNK 0 0 VIVDDLRMNRVFEFPCYAWITKPSETIIVCQKSIKKEEKEIIWQ 1 2 KIPLEIRLYTGDVPNAGTTAKVYL 12 HQKTTSENKEDVYTTPYIWLEDGNYDRNGVTLFSIDLPVPKFISPLSQLVIGHDNTGHSPSWFFDK 0 0 VQIYCPLNGVEQTFLYRKWLTSSKPGMKVEQILCEEKSLRKCTDK 1 2 KIPWEISVKTSPIANSYVTAAVSIIIFGSKDKTMKIQLDRSNLEVSSEWTKVSGNEVVEMFRPNVESKFRIYIKDIGIPCKLRIQHDNKGSNPNWHLQE 0 0 IVLTNLRTHEQYEFYCNRWLSTREDDGSILREIPAKGPGITNPLP 1 2 LYHYIIQIFTGNKPNSGTNANIFINIFGEKGDCGERWLGRSVNRNSELFQQNQ 00 MDEFVIEAVQLGSINKICIGHEERSPGYGWYLAKIVLTIKENPKYKLTFECYR 21 WFDVGEDDGQIVRELFAHSSLS 1 2 AIAYNVTVLTGTCRNAGTVANVFVHLYGLQGESKDMQLKHKETEITKFEAGKSEEFILACGKLGE 0 0 IKKIKIWHDSIAPHSGWFLDEVVVIDYFLGRRYVFHVNRWLAKNEADGLISLDLQPTSIINIER 1 2 KSSYEVIVKTGGRKYAGTDSHVYITMFGINSESKEYHLANSKTHNNKFEQNHEDLFN 0 0 LNAVGLGSLKKIRVRHTNTGIAPGWFLDYILIRESIEQKKMEYFFPCYQWLSATRLDGLVIRELSVASENIFERWKSGHDISDDSRLILE 1 2 DQSTIYHVRIHTGSQSNISSDANVQIQLYGDKDFTGNIRLYKAYYGKEDILVNKFQAGQIANFIVKAINIGNIKKIR 2 1 IEHDTAVGSARWFVERVEVEARKLGLLWKFECNRFITPQQPELILYPEPKDISFVKP 1 2 MTNYQIKTFTSDISGAETTSSVYIQLYGNDSLPSSIRRLHQNNDSEQRFQRNKIDTFYVELEELQEPFSKLRIWHNDKGSSTDWHLNKVEIRKIKTNQY 0 0 VFVTYIFPCNKWLSRNMDQAALERELIPSHMLQDQNGVLQFERQLSPKWLMNTYEVRITTGDKAYAGTDASVYITLFGENGDSGERKLTKSLTHRNKFERGQ 0 0 TDVFQLEIVDLGKINKVRIRHDNSGVNPSWYLSTIEVFNISKSQSVHGLKDQPNETHHLVKYIFNCEAWLSTEHDEKVLDRFFSAEIIKPQDLPGIVPKTSKDIVPHKLIQDLSQGKKALE 1 2 TIPYLITVITGSDRKAGTPGPVWISCVDKDKMNSEKFILCDCYNRTMLKRGTTRYFRFAGVKLNDLTEIQ 0 0 VGNDAPESPSMGWYIQSLYLSFPTIGKMYLFDCKEWLSTNRGNKKRTCNLKISNTNIVKFKP 1 2 KITYHLKITTANVKRAGTDCSINLQIFGTNGVTNCYILEKTSNRFAQGITDNISLEMEDVGKLLKMRIGHDNQ 0 0 GKNKHWNLSCVEVTVANTNQLYRFVYDDWLSLTYGKRKSLWADLPAMIGDTVQLK 1 2 ETCLDIFVKTGNMPASSTDANVYVQLFGEYGDSGEILLKQTVSNQKPFQNNS 0 0 IDHFKIPSILKLGNLARCRIWHDNKGSSPNWYCEWLEVKEVLIPGEKNLACNWKFAFNKWLSVSDDNKQLLRDAPCSEVYMNDSKGHRTIDQESIETLLTTADSMNISDLKDNPEGKL 1 2 VYEVVIETGNLKDSGTTCDAWIILEGKHGRSPKLELVNQVGNPILQINQMNTFQ 2 1 LPSFPLGDLETIRLGIQERNINKQTNPNDIQSQKWFCEKVSIKDPVSKRTYIFTIHQWLSVTPTSNLKKDVLVNYSKIIEDPYKKAFNELK 1 2 NKQTVMYKVSIYTGTKGCANTDANIFITMFSTTPGLNSGRIALKRENNNLFDRKQLDEFYVESIDL 1 2 ESIQRIIIEHDNTGVSPDWYLDKVLITNQTNNQIHLFQCYQWISKKKGDCRLWKELLVSN* 0 >Trichoplax adhaerens (trichoplax) Metazoa XM_002107971 17 PLAT domains blast 0 MSASLAMQELMSDPSGHFAPKPPPGRSKINRKRPQT 1 2 APGRQITTASKAPKLWRSSTYNEPNLKRRKFIVTLNHKKDISSISRNAPYANRAYTTAMATAARTYSNALNKGQQKEIIPIYNPLCDPHLNDYYARKFGLLNSRD 0 0 ESRKNRKQ 1 2 AGVAYQFGVKTGDKKGSDTDA 2 1 VYIQVIGTKDKIPKKRLFKKQETEKTERGNLFKFDKSTVEKFAVQHRDIGDPVKLIVE 0 0 HDGNEKRHGWFLEEITLTNIQSKKSWLFPCHKWLSKYEGDRKLCYELKPLAKAGKA 1 2 VYEVSVLTGDKRGAGTDANVSVTLFGKHTSSPKIQLLKS 2 1 SKHKNPFERNNTDEFKIRTRDVGKLSKIRIEHDNAGFGPGWFLDK 0 0 VIICNLEKPNVKYYCPCNQWLAKDVGDKSISRDLTAYTDPNAAPS 1 2 AYVYIVHTFTGNKRGAGTDANVYAVIFGDSGDTGEKRLDNSKNNFEKSR 2 1 KDTFKLSCSCVGKLERLRIRHDNTGLFAGWYLDKV 1 2 VVEDPQEQQSYTFYCRRWLSKTEDDGEICRDLIVSASGDDGDDSVAPK 1 2 GYPYHIHVTTSDVKNAGTDAEVYVVMHGEGKKSKELNSGKLVLANSEKKKNTFERAMTDIFHMECAEMLSPLTKLTVGHDNKGLAAGWHLDR 0 0 IVIDCPTTGIEQTFLCQQWLDRKAGDGLTERELVEAFDMRKTRRP 1 2 KQLWFAWIWTSDIRGAGTDANVSMQIYGDKGKSQEIKLGNNTDNFEQATLDKFK 0 0 LEIDQVGVPYKLRIGHDNSNAFPGWHLDKVKLENMNDKEQYLFNCNRWLSRSEEDNEIIRELPASGPNCPNYP 1 2 IVIYEVSVHTGNKMGGGTDANVFIKIYGELGDSGYRPLKSSKSHNNKFERNQVDVFHIEAVTLKALKKIKIGHDGNNP 1 2 GAGWFLDKVVIKELNGEASNEFPCNR 2 1 WLSKSEDDGQIVRELFLKSDTPLLK 1 2 TTSYHISVKTGDVRNAGTDANVFIQIFGAKDDTGRVRLKQSLNTSNKFERNRIDKFIIEAAQIGK 0 0 IEKIIIGHDGKGLGSGWFLDYIELDVPSVGRLYRFSCHQWFDSTEGDRKVERELYPSECIKSAA 1 2 KIPYQISVHTGDIRHAGTDSNVFAVIYGENGKTEELKLRNKSDNFERGQVDVFK 0 0 VECEDVGKLRKLRIGHDSAGMGSAWFLDK 0 0 VYVRRLPPKSGKKSKETDEREEETAKKDADEPEKLDENNYLFVANRWLSKEEGDRQTVIEISPVGVDGALA 1 2 EMTYTIRVITGNKFGCGTNANVFINMYGEEGDSGERQLKKSETHTDKFERNQ 0 0 EDVFKISCLSLGELKKIKIRHDNSGFRPAWFLDKVIIEVGESKYQFMCDRWLAKDEDDGQISRELLPQSDEQSRAEAIGASKDLQKK 