Bison: mitochondrial genomics: Difference between revisions
Tomemerald (talk | contribs) |
Tomemerald (talk | contribs) |
||
| Line 22: | Line 22: | ||
Bos frontalis (gayal) 0 29 0 20433524 18244904 17560527 | Bos frontalis (gayal) 0 29 0 20433524 18244904 17560527 | ||
Bos taurus (cattle) 168 500 1 19603063 19484124 19393053 19393048 19393045 20347826 | Bos taurus (cattle) 168 500 1 19603063 19484124 19393053 19393048 19393045 20347826 | ||
Bos primigenius (auroch) 1 17 0 18199470 19456314 | Bos primigenius (auroch) 1 17 0 18199470 19456314 20346116 | ||
Bos indicus (zebu) 3 463 0 12648092 19436739 19770222 20597883 18467841 12399392 | Bos indicus (zebu) 3 463 0 12648092 19436739 19770222 20597883 18467841 12399392 | ||
Revision as of 16:04, 8 December 2010
Introduction to bison and yak conservation genomics
(to be continued)
Phylogeny: bison and yak are sister groups
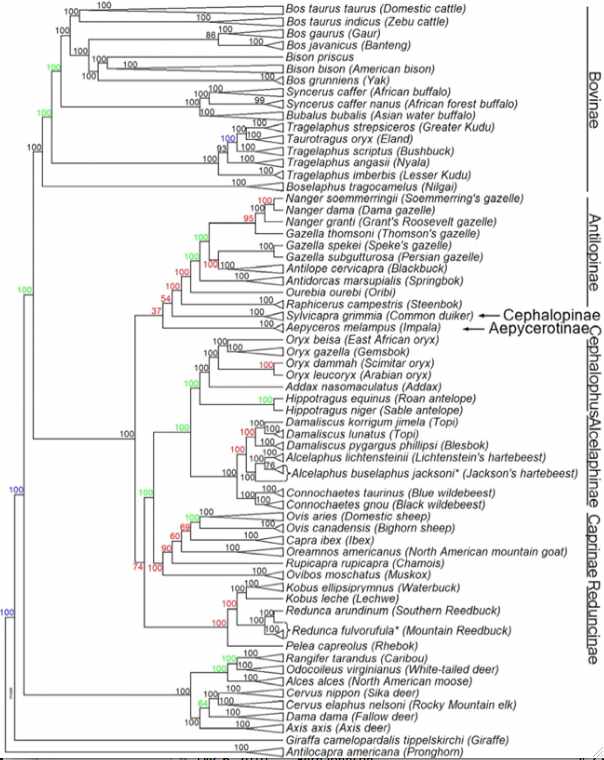
The evolutionary history of cetartiodactyls has taken decades to sort out. The position of whales, once controversial, has been settled (sister, together with hippopotamus, of Ruminantia), as has the non-intuitive branching order of pigs and lamas (Camelidae are basal).
Within pecoran ruminants, the issue arose not so much from conflicts between fossils morphology and molecular trees but rather rapid radiation of species (tree polytomy) that has only been resolved (we hope) with the advent of the bovine SNP bead chip. JE Decker et al evaluated 52,356 sites across the nuclear genome not only of cows but throughout the artiodactyls. The resulting tree (antelopes,(giraffes,(deer,((gazelles,sheep),bovinae)))) is critical to understanding the evolution of mitochondrial proteins.
Notice that Linnaean taxonomy requires substantial revision according to this tree -- genera such as Bos, Tragelaphus and Gazella are inconsistent with it. This could be remedied for bison by either placing them in Bos or putting yak, gaur and banteng in the genus Bison. Here the position of gaur and bantang (gayal and kouprey Bos sauveli were not considered) has less bootstrap support than other nodes and has long been contentious.
There may not be any simplistic resolution because of male introgression as illustrated in european bison (wisent) and zebu cattle. The speciation process can be far messier than indicated by bifurcating tree nodes. For example, subsequent to divergence, wandering bulls from one population can mix into another. While this does not affect mitochondrial lineages, it does result in continuing introgression into nuclear genome. Since domestication, cattle have been crossed with both yak and bison, indicating a full speciation barrier to hybridization still does not exist after several million years. Further, polymorphic alleles all represented in an ancestral population may sort out differently in descendant lineages.