1 2 VASTTYNVSVTTGDIKGAGTDANVHIVLYGEKDDTGLIHLKNSTTHSNKFERNQEDRFVVEAIDIGELKKIK 2 1 IGHDNKGGMAGWFLNKVEIDIPSLGRRLLFPCGRWIDKGKDDGALERELYPLNEAEETYRP 1 2 HIPYEVTVYTTDKRGASTGANVYVVIYGEENQTEQASLEPDKKRRKQYFKNGAIDKFVLE 0 0 LDDVGEEITKLRIGHDGKGWGAGWHLDKVEICRLLDGGKASKKFTFQCNRWLASDEDDGAIVRELVPSEIVEKSSKDGGQVKTKVTKPTDGLK 1 2 VKPYTIHVFTGDVDGAGTNANVFLTIFGESGDSGERKLAKSDTHYDKFERNQ 0 0 EDIFHIEAADLGRLFKVKIRHDNTGSLFSPAWFLNRIEIVDDENEETTAFPCERWLAKKKDDGKIDRTLFVKGWEGDTSSVATTRSK 1 2 ASFRSNSTDLGEMVSASGRRASSRKGPVMAPIDEK 1 2 CVPYNIKVTVGEESSKNFQETLHLELFGQIEEEKSGPIELSPEKKSDKTFYPGKITTFYVSAAEVNIIEKIQVS 1 2 HNSYMPDSGIYLKEIEVDVPTIGNKYIFPCNRWLAKDKDDSKTSRIFTAANAQVTSYTA 1 2 GTDSNIFVILFGTKGRTPEISLEKNEDRFERAKVDIIP 0 0 LELDDVGTIKKIRIGHDGKGSRTDWYLEK 0 0 ASIQRMDTLDMYMFRANQWFSKKIDDKKLVREIPAETSKEGATTIK 1 2 KINYVLSTHTSDKRGSGTDANVFVIIFGENGDSGEIALKKSETNWNKFEKGQTDVFLINDRLSLGRLQKLRIWHDNA 1 2 GFGASWHLASVDIVDESTGVKYTFPCDKWLSKSNGDKLILRELPCAETAGNTAASKATKQESKSGKA 1 2 EYEIAFTTGTEKRAGTNQDVAIVLKGKSEKSREFLIENNEDKKYFSKGKTNKFTYTCKPLGDITKAIVSHRESAIGEEPDSKNSSWYLKVVTVVHKASGTT 2 1 YKFPCNKWIDLDDDEENNSSVTLKCKSADVAASSKAKPV 1 2 ELKPVKYEVTVVTGDEKGAGTDANVSVILYGDNGDTGPRPLKKKFVNLFERNQHDKFTIEALDLGKLTKLHIEHDNKGWGASWLLDRVEVHNVDSNETIIFPCKQWLDKKKGDGQIAKDLLPES* 0 >Monosiga brevicollis (monosiga) Eukaryota|Choanoflagellates XM_001742822 15 PLAT domains MADLPVPTTALQRLQLERSRYSFAAADPPPSFVQRYTSADVLSHRLYAQGLASDVGPGSHSSLGMSRRVELPPYNPLNDPALANYFARKFEWNSTRSVGA GSGSRRSASASLTRTRQPVRGASAHGHTKSRTKSSHRKGHASLEIFTSKRSNPVSKSEKYITVVGTKGRSDAIQLAAPGVHFRAGNKDVFQVNLTGIGKP TKVILENTGTKRTDGWCVSKVVLVKKTDKGTRRYRFSGPVWLSKHHDEMKLKRVLHIDDEDIGSGNGYRIDCYTGDVANAGTDAVATIQLFGSKGQSPMV ELRRSDGQAFQRARVATFTLDDLANLGKLKKLVISHNGHGMASGWFLDKIIVTSLSSNKATVFPCDAWLDRKNGRSKELVARTAAAGAEGMTTFTIRVMT GDRRGAGTDANVQCTLFGEDGESGPHTLNTSRNDFRRGHTDVFAVSSRKIGTLKRLRIWHDNGGAGPAWFLDAVEVVDEASGQTYRFECNRWLAKDEDDG QISRELTCNGDSSWGLKSYKLTIFTGDKRNAGTSANVFCKLVGERGASDNVILENSSKNFQRDRTDIFTVEASDLGSLRHIVLGHDNHGMGAGWYVERFS LEVPSEGKLYNVDVKQWFATDMSDGAIERTFRLDNAETLDIAQRLDWKCTIYTSDVANAGTDANVFMQVYGKKGKTDVVPLKNKSDTFERGQTDELRVQL INVGSLRKLRVWHDNKGMASGWHLDRIVLSRDGEEYIFPCAEWLAVSEGDKEIVRELPATGPNVKKPLQLVEYTVRVATGHARFAGTNADVFVMLTGELG DSGKRALLRSQTNRNKFERGKEDVFTVAAVDLGKLTSVTVGHNNAGTSAGWFLDKIVVLDPRRGEEEEFPCHRWLAVDADDGQIERELVPKMAEHQAAAT TTYIVKIKTGDVRHAGTDANVFVQLFGKTGESTQLKLRNSETYSDAFERNKMDIFKFELLDLGDLSRILVGHDNKGMGAAWFLDYVEVEVPSIRTRWKFP CSRWFSKSQDDGLTEREIYAEKEAGEPMEEDVSAPYLFRFYTSDVAFAGTDANVSVVLYGDEGKTEELVVNNQSDNFERGKADDFKLACKPVGRPSKIRL SAHGGGMSADWHLEKIEVHELGQARIYTFEHNDWLRKGTKAKPFMVELPLRRIETVDDNGREVVEELALDANKRTYRVKVHTGDQKGAGTDANVYVNLHG SLGDSGDRHLKNSLTHTNKFQRKTVDEFDIDAVTLGDINKVKVWHDNAGLGAAWYLEKIEVVDTADDKTYIFPCAQWFAKSMGDGQIARELGVLEEQKPA DFQTKNVGFKYRISVHTSDVKHAGTDANVDIVLYGEKGDTGKIRLAKSETHRDMWERGNCDVFTVSAIELGDLKRVDIMHDGKGVGSGWHLNKVVVDAPQ AGKTWTFMCDAWLDKATDDGTMAKTLYASADAIEEYSAHVPYEIIIKTSDVRNAGTDANVFIDLYGRDQEERDLTAHHEFKDAVKAHFERNLEDRFNVEL PDVGSIYKIRLGHDGKGMSSSWHVASVVVINQRTHERFEFPCDAWLSKDKDDKKLVREFAVGEVKALEGDKVVRRESILSLQEAIYKIHVFTGDIKHAGT DANIYVQIFGDTGDSGEIKLEKSETYRDKFERGHEDIFTHRCLDLGPLRKIKVRSDGKGLMGGDWYLDRVEVHQENDLSEPPVRFVCQDWFKRGKQEGDT LEREITAQVDAAVAMKEATALEDEAARLQRKATTLSRKSTKGSPLRKGTGTTETNAELEEAQRQANIARRKADEAKERAGLAGADDTQLEYKVTVYTGTD TQAGTTANVWLQLFGEKDAPLTPAPPSPSKLSRSGSLFGRRRRASSDAQSVSSSLSAGASGPRETSTGRLQLNNAPADLQSGAKTTFTVTGLDVGELVGL EIGHDDERDKWYLEQVVVEVPKSATHAARRYEFKAGVWLAARADGSTSGSAKGKSKASVRLQPSEIHAGSERILEYTLKVYTASADGAGCTAVPQVQLFG DKHTTEALPLRAGGDILPASVVETQHRVPDLGALLKVRLMVPRGSSWTVEKVEFGRAGQTPITFVGSDGSAVTLGAERLSFDFLPAAAPASSGGKGKRRG STTSSVAVAQQTSYRVYVTTADERGTGTDANVSIILYGAMGDSGEHSLTKSETFDDPFERGNTDVFTLEVPDLGELQRARIWHDGKGMFSSWKLDKIVVV VEATQSRYELPCGQWLSKNKGDKQLTRDLAVASKRVNALTGTYKLEVATDSRAGGGCKGPVRIMLLDAEKNQLPLTLEPPGGEFAPGSVEHLVFDNVMLL GPLTELRIRRKPSGASSRRGAEDDDEDDNEASGASSQSGSAVSPWHLEHIIVKHLQSGQSFVFKGPSKGLSRSRAKLSVHTEATTEEAVAQATKASVAQY EVAVTTGTERGAGTDSNVFVTLFGKNGDSGERALAKSKTFRNMFESGNTDVFDVECQDLGELTKIEVKSDLKGFGAAWQLDKIKVTRTGSQNSWQFKCDQ WFDKKQGAEHTFSVAS*
>LOXHD1_homSap_full dna: each line is an exon ATGATGCCCCAGAAGAAAAGGCGGAGGAAGAAGGACATCGACTTCCTGGCCCTGTACGAAGCGGAGCTGCTGAACTACGCCTCGGAGGACGACGAGGGGGAGCTGGAACACGAGTACTACAAGGCCAGAG TGTATGAAGTGGTCACAGCCACGGGGGATGTTCGCGGTGCAGGGACGGATGCCAATGTCTTCATCACGCTTTTTGGAGAGAATGGGCTCTCTCCCAAGCTCCAGCTCACCAGCAA GAGCAAGTCTGCCTTTGAGAAGGGCAACGTCGATGTGTTCCGGGTGAGAACCAACAATGTGGGCCTCATCTATAAAGTCAG GATTGAGCATGACAACACGGGCTTGAATGCCAGCTGGTACCTGGACCATGTGATTGTGACCGACATGAAGAGGCCTCATCTCCGTTACTACTTCAACTGCAACAACTGGCTGAGCAAGGTGGAAGGTGACCGCCAGTGGTGCCGTGACCTGCTGGCCAGCTTCAACCCCATGGACATGCCCAGAG GTAATAAGTATGAAGTCAAGGTATACACTGGTGATGTAATTGGTGCAGGGACAGATGCTGATGTCTTCATCAATATTTTTGGAGAGTATGGAGACACAG GGGAGCGTAGGCTAGAAAATGAAAAGGACAACTTTGAAAAGGGAGCTGAAGACAGGTTCATCCTGGATGCCCCGGATTTGGGGCAGCTGATGAAGATCAATGTTGGCCACAACAATAAGGGGGGCTCTGCAGGTTGGTTCCTGTCCCAG ATAGTCATTGAAGATATTGGGAACAAAAGAAAATATGACTTCCCCCTTAACCGCTGGCTGGCCTTGGACGAAGACGATGGCAAAATCCAAAGGGATATCTTAGTGGGCGGAGCTGAGACCACAG CTATTACGTATATTGTCACCGTCTTCACTGGGGATGTCCGGGGGGCTGGTACCAAATCCAAAATCTACTTGGTCATGTATGGGGCCAGAGGGAATAAGAACAGTGGGAAAATCTTCCTGGAGGGCGGCGTGTTTGACCGAGGCCGCACGGACATCTTCCACATCGAGCTGGCTGTCCTCCTTAGCCCCCTGAGTCGGGTCTCCGTCGGGCATGGCAATGTGGGTGTCAACAGAGGCTGGTTCTGTGAGAAG GTGGTGATTCTGTGCCCCTTCACTGGTATCCAGCAGACCTTCCCTTGTAGCAACTGGCTGGATGAGAAGAAAGCGGATGGGTTGATTGAGAGGCAGCTCTATGAGATGGTGTCTCTCAGGAAGAAGCGGCTGAAAA AATTCCCTTGGTCCCTGTGGGTCTGGACAACCGACCTAAAGAAAGCTGGTACCAACTCTCCCATCTTCATCCAGATTTATGGGCAGAAGGGGCGGACAGATGAGATTCTCCTGAATCCCAACAACAAGTGGTTCAAACCCGGCATAATCGAGAAGTTTAGG ATTGAGCTCCCGGATCTTGGCAGGTTTTATAAGATTCGAGTATGGCATGATAAAAGGAGTTCTGGTTCTGGATGGCATTTAGAAAGG ATGACCCTGATGAACACTCTGAACAAAGACAAGTACAACTTCAATTGCAACCGCTGGCTGGATGCCAATGAGGATGACAATGAGATAGTGAGGGAAATGACTGCAGAAGGCCCAACAGTGCGCAGGATCATGGGCA TGGCCCGGTACCATGTGACTGTGTGCACAGGTGAACTTGAAGGTGCTGGGACCGATGCCAACGTCTATCTCTGCCTTTTTGGTGATGTGGGGGACACGGGGGAACGGCTGCTCTACAACTGCAGGAATAACACAGACCTGTTTGAAAAGGGCAAT GCTGACGAGTTCACTATCGAGTCTGTCACCATGCGGAATGTGAGGCGGGTGAGGATCAGACACGATGGCAAAGGCTCCGGCAGCGGCTGGTACCTGGACAGAGTGCTGGTGAGAGAGGAGGGGCAGCCTGAGAGCGACAACGTGGAGTTCCCATGTCTCAG GTGGTTGGACAAGGATAAGGATGATGGGCAGCTGGTCCGAGAGTTGCTACCCAGTGACAGCAGCGCGACACTGAAGA ACTTTCGCTATCACATCAGCTTGAAGACTGGGGATGTCTCTGGGGCCAGCACGGATTCTAGAGTCTACATCAAGCTCTATGGGGATAAATCTGACACCATCAAGCAAGTTCTTCTTGTCTCTGACAACAACCTCAAAGACTACTTTGAACGTGGCCGGGTGGATGAGTTCACCCTCGAGACCCTGAACATTGGAAAT ATCAACCGGCTGGTGATTGGGCATGACAGCACTGGCATGCATGCCAGCTGGTTCCTGGGCAGCGTTCAGATCCGTGTGCCCCGTCAAGGCAAGCAGTACACCTTTCCCGCCAACCGCTGGCTGGACAAGAACCAGGCTGACGGGCGCCTGGAGGTGGAGCTGTATCCCAGCGAGGTGGTGGAGATCCAGAAAT TGGTCCACTATGAGGTTGAGATTTGGACAGGAGATGTGGGTGGCGCAGGCACCAGTGCCCGAGTCTACATGCAGATCTATGGAGAGAAAGGCAAGACAGAAGTGCTCTTCCTCTCCAGCCGCTCAAAAGTTTTTGAACGGGCGTCCAAGGACACATTCCAG CTTGAGGCGGCCGACGTGGGCGAGGTCTATAAGCTCCGGCTCGGGCACACGGGCGAGGGCTTTGGGCCCAGCTGGTTCGTGGACACCGTGTGGCTGCGGCACCTGGTGGTGCGGGAGGTGGACCTCACGCCGGAGGAGGAGGCCCGGAAGAAGAAGGAGAAGGACAAGCTGCGGCAGCTGCTCAAGAAGGAGCGGCTGAAGGCCAAGCTGCAGAGGAAGAAGAAGAAGAGGAAGGGCAGCGACGAAGAGGACGAGGGGGAGGAAGAGGAGTCGTCCTCATCAGAGGAGTCCTCGTCAGAGGAGGAGGAGATGGAAGAAGAGGAGGAAGAGGAGGAGTTTGGGCCGGGGATGCAGGAGGTGATTGAGCAGCACAAGTTCGAAGCCCACCGCTGGCTGGCCCGGGGCAAGGAGGACAACGAACTTGTCGTGGAGTTGGTGCCAGCTGGCAAGCCGGGTCCTGAGC GAAACACCTATGAGGTTCAGGTGGTCACGGGGAATGTGCCCAAGGCCGGCACTGATGCTAACGTCTACCTAACCATCTACGGCGAGGAGTATGGAGACACGGGCGAACGACCCCTGAAGAAGTCAGACAAGTCCAACAAATTTGAGCAGGGGCAG