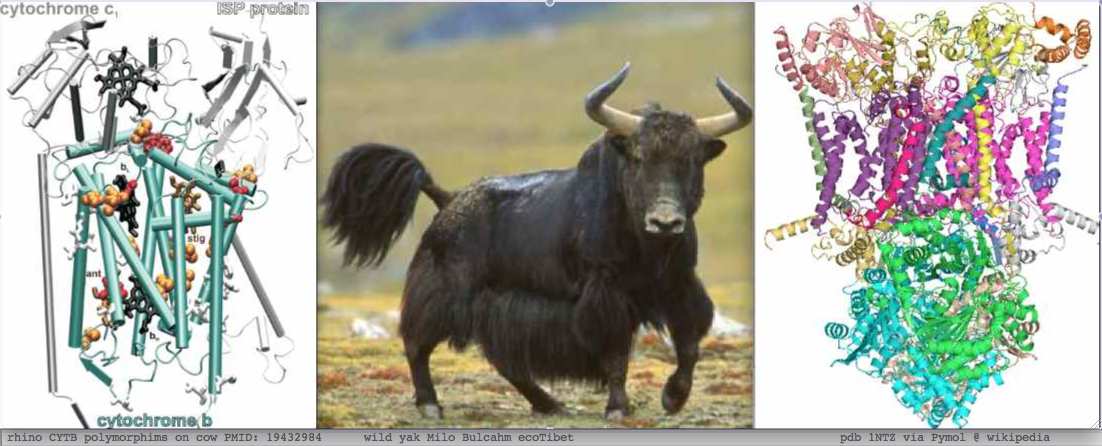
The data situation is otherwise very favorable with over 214 complete mammalian mitochondrial genomes and high multiplicities for individual species such as cow, bison, and yak. Certain genes such as CYTB have considerable additional data from targeted studies.
Species Common Mito CYTB Nuc PubMed Bos sauveli (kouprey) 0 6 0 15522811 16439342 17848372 Bos frontalis (mithun) 0 25 0 20331596 Bos gaurus (gaur) 0 14 0 19436739 19777782 19367625 17986322 Bos javanicus (bantang) 2 39 0 18937038 18937038 17614913 16922247 12522420 Bos frontalis (gayal) 0 29 0 20433524 18244904 17560527 Bos taurus (cattle) 168 500 1 19603063 19484124 19393053 19393048 19393045 20347826 Bos primigenius (auroch) 1 17 0 18199470 19456314 20346116 Bos indicus (zebu) 3 463 0 12648092 19436739 19770222 20597883 18467841 12399392 Bos grunniens (yak) 72 53 * 19917041 17257194 18439980 16942892 12137333 Bison bison (plains bison) 33 7 0 20870040 20637048 19414501 Bison athabascae (woods bison) 2 3 0 20808568 18191321 Bison bonasus (wisent eurobison) 3 6 0 14739241 19623210 17177698 15125253 14703870 Bison priscus (steppe bison) 0 0 0 15567864 20409351 20212118 18653730 18199470 Bubalus bubalis (waterbuffalo) 1 342 0 17459014 15621663 11212504 19140976 19462514 19207933 Syncerus caffer (cape buffalo) 0 10 0 10603253 9126673 9987926 17313588 17459014 14715223 Tragelaphus (kudo, eland) 0 204 0 10222159 7723053
Yak nuclear genome sequencing is in progress at Beijing Genomics Institute. Other cetartiodactyl genomes include Camelus bactrianus and Ovis aries (also in progress) and Camelus dromedarius and Pantholops hodgsonii (completed but not released). Other genomes said to be underway include Bubalus bubalis, Addax nasomaculatus, Muntiacus muntjak, Hippopotamus amphibius, and Balaena mysticetus. Cow, pig, sheep, and vicuna genomes have long been available for blast search.
These additional genomes would allow fossil nuclear numts to contribute to understanding of mitochondrial gene evolution, making the mitochondrial proteome of ancestral species such as Leptobos (last common ancestor to cows and bison) easy to work out.
Note too that the mitochondrial genome, although not targeted, gets sequenced to very high multiplicity as a byproduct. To date, such projects have produced single mitochondrial genomes. This however must be wrong in view of the prevalence of heteroplasmy. In other words, most species will host a population of slightly different mitochondrial genomes. Thus these genome projects are a golden opportunity to characterize mitochondrial genome diversity within single species.
Interpreting bison CYTB variation
Bison mitochondrial genomes became well-represented at GenBank with the 1 Dec 10 release by the Derr group of 31 complete genomes from 6 herds including two woods bison (Bison bison athabascae) from the non-admixed Elk Island herd (along with various cow-bison hybrid and cow breed genomes). The cow-bison hybrids represent crossing of a bison male with a domestic cow (or rather a continuous line of female descent from such a cross) and so have strictly cow mitochondrial dna, not relevent to this section. The haplotype of all hybrids studied (from an unnamed private ranch in Montana, presumably Turner's Flying D) cluster with cow haplotype cHap32.
Bison accession numbers: GU946976 GU946977 GU946978 GU946979 GU946980 GU946981 GU946982 GU946983 GU946984 GU946985 GU946986 GU946987 GU946988 GU946989 GU946990 GU946991 GU946992 GU946993 GU946994 GU946995 GU946996 GU946997 GU946998 GU946999 GU947000 GU947001 GU947002 GU947003 GU947004 GU947005 GU947006
The CYTB sequences retrieved from these genomic entries (they are not yet in the database used by blastp) show haplotype notation. The 15 previously existing bison sequences at GenBank (some just fragments are also provided. Older fragmentary sequences are demonstrably error-prone and will be used here only as support -- never as sole source -- of a polymorphism. Redundancy introduced via non-standard SwissProt (UniProt) entries also has to be manually removed -- the Swiss did no sequencing on their own, simply deriving protein sequences from existing GenBank entries. This leaves 5 older complete sequences for Bison bison and 4 fragments, 2 attributed to Bison bonasus and 1 fossil dna sequence from Bos primigenius to serve as outgroup (rather than an inbred domestic cow).