ACAGACACCTTCACCATCTATGCCATTGACCTGGGGGCCCTGACCAAGATTCGGATTCGCCACGACAACACAGGCAACAGAGCAGGCTGGTTCCTGGACAGAATAGACATTACTGACATGAACAACGAGATCAC GTACTACTTTCCATGCCAACGTTGGCTGGCAGTGGAGGAAGATGATGGCCAGCTGTCCAGGGAGCTGTTGCCAGTGGATGAGTCCTATGTGCTGCCACAGAGCGAGGAGGGTAGGGGAGGCGGTGACAACAACCCCCTCGACAACCTGGCCCTGGAGCAGAAAG ATAAATCTACCACATTCTCAGTGACCATAAAGACTGGGGTTAAGAAGAATGCGGGCACAGATGCTAATGTCTTCATCACACTCTTTGGCACACAGGATGACACTG GAATGACCCTCCTGAAGTCCTCCAAGACAAACAGCGATAAGTTTGAGAGGGACAGCATTGAAATCTTCACGGTGGAGACGCTGGATCTGGGAGACCTGTGGAAAGTCCGGCTTGGCCATGACAACACAG GCAAGGCCCCAGGCTGGTTTGTAGACTGGGTAGAGGTGGATGCCCCATCTCTTGGGAAGTGCATGACGTTTCCCTGTGGCCGCTGGCTGGCCAAAAACGAAGACGACGGGTCCATCATCAGAGACCTCTTCCATGCAGAGCTTCAGACGAGGCTGTACACACCAT TTGTTCCTTACGAGATCACCCTCTACACCAGTGATGTCTTTGCTGCTGGGACAGATGCCAACATCTTCATCATCATCTATGGCTGCGATGCCGTGTGCACCCAGCAGAAGTATCTGTGTACCAACAAGAGGGAACAGAAGCAGTTCTTTGAGAGGAAGTCTGCCTCCCGCTTCATCGTAGAG TTAGAAGATGTGGGAGAAATCATTGAAAAAATTCGGATTGGCCATAATAACACGGGCATGAATCCTGGGTGGCACTGCTCTCACGTGGACATCCGCAGGCTCCTCCCGGATAAAGAC GGTGCAGAGACCTTGACTTTCCCATGCGATCGGTGGCTTGCCACCTCTGAGGATGACAAAAAGACCATTCGAGAACTGGTTCCATATGACATCTTCACTGAGAAATACATGAAAGATGGGTCCTTACGGCAAGTCTACAAGGAAGTAGAAGAGCCTCTGGACa TTGTGCTGTACTCGGTGCAGATCTTCACAGGGAACATTCCTGGGGCAGGGACGGATGCCAAGGTGTACATCACCATCTATGGAGACCTCGGGGACACTGGGGAGCGATACCTTGGCAAGTCAGAGAACCGGACCAACAAGTTCGAGAGAGGAACG GCTGACACCTTCATCATCGAGGCCGCTGACCTAGGCGTCATCTACAAGATCAAGCTCCGCCATGACAACTCCAAGTGGTGCGCAGACTGGTACGTGGAGAAGGTGGAGATCTGGAATGACACCAACGAGGACGAGTTCCTGTTCCTATGCGGGCGCTGGCTCTCCCTGAAGAAGGAGGATGGGCGACTCGAGAGGCTCTTTTACGAGAAG GAGTACACTGGGGACCGCAGCAGCAACTGCAGCAGCCCTGCTGACTTCTGGGAGATCGCCCTGAGCTCCAAGATGGCCGATGTCGACATCAGCACAGTGACCGGGCCCATGGCTGACTACGTTCAAGAGGGCCCAA TTATTCCCTACTATGTGTCAGTCACCACTGGGAAGCACAAGGACGCGGCCACTGACAGCCGAGCCTTCATCTTTCTCATCGGGGAGGATGATGAACGTAGTAAGCGCATCTGGTTGGACTACCCCCGAGGGAAGAGGGGCTTCAGCCGTGGCTCTGTGGAGGAGTTCTACGTCGCAGGCTTGGATGTGGGCATCATCAAGAAAATAGAG CTGGGCCATGACGGGGCCTCCCCTGAGAGCTGCTGGCTGGTGGAAGAGTTGTGTTTGGCAGTGCCCACCCAGGGCACCAAGTACATGTTGAACTGTAACTGCTGGCTGGCCAAGGACAGAGGCGACGGCATCACCTCCCGTGTCTTCGACCTCTTGGATGCCATGGTGGTGAACATTGGGGTGAAG GTTCTCTATGAAATGACGGTGTGGACAGGGGATGTGGTTGGCGGGGGCACTGACTCCAACATCTTCATGACCCTCTACGGCATCAACGGGAGCACAGAGGAGATGCAGCTGGACAAAAAGAAAGCCAG GTTTGAGCGGGAGCAGAACGACACCTTCATCATGGAGATCCTAGACATTGCTCCATTCACCAAGATGCGGATCCGGATTGATGGCCTGGGCAGTCGGCCGGAGTGGTTCCTGGAGAGG ATCCTACTGAAGAACATGAACACTGGAGACCTGACCATGTTCTACTATGGAGACTGGCTGTCCCAGCGGAAGGGCAAGAAGACCCTGGTGTGTGAAATGTGTGCCGTTATCGATGAGGAAGAAATGATGGAGTGGACCTCCTACACCGTCGCAGTTAAGACCAGCGACATCCTGG GAGCAGGCACTGATGCCAACGTGTTCATCATCATCTTCGGGGAGAACGGGGATAGTGGGACACTGGCCCTGAAGCAGTCGGCAAACTGGAACAAGTTTGAGCGGAACAACACGGACACATTCAACTTCCCTGACATGCTGAGCTTGGGCCACCTCTGCAAGCTGAGGGTCTGGCACGACAACAAAG GGATATTTCCTGGCTGGCATCTGAGCTATGTCGATGTGAAGGACAACTCCCGCGACGAGACCTTCCACTTCCAGTGTGACTGCTGGCTCTCCAAGAGTGAGGGTGACGGGCAGACGGTCCGCGACTTTGCCTGTGCCAACAACAAGATCTGTGATGAGCTGGAAGAGACCA CCTACGAGATCGTCATAGAAACGGGCAACGGAGGCGAAACCAGGGAGAACGTCTGGCTCATCCTGGAGGGCAGGAAGAACCGATCCAAAGAGTTTCTCATGGAAAATTCTTCTAGGCAGCGGGCCTTTAGGAA GGGGACCACAGACACGTTTGAGTTTGACAGCATCTACTTGGGGGACATTGCCTCCCTCTGTGTGGGCCACCTTGCCAGGGAAGACCGGTTTATCCCCAAGAGAGAACTTGCCTGGCATGTCAAGACCATCACCATCACCGAGATGGAGTACGGCAATGT GTACTTCTTTAACTGTGACTGCCTCATCCCCCTCAAGAGGAAGAGGAAGTACTTCAAGGTATTCGAGGTTACCAAGACGACAGAGAGCTTTGCCAGCAAGGTCCAGAGCCTGGTGCCCGTCAAGTACGAAGTCATCGTGACAACAGGCTATGAGCCAGGGGCAGGCACTGATGCCAACGTCTTCGTGACCATCTTTGGGGCCAACGGAGACACAGGCAAGCGGGAGCTGAAGCAGAAAATGCGCAACCTCTTCGAGCGGGGCAGCACAGACCGCTTCTTCCTGGAGACGCTGGAGCTGGGTGAGCTGCGCAAGGTGCGCCTGGAGCACGACAGCAGTGGCTACTGCTCAGGCTGGCTGGTGGAGAAGGTGGAGGTCACCAACACCAGCACCGGCGTGGCCACCATCTTCAACTGTGGCAGGTGGCTGGACAAGAAGCGGGGGGATGGACTCACCTGGAGAGACCTCTTCCCTTCTGTC
Reference PLAT domain sets
>LOXHD1_homSap_PLAT:01 RVYEVVTATGDVRGAGTDANVFITLFGENGLSPKLQLTSKSKSAFEKGNVDVFRVRTNNVGLIYKVRIEHDNTGLNASWYLDHVIVTDMKRPHLRYYFNCNNWLSKVEGDRQWCRD >LOXHD1_homSap_PLAT:02 NKYEVKVYTGDVIGAGTDADVFINIFGEYGDTGERRLENEKDNFEKGAEDRFILDAPDLGQLMKINVGHNNKGGSAGWFLSQIVIEDIGNKRKYDFPLNRWLALDEDDGKIQRDILVGG >LOXHD1_homSap_PLAT:03 ITYIVTVFTGDVRGAGTKSKIYLVMYGARGNKNSGKIFLEGGVFDRGRTDIFHIELAVLLSPLSRVSVGHGNVGVNRGWFCEKVVILCPFTGIQQTFPCSNWLDEKKADGLIER >LOXHD1_homSap_PLAT:04 FPWSLWVWTTDLKKAGTNSPIFIQIYGQKGRTDEILLNPNNKWFKPGIIEKFRIELPDLGRFYKIRVWHDKRSSGSGWHLERMTLMNTLNKDKYNFNCNRWLDANEDDNEIVREM >LOXHD1_homSap_PLAT:05 ARYHVTVCTGELEGAGTDANVYLCLFGDVGDTGERLLYNCRNNTDLFEKGNADEFTIESVTMRNVRRVRIRHDGKGSGSGWYLDRVLVREEGQPESDNVEFPCLRWLDKDKDDGQLVRELLPS >LOXHD1_homSap_PLAT:06 FRYHISLKTGDVSGASTDSRVYIKLYGDKSDTIKQVLLVSDNNLKDYFERGRVDEFTLETLNIGNINRLVIGHDSTGMHASWFLGSVQIRVPRQGKQYTFPANRWLDKNQADGRLEV >LOXHD1_homSap_PLAT:07 VHYEVEIWTGDVGGAGTSARVYMQIYGEKGKTEVLFLSSRSKVFERASKDTFQLEAADVGEVYKLRLGHTGEGFGPSWFVDTVWLRHLVVREVDLTPEEEARKKKEKDKLR >LOXHD1_homSap_PLAT:08 NTYEVQVVTGNVPKAGTDANVYLTIYGEEYGDTGERPLKKSDKSNKFEQGQTDTFTIYAIDLGALTKIRIRHDNTGNRAGWFLDRIDITDMNNEITYYFPCQRWLAVEEDDGQLSRELLPV >LOXHD1_homSap_PLAT:09 TTFSVTIKTGVKKNAGTDANVFITLFGTQDDTGMTLLKSSKTNSDKFERDSIEIFTVETLDLGDLWKVRLGHDNTGKAPGWFVDWVEVDAPSLGKCMTFPCGRWLAKNEDDGSIIRDLFHA >LOXHD1_homSap_PLAT:10 VPYEITLYTSDVFAAGTDANIFIIIYGCDAVCTQQKYLCTNKREQKQFFERKSASRFIVELEDVGEIIEKIRIGHNNTGMNPGWHCSHVDIRRLLPDKDGAETLTFPCDRWLATSEDDKKTIRELVP >LOXHD1_homSap_PLAT:11 VLYSVQIFTGNIPGAGTDAKVYITIYGDLGDTGERYLGKSENRTNKFERGTADTFIIEAADLGVIYKIKLRHDNSKWCADWYVEKVEIWNDTNEDEFLFLCGRWLSLKKEDGRLERLFYEK >LOXHD1_homSap_PLAT:12 IPYYVSVTTGKHKDAATDSRAFIFLIGEDDERSKRIWLDYPRGKRGFSRGSVEEFYVAGLDVGIIKKIELGHDGASPESCWLVEELCLAVPTQGTKYMLNCNCWLAKDRGDGITSRVFD >LOXHD1_homSap_PLAT:13 VLYEMTVWTGDVVGGGTDSNIFMTLYGINGSTEEMQLDKKKARFEREQNDTFIMEILDIAPFTKMRIRIDGLGSRPEWFLERILLKNMNTGDLTMFYYGDWLSQRKGKKTLVC >LOXHD1_homSap_PLAT:14 TSYTVAVKTSDILGAGTDANVFIIIFGENGDSGTLALKQSANWNKFERNNTDTFNFPDMLSLGHLCKLRVWHDNKGIFPGWHLSYVDVKDNSRDETFHFQCDCWLSKSEGDGQTVRDFAC >LOXHD1_homSap_PLAT:15 TTYEIVIETGNGGETRENVWLILEGRKNRSKEFLMENSSRQRAFRKGTTDTFEFDSIYLGDIASLCVGHLAREDRFIPKRELAWHVKTITITEMEYGNVYFFNCDCLIPLKRKRKYFKVF >LOXHD1_homSap_PLAT:16 VKYEVIVTTGYEPGAGTDANVFVTIFGANGDTGKRELKQKMRNLFERGSTDRFFLETLELGELRKVRLEHDSSGYCSGWLVEKVEVTNTSTGVATIFNCGRWLDKKRGDGLTWRDLFPS >PKD1L1_homSap_PLAT chr7: 47780815 polycystin-1L1 GPS-TM-LH2-TM pdb:3CWZ QLYAVVIDTGFRAPARLTSKVYIVLCGDNGLSETKELSCPEKPLFERNSRHTFILSAPAQLGLLRKIRLWHDSRGPSPGWFISHVMVKELHTGQGWFFPAQCWLSAGRHDGRVERELTC >PKD1_homSap_PLAT chr16: 2078712 polycystin 1 GPS-TM-LH2-TM pdb:3CWZ FKYEILVKTGWGRGSGTTAHVGIMLYGVDSRSGHRHLDGDRAFHRNSLDIFRIATPHSLGSVWKIRVWHDNKGLSPAWFLQHVIVRDLQTARSAFFLVNDWLSVETEANGGLVEK >PKD1L2_homSap_PLAT chr16: 79691985 polycystin 1-like 2 signal...GPS-TM-LH2-TM pdb:3CWZ YHYLVTVYTGHRRGAATSSKVTVTLYGLDGEREPHHLADPDTPVFERGAVDAFLLSTLFPLGELRSLRLWHDNSGDRPSWYVSRVLVYDLVMDRKWYFLCNSWLSINVGDCVLDKVFPVA >PKDREJ_homSap_PLAT chr22: 45030224 receptor for egg jelly-like signal...GPS-TM-LH2-TM pdb:3CWZ CYLVTIFTGSRWGSGTRANVFVQLRGTVSTSDVHCLSHPHFTTLYRGSINTFLLTTKSDLGDIHSIRVWHNNEGRSPSWYLSRIKVENLFSRHIWLFICQKWLSVDTTLDRTFHVT >DENND5A_homSap_PLAT chr11: 9116949 RAB6 interacting 1 RUN-LH2-RUN pdb:3CWZ HILIVPSKKLGGSMFTANPWICISGELGETQIMQIPRNVLEMTFECQNLGKLTTVQIGHDNSGLYAKWLVEYVMVRNEITGHTYKFPCGRWLGKGMDDGSLER >DENND5B_homSap_PLAT chr12: 31426424 RAB6 interacting 2 RUN-LH2-RUN pdb:3CWZ RSVIIPIKKLSNAIITSNPWICVSGELGDTGVMQIPKNLLEMTFECQNLGKLTTVQIGHDNSGLLAKWLVDCVMVRNEITGHTYRFPCGRWLGKGIDDGSLER >ALOX5_homSap_PLAT chr10: 45189635 arachidonate 5-lipoxygenase NH2-LH2-lipox pdb:2P0M