Here it is necessary to pick a terminology. This must accommodate NCBI taxonomy -- regardless of its correctness -- because otherwise blastp searches cannot be restricted by taxon. Note although bison are definitely sistered with yak to the exclusion of all other extant species, that creates problems because yak has been put in the genus Bos. Many relic wild cattle have no english language common name but rather that of a local language. Terminology table must show synonyms to allow PubMed and google searches -- especially important in a fast-moving field to locate preprints and conference proceedings. The table below does not attempt to implicitly resolve any scientific issue; it simply states preferred terminology at this site along with synonyms in common use.
(sequence annotation to be continued) YP_002791041 Bison bison Q9T9C1 Bison bison YP_003587278 Bison bonasus ACE76876 Bos primigenius YP_003541096 Bos primigenius O20998 Bison bonasus ADQ12704 Bison bonasus GenBank error? 61 ......T.....ETTAEF.. 120 AAL85955 Bison bison AAL85956 Bison bonasus ADM87433 Bison bison AAW28803 Bison bison AAW28804 Bison bison AAW28802 Bison bonasus AAN28295 Bison bison CAA76013 Bison bonasus
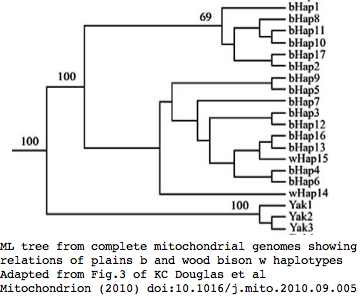
Sequences are color clustered according to the phylogenetic tree above. bHap1 is not shown. Note the woods bison cannot be resolved from the plains bison even though the Elk Island woods bison are a relic herd that did not mix with 7,000 plains bison imported from the Flathead Reservation in Montana up to Canada's Wood Buffalo National Park in the 1920's.
>CYTB_bisBis.GU946988 bHap8 plains bison b973 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946994 bHap11 plains bison b1031 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946990 bHap10 plains bison b985 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU947000 bHap10 plains bison bFN5 Niobrara MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946991 bHap10 plains bison b1005 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU947004 bHap17 plains bison bYNP1586 Yellowstone NP MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946976 bHap2 plains bison b790 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946977 bHap2 plains bison b853 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946978 bHap2 plains bison b854 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946981 bHap2 plains bison b880 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946983 bHap2 plains bison b925 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946984 bHap2 plains bison b929 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946993 bHap2 plains bison b1029 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946995 bHap2 plains bison b1050 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946996 bHap2 plains bison b1051 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU947001 bHap2 plains bison bNBR1 National Bison Range MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946986 bHap2 plains bison b959 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHAGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946989 bHap9 plains bison b979 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946997 bHap9 plains bison b1091 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946982 bHap5 plains bison b897 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946982 bHap5 plains bison b897 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946987 bHap7 plains bison b961 Montana MTSLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946979 bHap3 plains bison b855 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946992 bHap3 plains bison b1018 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946992 bHap3 plains bison b1018 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946998 bHap12 plains bison b1191 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU947003 bHap16 plains bison bTSBH1005 Texas State Bison Herd MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946999 bHap13 plains bison b1428 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU947002 bHap13 plains bison bTSBH1001 Texas State Bison Herd MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisAth.GU947005 wHap15 woods bison wEI1 Elk Island MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946980 bHap4 plains bison b877 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisBis.GU946985 bHap6 plains bison b935 Montana MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKW >CYTB_bisAth.GU947006 wHap14 woods bison wEI14 Elk Island MTNLRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGMCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTMMATAFMGYVLPWGQMSF WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIIMAIAMVHLLFLHETGSNNPTGISSDMDKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQMASIMYFLLILVLMPTAGTIENKLLKWDEFINITION
Interpreting CYTB variation in yak
Yaks are the closest living sister species to bison. Although 15,000 wild yaks still persist, they have been subject to very similar pressures to those experienced by bison: bottlenecks, population fragmentation, introgression from long domesticated yaks and hybridization with cattle. Adaptations specific to mitochondria may exist as yak live at altitudes exceeding 4000 meters with average annual temperatures in rearing areas –8°C, with animals surviving winter temperatures of –40°C.
Because yaks provide the immediate outgroup for bison genetics (and vice versa), their parallel mitochondrial proteomics are investigated in depth here. This further enables reconstruction of mitochondrial proteins of their last common ancestor (after consideration of lineage sorting) and correct placement of Pleistocene genomic sequences.
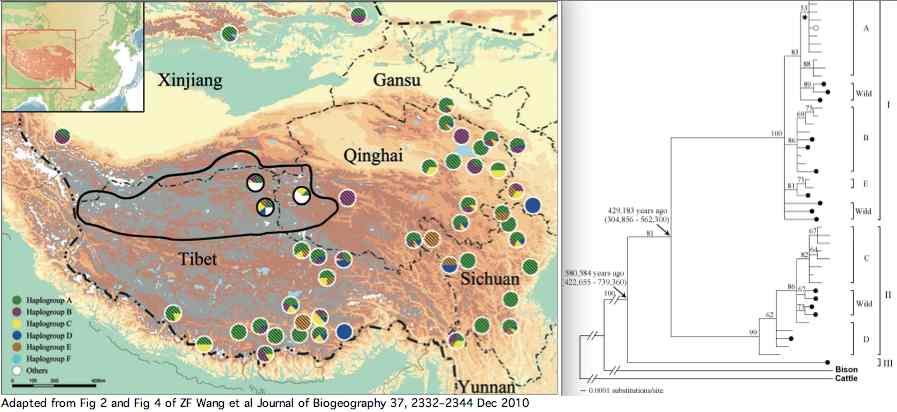
Data availability for yaks was greatly improved by a Dec 2010 paper by Zhaofeng Wang et al. that investigated yak phylogeographical structure and demographic history on the Qinghai-Tibetan Plateau. Complete mitochondrial genomes were determined for 48 domesticated and 21 wild yaks. The three lineages shown in article supplemental established diverged at 420 kyr and 580 kyr in accordance with extended but temporary allopatric migration barriers created by two large plateau glaciations.
The wild yaks are found in all three branches of the tree (solid circles in figure). Their entries at GenBank are distinguished by a W (for wild) prefix, eg isolate W77 GQ464266. There is potential for confusion here because NCBI taxonomy uses Bos grunniens mutus for wild yak, yet the subspecies concept is contradicted by the mixed distribution of wild and domestic yaks in the mitochondrial tree. Related taxa such as Bos mutus (Przewalski, 1883), Bos mutus grunniens, and Poephagus mutus also conflict with the facts. Yak and bison -- diverging at 2.5 million years -- need to reside in the same genus.
The primary focus here are protein polymorphisms in wild yak because domesticated animals may exhibit inbreeding issues and other evolutionary artifacts due to their estrangement from darwinian selection. Consequently it is important to track which GenBank entries reference wild yaks.
Bos grunniens mutus has three GenBank entries relevant to cytochrome b: proteins AAX53006 and AY955226 both containing unique V195A, I348F mutations in an otherwise wildtype background and CAA76015, an older fragmentary wildtype sequence not considered further here. The first two animals add samples to the large, remote Xinjiang province but remain unpublished (Liu,Q Wu,M Li,Y) despite the 27 Mar 2005 submission date at GenBank. (A number of D-loop sequences submitted for this taxon on 19 Jan 2009 by 27-MAR-2005 by Ma,ZJ also remain unpublished.)
The Myanmar/Bhutan mithun sequence BAJ05329 attributed to Bos grunniens at GenBank has 12 differences to wild yak but is 100% identical to 94 Bos indicus entries, ie it is a hybrid and its mitochondrial genome is irrelevant here. Such GenBank errors are all but impossible to correct.
The 21 new genome accessions of wild yak are GQ464266, GQ464265, GQ464264, GQ464263, GQ464262, GQ464261, GQ464260, GQ464259, GQ464258, GQ464257, GQ464256, GQ464255, GQ464254, GQ464253, GQ464252, GQ464251, GQ464250, GQ464249, GQ464248, GQ464247, GQ464246. These were not mapped to the published tree.
In terms of protein accessions (which will be shown at NCBI blastp output), these are ACU81659, ACU81646, ACU81633, ACU81620, ACU81607, ACU81594, ACU81581, ACU81568, ACU81555, ACU81542, ACU81529, ACU81516, ACU81503, ACU81490, ACU81477, ACU81464, ACU81451, ACU81438, ACU81425, ACU81412, ACU81399 to which AAX53006 and AY955226 can be added.
Of these, 16 fall in the main reference sequence group (wildtype) but 5 wild plateau yaks exhibit polymorphisms that cannot be attributed to domestication. As noted, two additional wild yaks from extreme NW China have additional double mutations but no associated PubMed publication nor tissue source indicated. As either change alone would inactivate an essential enzyme, these represent either heteroplasmic oddities or sequence error (to be pursued as other proteins are considered). The remaining sequences were derived from muscle and skin dna.
There is no overlap between wild yak polymorphism sites and the five of domestic yak. Alleles occurring in full length sequences are analyzed further below.