PSYTVTVATGSQWFAGTDDYIYLSLVGSAGCSEKHLLDKPFYNDFERGAVDSYDVTVDEELGEIQLVRIEKRKYWLNDDWYLKYITLKTPHGDYIEFPCYRWITGDVEVVLRDG >ALOX15_homSap_PLAT chr17: 4480963 arachidonate 15-lipoxygenase NH2-LH2-lipox pdb:2P0M GLYRIRVSTGASLYAGSNNQVQLWLVGQHGEAALGKRLWPARGKETELKVEVPEYLGPLLFVKLRKRHLLKDDAWFCNWISVQGPGAGDEVRFPCYRWVEGNGVLSLPEG >ALOX12_homSap_PLAT chr17: 6840108 arachidonate 12-lipoxygenase NH2-LH2-lipox pdb:2P0M GRYRIRVATGAWLFSGSYNRVQLWLVGTRGEAELELQLRPARGEEEEFDHDVAEDLGLLQFVRLRKHHWLVDDAWFCDRITVQGPGACAEVAFPCYRWVQGEDILSLPEG >ALOX15B_homSap_PLAT chr17 7,888,129 arachidonate 15-lipoxygenase NH2-LH2-lipox pdb:2P0M AEFRVRVSTGEAFGAGTWDKVSVSIVGTRGESPPLPLDNLGKEFTAGAEEDFQVTLPEDVGRVLLLRVHKAPPVLPLLGPLAPDAWFCRWFQLTPPRGGHLLFPCYQWLEGAGTLVLQEG >ALOX12B_homSap_PLAT chr17: 7916679 arachidonate 12R-lipoxygenase NH2-LH2-lipox pdb:2P0M ATYKVRVATGTDLLSGTRDSISLTIVGTQGESHKQLLNHFGRDFATGAVGQYTVQCPQDLGELIIIRLHKERYAFFPKDPWYCNYVQICAPNGRIYHFPAYQWMDGYETLALREA >ALOXE3_homSap_PLAT chr17: 7939943 arachidonate lipoxygenase 3 NH2-LH2-lipox pdb:2P0M AVYRLCVTTGPYLRAGTLDNISVTLVGTCGESPKQRLDRMGRDFAPGSVQKYKVRCTAELGELLLLRVHKERYAFFRKDSWYCSRICVTEPDGSVSHFPCYQWIEGYCTVELRPG >LOXHD1_schMan_PLAT:01 Schistosoma mansoni (flatworm) Bilatera; Platyhelminthes; Trematoda XP_002576380 from contig NS_000200 TLYKVVLTTADVPGAGTSAQIYITLKGEWGSSTRQKLRKEKVPTRNLRFYFYPGSTNTFSVVSPDLGGLHSVFIEHDSLRKSDSWLLESVQVFHPLTKKRYMFMCNHWFSLYKEDGLIARELF >LOXHD1_schMan_PLAT:02 TKYSIVTVTGDQEGSGTNSKVYITIYGRTGITPRIELSQENKSTGKDILCAPFGRGTSTKFIVKAPNVGAITNIRIKQDESGNEPHWFLERVVVTDMSYPQWTYYFHLSCWLSSKYGDGKSCR >LOXHD1_schMan_PLAT:03 TEYKLTFYTSDQKDAGTTGEVYVKLYGEKDSSREIWVNSINQKNRRQPTSYHFTRGSTVEVYLPPCPQLGEITKLKVGHNRAGSSPSWFLNKVIVDDLRMNRVFEFPCYAWITK >LOXHD1_schMan_PLAT:04 IPLEIRLYTGDVPNAGTTAKVYLHQKTTSENKEDVYTTPYIWLEDGNYDRNGVTLFSIDLPVPKFISPLSQLVIGHDNTGHSPSWFFDKVQIYCPLNGVEQTFLYRKWLTSSKPGMKVEQILCEE >LOXHD1_schMan_PLAT:05 IPWEISVKTSPIANSYVTAAVSIIIFGSKDKTMKIQLDRSNLEVSSEWTKVSGNEVVEMFRPNVESKFRIYIKDIGIPCKLRIQHDNKGSNPNWHLQEIVLTNLRTHEQYEFYCNRWLSTREDDGSILREIPAK >LOXHD1_schMan_PLAT:06 YIIQIFTGNKPNSGTNANIFINIFGEKGDCGERWLGRSVNRNSELFQQNQMDEFVIEAVQLGSINKICIGHEERSPGYGWYLAKIVLTIKENPKYKLTFECYRWFDVGEDDGQIVRELFA >LOXHD1_schMan_PLAT:07 IAYNVTVLTGTCRNAGTVANVFVHLYGLQGESKDMQLKHKETEITKFEAGKSEEFILACGKLGEIKKIKIWHDSIAPHSGWFLDEVVVIDYFLGRRYVFHVNRWLAKNEADGLISLDLQPT >LOXHD1_schMan_PLAT:08 SYEVIVKTGGRKYAGTDSHVYITMFGINSESKEYHLANSKTHNNKFEQNHEDLFNLNAVGLGSLKKIRVRHTNTGIAPGWFLDYILIRESIEQKKMEYFFPCYQWLSATRLDGLVIREL >LOXHD1_schMan_PLAT:09 TIYHVRIHTGSQSNISSDANVQIQLYGDKDFTGNIRLYKAYYGKEDILVNKFQAGQIANFIVKAINIGNIKKIRIEHDTAVGSARWFVERVEVEARKLGLLWKFECNR >LOXHD1_schMan_PLAT:10 TNYQIKTFTSDISGAETTSSVYIQLYGNDSLPSSIRRLHQNNDSEQRFQRNKIDTFYVELEELQEPFSKLRIWHNDKGSSTDWHLNKVEIRKIKTNQYVFVTYIFPCNKWLSRNMDQAALERELIPS >LOXHD1_schMan_PLAT:11 NTYEVRITTGDKAYAGTDASVYITLFGENGDSGERKLTKSLTHRNKFERGQTDVFQLEIVDLGKINKVRIRHDNSGVNPSWYLSTIEVFNISKSQSVHGLKDQPNETHHLVKYIFNCEAWLSTEHDEKVLDR >LOXHD1_schMan_PLAT:12 IPYLITVITGSDRKAGTPGPVWISCVDKDKMNSEKFILCDCYNRTMLKRGTTRYFRFAGVKLNDLTEIQVGNDAPESPSMGWYIQSLYLSFPTIGKMYLFDCKEWLSTNRGNKK >LOXHD1_schMan_PLAT:13 ITYHLKITTANVKRAGTDCSINLQIFGTNGVTNCYILEKTSNRFAQGITDNISLEMEDVGKLLKMRIGHDNQGKNKHWNLSCVEVTVANTNQLYRFVYDDWLSLTYGKRKSLWADLPAM >LOXHD1_schMan_PLAT:14 IFVKTGNMPASSTDANVYVQLFGEYGDSGEILLKQTVSNQKPFQNNSIDHFKIPSILKLGNLARCRIWHDNKGSSPNWYCEWLEVKEVLIPGEKNLACNWKFAFNKWLSVSDDNKQLLRDAPCS >LOXHD1_schMan_PLAT:15 VYEVVIETGNLKDSGTTCDAWIILEGKHGRSPKLELVNQVGNPILQINQMNTFQLPSFPLGDLETIRLGIQERNINKQTNPNDIQSQKWFCEKVSIKDPVSKRTYIFTIHQWLS >LOXHD1_schMan_PLAT:16 VMYKVSIYTGTKGCANTDANIFITMFSTTPGLNSGRIALKRENNNLFDRKQLDEFYVESIDLESIQRIIIEHDNTGVSPDWYLDKVLITNQTNNQIHLFQCYQWISKKKGDCRLWKELLVS
Reference PLAT:07 domains