The summary table of yak CYTB amino acid polymorphisms below arises from alignment of 5000 full-length mammalian cytochrome b orthologs. Magenta indicates a deleterious change at an invariant position,red a deleterious mutation at a naturally polymorphic site, green a possibly acceptable change but of restricted distribution and fitness, and blue a near-neutral substitution. Gray is reserved for probable sequencing error. It can be seen that the smallish yak population sampled (72 animals) already contains 5 deleterious alleles in CYTB which represents only 10% of the amino acids of the mitochondrial proteome.
In summary, out of 70 individual yaks, 10 are carrying deleterious mutations at five sites. That seems like an extraordinary number for a central enzyme in energy metabolism for which it is difficult to envision compensation by another gene. Restricting to the 21 wild yaks, 3 have deleterious polymorphism and 1 has a marginal change. Overall 1 in 7 animals is affected just in this one gene. However CYTB is but one of 13 encoded by the mitochondrial genome -- what sort of genetic burden are yaks carrying overall?
1 ACU81568 A017T wild yak isolate W50 GQ464259 2 ACU81399 I192T wild yak isolate W02 GQ464246 ACU81633 I192T wild yak isolate W75 GQ464264 3 ACU81555 D214N wild yak isolate W40 GQ464258 4 AAX53006 V195A I348F mutus isolate Xinjiang01 unpublished Liu,Q Wu,M Li,Y AAX53007 V195A I348F mutus isolate Xinjiang02 unpublished Liu,Q Wu,M Li,Y 5 ACU81529 V329M wild yak isolate W1313 GQ464256 6 ABI15999 V039I A067T domestic yak fragment PUBMED:17257194 Poephagus 7 ABI16000 V039I A067T domestic yak fragment PUBMED:17257194 Poephagus ACU82153 A084T domestic yak isolate HY5 8 ACU82101 V098L domestic yak isolate HY1 9 AAU89116 I118T domestic yak =SP:Q5Y4Q0 PUBMED:16942892 ACU81711 I118T domestic yak isolate HZ3 ACU81737 I118T domestic yak isolate MQ1 AAS93096 I118T domestic yak fragment PUBMED:17257194 AAS93099 I118T domestic yak fragment PUBMED:17257194
Although the mitochondria encodes the usual 20 amino acids, only a subset of chemically similar residues ever appear at a given position in a given protein -- its reduced alphabet. This subset describes the evolutionarily acceptable substitutions that do not significantly disrupt protein functionality. Discovery of this reduced alphabet can be achieved with greater precision the higher the number of available species and individual sequences multiplicities. For mitochondrial proteins, that sensitivity is 1 in 10,000 (0.01% occurrence frequency) for a given amino acid, much better than even the much-studied human nuclear genome.
Interpretive certainty is never attained without experimentation (yeast is a surprisingly informative model system) but improves up to a point with more sequence data. Here it is important to check whether less common substitutions have persisted over evolutionary time in a phylogenetically coherent manner (ie a sub-clade) or are novel adaptations perhaps in conjunction with a co-evolving residue at another site (or another protein, perhaps nuclear-encoded). After these considerations, the remaining rare changes are mostly deleterious (or sequencing error) but rarely adaptive. Polymorphism significance can be pursued at the xray structural level for only 3 of the 13 mitochondrial proteins (CYTB, COX2, COX1) and even this is complicated in the case of CYTB by its oligomeric association with 3 nuclear encoded proteins.
Aligning CTYB from the 72 complete yak mitochondrial genomes available on 1 Dec 10 shows variation at just 9 sites along the protein (ie 9 nsSNPs). These are quickly found when the web alignment tool retains input sequence order, displays residues identical to the top sequence as dots, gaps fragmentary data correctly, and allows a wide display permitting effective cross-species comparisons.
Yak and bison -- despite being sister species -- share variation only at one site, position 98. Here yak is exclusively valine with the exception of a single deleterious occurrence (see below) of leucine, whereas bison have a mix of valine and alanine (which otherwise is very rare at this position in mammals), ie the ancestral residue was valine. Thus no lineage sorting occurred at any amino acid position in CYTB at the time these two species diverged at 2.5 myr. Lineage sorting however may be important in the overall evolution of the Bovini: 53 ancient polymorphisms (at the dna level) are said to have persisted since Bos and Bison diverged from Bubalus 5–8 million years ago.
The changes can also be displayed in context by coloring the appropriate residues in a reference sequence relative to a composite sequence consolidating all the polymorphisms from distinct animals (no one animal has more than two of the 9; V195A + I348F occurs in two animals). The composite sequence is quite useful in comparing polymorphism sites across species as explained in the annotation tricks section.