The seventh PLAT domain (human numbering) is unique among PLAT domains in having a large expansion in the middle of the domain. It is a unique signature of the LOXHD1 orthology class that would distinguish it from other poly-PLAT domain concatenates of independent origin. That issue is completely hypothetical because no other protein in any vertebrate genome contains more than a single PLAT domain. Indeed, the LOXHD1 gene is single-copy in all species that retain it (including teleost fish). Thus even with very ancient eukaryotic divergencs, the corresponding genes can be taken as straightforward human orthologs.
These other proteins consist of 10 genes, namely four polycystins (PKD1L2, PKDREJ, PKD1L1, PKD1), two RAB6 (DENND5B, DENND5A), and five arachidonate 5-lipoxygenases (ALOX5, ALOXE3, ALOX15, ALOX12B, ALOX12) plus three bogus misannotations (AK310228, FLJ00285 and NPIP) found in many database compilations. In these other genes the PLAT domain occurs in the context of GPS-TM-PLAT-TM (polycystins), RUN-LH2-RUN (RAB6), and N-terminally (lipoxygenases).
Presumably a loop out of the beta barrel, it consists of an extremely high AG content at the dna level and encodes charged residues, primarily lysine K and glutamate E residues. These are asymmetrically distributed with the lysines N-terminal and the glutamates distal. The region is called the KE loop of PLAT:07 in this article.
Despite minimal net charge, this region may not be able to interact with (hydrophobic) membranes. Yet the split PLAT domain retains the potential to form a standard beta barrel. The region is quite variable even within mammals and must be hand-annotated to ensure accuracy (because its quasi-repetitive nature wreaks havoc with alignment). This section uses comparative genomics within eukaryotes to answer the following questions:
- when did the high GA section arise in phylogenetic time? [early eukaryotes]
- what is its source? [internal expansion, replication slippage]
- did it arise from intron or retroposon retention? [species lacking the repeat also have a single exon]
- can the boundaries of 7th PLAT domain be better located? [conservation on both sides of the KE region allow definition of the PLAT domain segments]
- is the 44-species alignment accurate in this region? [corrections are needed when longer than human, eg platypus]
- does the KE domain disrupts PLAT function? [evidently not: the split domain is highly conserved and can form a normal beta barrel]
- is it a mammalian innovation or an unwanted intrusion? [this ancient expansion has variable expansion and contraction in different clades]
homSap LEAADVGEVYKLRLGHTGEGFGPSWFVDTVWLRHLVVREVDLTPEEEARKKKEKDKLRQLLKKERLKAKLQRKKKKRKGSDEEDEGEEEESSSSEESSSEEEEMEEEEEEEE FGPGMQEVIEQHKFEAHRWLARGKEDNELVVELVPAGKPGPE 1 ornAna LEATDVGEIYKVRLGHSGEGFGSGWFIESLVLKRLVLKEVEPNPEEEKRKAKERERAREQRRKERLKAKQQRKKKKKMKKSSDDEDSEAEDSEEEEGSSEEESSSSSSEEEVEEEFGPGIKEVIDVYKFEAHRWLARDEDDKELIVELEPANRPGPE 1 galGal VEAADVGKIYKIRIGHDGKGIGDGWFLESVTLKRLATKMDETD KKKKKKKKKSEEEEEEEE TKVEEVMDVYTFVAHRWLAKDEGDKELVVELVPDGESELE 1 taeGut REAADVGKIYKIRIGHDGTGIGDGWFLESVTLKRLATKTEGSD KKKKKKKKSEEEETKE EEGMDVYTFVAHRWLAKDEGDKELVVELVPDGESDLE 1 anoCar VEAVDVGKVYKIRIGHDGKGFGDGWFLDSVVVKKLPTKVP KKKKKKKKKKTPEEEEAEE GPGIMEVYNFTPCRWLASDEEDKELVVELVPDEGSELE 1 oryLat IEALDVGKIYKIRIYHDGSGIGDGWFLETVDIKRLTMALVQVEV KKEEAPKKDKKKDKKKKKKEEEEVEIIEE MQEVVETFTFTCNRWLARDEEDGEIVVELLTEENEDLE 1 gasAcu IEAKDVGKIFKIRIGHDGSGIGSGWFLETVDVKRLILALVPKEK KKEDKKKKKKKKEDVDEEGGEE MQEVVLTYSFPCSRWLAGGEEDGELVVELLPDDAKELE 1 takRub IEAKDVGKIFKIRIGHDGLGIGSGWFLEKVYVKHLIMALVPREN