>CYTB_bosGruR Bos grunniens cytochrome b ref seq taken as gi|147744503
MTNIRKSHPLMKIVNNAFIDLPAPSNISSWWNFGSLLGVCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHANGASMFFICLYMHVGRGLYYGSYTFLETWNIGVILLLTVMATAFMGYVLPWGQMSF
WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIITAIAMVHLLFLHETGSNNPTGISSDADKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI
LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLVADLLTLTWIGGQPVEHPYIIIGQLASIMYFLLILVLMPTAGTIENKLLKW
>CYTB_bosGruP Bos grunniens composite polymorphisms: A017T A084T V098L I188T I192T V195A D214N V329M I348F
MTNIRKSHPLMKIVNNTFIDLPAPSNISSWWNFGSLLGVCLILQILTGLFLAMHYTSDTTTAFSSVAHICRDVNYGWIIRYMHTNGASMFFICLYMHLGRGLYYGSYTFLETWNIGVTLLLTVMATAFMGYVLPWGQMSF
WGATVITNLLSAIPYIGTNLVEWIWGGFSVDKATLTRFFAFHFILPFIITATAMAHLLFLHETGSNNPTGISSNADKIPFHPYYTIKDILGALLLILALMLLVLFTPDLLGDPDNYTPANPLNTPPHIKPEWYFLFAYAI
LRSIPNKLGGVLALAFSILILALIPLLHTSKQRSMIFRPLSQCLFWTLMADLLTLTWIGGQPVEHPYFIIGQLASIMYFLLILVLMPTAGTIENKLLKW
wild dom dom dom wild dom wild wild dom
A017T A084T V098L I118T I192T V195A D214N V329M I348F
4018S 4994A 4522V 4309I 4353L 4528V 4429D 4610V 4232I
927A* 3T 430I 667S 505M 427I 512N 188T 651V
46T 1P 34M 14I 94I* 25T 43E 133A 63T
3L 1V 11A 1T 31T 4G 8S 44I 45M
3M 1L 3F 4M 2Y 22M 4N
1F 1N 2V 1A 1H 2G 2F
1P 1A 1E 1A
1S
A017T: At position 98, the mammalian reduced alphabet consists primarily of serine but with yak alanine also well represented at 18%. Threonine occurs in 46 sequences so cannot be sequence error or serious mutation. Bulk seems to be the main criterion at this site rather than polarity -- threonine though polar is bulkier residue than serine or alanine. To determine whether it has arisen multiple times or just in one clade, the phylogenetic distribution of the 46 occurrences needs consideration.
It can be seen from the graphic at left that A017T has arisen multiple times with no common denominator (such as high elevation lifestyle) but -- with the exception of monotremes -- never in a deep stem ancestor. That is, A017T occurs here and there but only in recently speciated clades. This suggests that while not lethal, over time it gets replaced by more adaptive serine or alanine.
A017T
4018 S
927 A yak do not have the most common amino acid at position 17
46 T
3 L
3 M
1 F
1 P
A084T: At position 84, alanine is strictly invariant. Thus threonine is an unmistakable deleterious mutation in domestic yak.
A084T
4994 A
3 T
1 P
1 V
V098L: At position 98, the reduced alphabet consists of valine 90% of the time regardless of mammalian clade with the similar (branched chain aliphatic) isoleucine having substantial dispersed representation at nearly 9%. The 430 species in which it occurs are scattered incoherently within mammal clades, meaning that it has arisen independently many times. V098I may be slightly suboptimal as there is an evident bias (at some level) against equal occurrence. It likely co-exists with valine in most non-bottlenecked populations of mammals, observed if enough individuals of a given species are sequenced.
However leucine, the seemingly similar third aliphatic residue, occurs one once despite being but a single base change transition away from the dominant residue. Were leucine a near-neutral substitution, its incidence would be vastly higher. Thus the change V098L reported for yak represents either a deleterious mutation or an unprecedented adaptation (eg to high altitude) or sequencing error in GenBank entry ACU82101. The same can be said for the more overtly radical change V098N in lemur AAS00156. The 34 methionines occur sporadically in the phylogenetic tree suggesting they are sub-adaptive and blink out over time. Indeed, canine spongiform leukoencephalomyelopathy is attributed to V98M. Dog CYTB is 89% identical to that of yak and numbering corresponds.
V098L 4522 V 430 I 34 M 11 A bison 1 L yak 1 N lemur
I118T: At position 118, the reduced alphabet consists predominantly of ILV with some A and M, a very common occurrence proteomewide. TSF are all deleterious mutations in domestic yak.
I118T
2597 I
1843 L
404 V
87 A
61 M
6 T (all yak)
1 S
1 F
I192T: At position 192 of wild yak, the dominant residue is leucine instead of the yak ancestral value isoleucine, which is disfavored relative to methionine, ie isoleucine is a mild polymorphism in its own right but the associated taxonomy shows it narrowly restricted to 83 sequences in Bos, Bison, and separately in 5 Kobus (waterbucks), too persistent to be dysfunctional and indeed a candidate for adaptive. However change to polar threonine is seen in 31 nominal species but after removal of redundancy, only in two species of pocket mice. Thus the yak change is deleterious.
I192T
4353 L
505 M
94 I
31 T
3 F
2 V
1 A
1 S
V195A: This allele occurs together with I348F in two wild yaks from a remote region in NW China. Despite sequence submission, no article has appeared in the three subsequent years. It can be seen from the reduced alphabet frequencies that this is a severe mutation (as is I348F) so taken together likely sequence just error. No further analysis will be done here until such time as the polymorphisms are confirmed.
V195A
4528 V
427 I
25 T
4 G
4 M
1 A
D214N: This polymorphism of wild yak is seen quite widely, in some 10% of mammals. The 223 taxa with D214N are mostly confined to laurasiatheres and glires but are not a hallmark of these clades. Nor do the species with asparagine have any common lifestyle denominator. Asparagine is an acceptable variation for aspartate at this site if perhaps not optimal.
D214N
4429 D
512 N
43 E
8 S
4 X
2 Y
1 H
V329M: This allele occurs in wild yak. Methionine is not a radical substitution in terms of physical/chemical properties and similar additional amino acids appear at low levels, even though valine occurs in a huge majority of species. Methionine occurs in 17 other species phylogenetically scattered species include Bos javanicus, Ovis, Budorcas, Naemorhedus, Mus, Rattus, bats and sloth. Thus it is likely suboptimal but not significantly deleterious.
V329M
4610 V
188 T
133 A
44 I
22 M
2 G
1 E
I348F: This allele occurs together with V195A in two wild yaks from a remote region in NW China. Despite sequence submission, no article has appeared in the three subsequent years. In can be seen from the reduced alphabet frequencies that this is a severe mutation but more likely sequence error, as is V195A.
I348F
4232 I348F
651 V
63 T
45 M
4 N
2 I348F
1 A
Human CYTB polymorphism and disease
Polymorphisms and pathogenic mutations disease for human CYTB have been very helpfully compiled by mtDB and MitoMap, with other mammals at OMIA. Fortunately, numbering systems carry over without change to bison and yak since no indels occur in this gene within mammals.
A poor tradition in mitochondrial research allows amino acid changes to be described just by a single nucleotide coordinate relative to the Cambridge Reference Sequence, NC_012920. That requires the user to have a numbered translation via the mitochondrial genetic code showing in-frame amino acids; however the change from the protein perspective (eg V98T) is often conveniently displayed at Uniprot. Coordinates for all mitochondrial features are tabulated here; CYTB extends from position 14747-15887.
Given over 7000 complete human mitochondrial genomes and a high mutation rate, some human polymorphic sites will inevitably overlap with yak and bison alleles. Thus any information about associated human disease at the 16 known disease sites might be transferable. However many rare and obviously dysfunctional alleles were collected for population haplotype mapping and no disease information was collected.
Annotation transfer is vastly complicated by heteroplasmy, experimentalist inability to establish the heritability of the allele, and differences in tissues used to obtain dna for sequencing and so neglects the possibly compensatory effect of changes elsewhere in this gene (or nuclear genes that interact with it), a substantial issue in a protein like cytochrome b where 10% of the residues between bovids and human are non-identical. At such sites (eg H214Y human, D214N yak), transfer of phenotypic information is dubious.
brown in the human allele table below indicates human polymorphisms corresponding to an allele of concern in yak or bison. In two significant cases -- both in domestic yak -- the initial and final residue of human are identical to that of yak, namely A084T* and I118T*. Both are predicted to be deleterious in both human and yak. Unfortunately no clinical information was collected on the human side and the health status of the yaks is unknown (eg level of exercise intolerance).
However even A084T (a strongly invariant site in all mammals) was evidently not early-lethal for its adult human carrier (dna samples are collected from adult volunteers whose health status is not recorded). Here the vast and still unsettled complexities of mitochondrial genomics may come into play:
- a single mitochondrion may up to 10 replicated copies of its genome which need not be identical
- cells can carry thousands of mitochondria inherited erratically during embryogenesis and later stem cells
- dna samples, not being collected from germline cells, may represent non-heritable somatic mutations in restricted descendent cells of the tissue sampled
- disease onset is often in late adulthood due to the nature of mitochondrial replication and dispersal to daughter cells and so may not be applicable to shorter-lived species
Symptoms of severe heteroplasmic mitochondrial disorders frequently do not appear until adulthood because many cell divisions and much time is required for a cell to receive enough mitochondria containing the mutant alleles to cause symptoms. An example of this phenomenon is Leber optic atrophy (LHON). Affected individuals may not experience vision difficulties until they have reached adulthood. Another example is MERRF syndrome (Myoclonic Epilepsy with Ragged Red Fibers). Heteroplasmy here explains the variation in severity of the disease among siblings. The incidence of heteroplasmy in human mtDNA is unknown, as the number of individuals who have been subjected to mtDNA testing for reasons other than the diagnosis of mitochondrial disorders is small."
The oft-observed disease Leber Hereditary Optic Neuropathy (LHON) is genetically heterogeneous, arising from mutations in other mitochondrial genes (R340H in ND4, A52T in ND1 and M64V in ND6, subunits of complex I of the oxidative phosphorylation chain in mitochondria) as well as from CYTB variants A29T and secondarily D171N and V356M.
tRNA disruptions in bison were analyzed by Douglas et al. Here it is known the human disease MERRF disrupts mitochondrial tRNA-Lys in 80% of cases and so biosynthesis of mitochondrial proteins essential for oxidative phosphorylation. It too is genetically heterogeneous as tRNAs for leucine, histidine, serine and phenylalanine can be affected in other individuals.