KKDDKKKKKKKKKDKEDEEEVGGEE MQEVVVTYHFPCSRWLASGEDDDDLVVELLPEDAEELE 1 petMar VEAMDVGKVVKLRVGHDNSGMGSGWFLDSIVIRRLRQSSPHRPQPVDA EEDEDEEDDEEAEDED VQTYTFPCKRWLARDEDDGEIVRELLPQDCAEME 1 sacKow IEAADVAMLTKIRIGHDNSGRSAGWYLERVIIERFPPKRKM KRKRSGTPRRRGEYDEEDYDD IPETNVVNFVCNRWFAKDEEDHQIVRELLPTDEEALKGH 1 triAdh VECEDVGKLRKLRIGHDSAGMGSAWFLDKVYVRRLPPKSGKKSKET DEREEETAKKDADEPEKLDE NNYLFVANRWLSKEEGDRQTVIEISPVGVDGALA 1 monBre LACKPVGRPSKIRLSAHGGGMSADWHLEKIEVHELGQAR IYTFEHNDWLRKGTKAKPFMVELPLRRIETVDDN 1 >PLAT:07_homSap Homo sapiens (human) 2 LVHYEVEIWTGDVGGAGTSARVYMQIYGEKGKTEVLFLSSRSKVFERASKDTFQ 0 0 LEAADVGEVYKLRLGHTGEGFGPSWFVDTVWLRHLVVREVDLTPEEEARKKKEKDKLRQLLKKERLKAKLQRKKKKRKGSDEEDEGEEEESSSSEESSSEEEEMEEEEEEEE FGPGMQEVIEQHKFEAHRWLARGKEDNELVVELVPAGKPGPE 1 >PLAT:07_monDom Monodelphis domestica (opossum) 2 LIHYEVEIVTGDMGNASTSARVYMQIYGEEGKTEVLFLKSRSKVFQRGNTDTFQ 0 0 LQEAEVTPEEEARKKKEMDKLRQLMRKERLKAKEEQKRKRKLAKGSDVEDEEESSSEESSEESEEETEIEEEDEE FQEVVDVYRFHAGRWLATDEEDKDLDMELAPSGRSGPQR 1 >PLAT:07_ornAna Ornithorhynchus anatinus (platypus) 2 LIHYEVEIVTGDMGYAGTNARVYMQIYGELGKTEVLHLTSRTNVFEQGATDTFQ 0 0 LEATDVGEIYKVRLGHSGEGFGSGWFIESLVLKRLVLKEVEPNPEEEKRKAKERERAREQRRKERLKAKQQRKKKKKMKKSSDDEDSEAEDSEEEEGSSEEESSSSSSEEEVEEEFGPGIKEVIDVYKFEAHRWLARDEDDKELIVELEPANRPGPE 1 >PLAT:07_galGal Gallus gallus (chicken) 2 LINYEVSVVTGDVRAAGTNAKVFMQIYGETGKTELIILENRSNNFERGATDIFE 0 0 VEAADVGKIYKIRIGHDGKGIGDGWFLESVTLKRLATKMDETDKKKKKKKKKSEEEEEEEETKVEEVMDVYTFVAHRWLAKDEGDKELVVELVPDGESELE 1 >PLAT:07_taeGut Taeniopygia guttata (finch) 2 LINYEVSVVTGDVRAAGTNAKVFMQIYGETGKTELIILENRSNNFERGATDIFK 0 0 REAADVGKIYKIRIGHDGTGIGDGWFLESVTLKRLATKTEGSDKKKKKKKKSEEEETKEEEGMDVYTFVAHRWLAKDEGDKELVVELVPDGESDLE 1 >PLAT:07_anoCar Anolis carolinensis (lizard) 2 LINYEVCVATGDVRNAGTNANVFMQIYGELGKTELLILKNRSNNYERGASETFR 0 0 VEAVDVGKVYKIRIGHDGKGFGDGWFLDSVVVKKLPTKVPKKKKKKKKKKTPEEEEAEEGPGIMEVYNFTPCRWLASDEEDKELVVELVPDEGSELE 1 >PLAT:07_oryLat Oryzias latipes (medaka) 2 VINYEIAVVTGDVRAGGTNASVFCQIYGEEGKTEVLNLKSRSNNFERGTTEIFK 0 0 IEALDVGKIYKIRIYHDGSGIGDGWFLETVDIKRLTMALVQVEVKKEEAPKKDKKKDKKKKKKEEEEVEIIEEMQEVVETFTFTCNRWLARDEEDGEIVVELLTEENEDLE 1 >PLAT:07_gasAcu Gasterosteus aculeatus (stickleback) 2 VINYEVTVVTGDVTFAGTNARVIIQIYGDKGKTEVIILESRSNNYERNTTEIFK 0 0 IEAKDVGKIFKIRIGHDGSGIGSGWFLETVDVKRLILALVPKEKKKEDKKKKKKKKEDVDEEGGEEMQEVVLTYSFPCSRWLAGGEEDGELVVELLPDDAKELE 1 >PLAT:07_takRub Takifugu rubripes (fugu) 2 VINYEVTVVTGDVTFAGTNAKVFVQIYGDKGKTEVIMLESRSNNYERNAMEIFK 0 0 IEAKDVGKIFKIRIGHDGLGIGSGWFLEKVYVKHLIMALVPRENKKDDKKKKKKKKKDKEDEEEVGGEEMQEVVVTYHFPCSRWLASGEDDDDLVVELLPEDAEELE 1 >PLAT:07_petMar Petromyzon marinus (lamprey) 2 GINYQVSVHTGNVRSAGTNANVFIQLYGDMGKTEVHNLRNRSDNFERASNDIFK 0 0 VEAMDVGKVVKLRVGHDNSGMGSGWFLDSIVIRRLRQSSPHRPQPVDAEEDEDEEDDEEAEDEDSEEVQTYTFPCKRWLARDEDDGEIVRELLPQDCAEME 1 >PLAT:07_sacKow Saccoglossus kowalevskii (acornworm) Bilateria|Deuterostomia|Hemichordata 2 GIPYEITVITGDKSGAGTDAQVFIRMYGLKGKTDEFILNNRTDNFERGMTDKFK 0 0 IEAADVAMLTKIRIGHDNSGRSAGWYLERVIIERFPPKRKMKRKRSGTPRRRGEYDEEDYDDIPETNVVNFVCNRWFAKDEEDHQIVRELLPTDEEALKGH 1 >PLAT:07_strPur Strongylocentrotus purpuratus (sea_urchin) 2 GIPYEITVITGDKSGAGTDANVFLTMYGDDGAKTEEFSLRNRTDNFEKNMTDKFK 0 >PLAT:07_triAdh Trichoplax adhaerens (placozoan) Eukaryota|Metazoa|Placozoa 2 KIPYQISVHTGDIRHAGTDSNVFAVIYGENGKTEELKLRNKSDNFERGQVDVFK 0 0 VECEDVGKLRKLRIGHDSAGMGSAWFLDK 00 VYVRRLPPKSGKKSKETDEREEETAKKDADEPEKLDENNYLFVANRWLSKEEGDRQTVIEISPVGVDGALA 1 >PLAT:07_monBre Monosiga brevicollis (choanoflagellate) Eukaryota|Choanoflagellida 2 PMEEDVSAPYLFRFYTSDVAFAGTDANVSVVLYGDEGKTEELVVNNQSDNFERGKADDFK 0 0 LACKPVGRPSKIRLSAHGGGMSADWHLEKIEVHELGQARIYTFEHNDWLRKGTKAKPFMVELPLRRIETVDDNGREVVEELALDANKRTYR 1