Mitochondrial diseases arise frequently: 1 in 4000 individuals is at risk of developing a mitochondrial disease sometime in their lifetime. Half of those affected are children who show symptoms before age five, and approximately 80% of them will die before age 20. The mortality rate is roughly that of cancer... The mutation rate of the mitochondrial genome is 10–20 times greater than of nuclear DNA, and mtDNA is more prone to oxidative damage than is nuclear DNA. Mutations in human mtDNA cause premature aging, severe neuromuscular pathologies and maternally inherited metabolic diseases, and influence apoptosis.
human yak A084T A084T* seen twice in Japanese population I098V V098L I118V I118T I118T I118T* seen once in Japan and once in India H214Y D214N A329T V329M T2A S56A I117V D171N I211T G251S M316T A354T T2I S56L I118V D171G T212A E251D Y325H V356M M4V T61A I118T S172N T212I Y256H A329T V356A M4T T70A L121F P173S H214Y T257I A330T T360A R5G Y75C A122T T174A T219A L258P A330V T360M I7T I78V T123A F181L T219I A259T I334V T368A N8S I78T A125T I184V I226V N260D T336A T368I N15S L82F E136D L185S A229T V284I I338V I369V H16R A84T F140L I189V L230F V291A P342S I369T F18L G86S L149M A190T L233V S297P V343M I372V I19M C93Y I153T A190V F235L I300T V343A M376V A29T I98V Y155H A191T L236I I304T S344G A380T A39T G101S I156V A191D S238P I306V S344N A380V A39V Y109H I156T A193T S238F I306T Y345F ---- I42V E111K T158A T194A T241A M309V T348I ---- I42T T112A D159N T194V T241M M309T I349V ---- F50L W113R I164V F199L T243A S310P I349T ---- F50L I115T G167S I211V F245L M316V V353M ---- Of known disease mutations, only V98M corresponds to a bison allele: A29T LHON Leber hereditary optic neuropathy G34S mitochondrial myopathy; sporadic S35P exercice intolerance V98M* dog leukoencephalomyelopathy S151P exercise intolerance G166E hyperthrophic cardiomyopathy D171N secondary LHON G231D 16026996 mouse G251D CMIH G251S obesity N255H cardiomyopathy Y278C multisystem disorder G290D exercise intolerance S297P neonatal polyvisceral failure G339E mitochondrial myopathy V356M secondary LHON
- See abstracts for all 16 disease sites
- Adaptive rates of evolution in all 13 genes from an alignment of 214 mammalian mitochondrial genomes
- Adaptive evolution of the mammalian mitochondrial genome
- A series of 12 papers on mitochondrial dna inheritance issues
Cytochrome b mutations in Leber hereditary optic neuropathy CYTB:D171N CYTB:V356M ND5:A458T New mutations were discovered in the apocytochrome b gene in Leber hereditary optic neuropathy probands who did not harbor either of the two known Complex I mutations (positions 3,460 and 11,778). A mutation at position 15,257 was found in eight independent probands which changed a highly conserved D to N, was not found in controls, and appears to be pathogenetically significant. The 15,257 mutation occurred in association with a known synergistic mutation at position 13,708 in 7/8 probands (ie ND5 A458T) and in association with a new apocytochrome b mutation at position 15,812 (ie V356M) in 4/8 probands. Mutations in Complex III genes may be involved in Leber hereditary optic neuropathy and multiple, simultaneous mutations occur frequently.
Mazunin IO (2010) Mitochondrial genome and human mitochondrial diseases. Molecular Biology 44(5) Today there are described more than 400 point mutations and more than hundred of structural rearrangements of mitochondrial DNA associated with characteristic neuromuscular and other mitochondrial syndromes, from lethal in the neonatal period of life to the disease with late onset. The defects of oxidative phosphorylation are the main reasons of mitochondrial disease development. Phenotypic diversity and phenomenon of heteroplasmy are the hallmark of mitochondrial human diseases. It is necessary to assess the amount of mutant mtDNA accurately, since the level of heteroplasmy largely determines the phenotypic manifestation. In spite of tremendous progress in mitochondrial biology since the cause-and-effect relations between mtDNA mutation and the human diseases was established over 20 years ago, there is still no cure for mitochondrial diseases.
Pathogenic mitochondrial DNA mutations in protein-coding genes
Lee-Jun C. Wong PhD Muscle Nerve, 2007
More than 200 disease-related mitochondrial DNA (mtDNA) point mutations have been reported in the Mitomap (http://www.mitomap.org) database. These mutations can be divided into two groups: mutations affecting mitochondrial protein synthesis, including mutations in tRNA and rRNA genes; and mutations in protein-encoding genes (mRNAs). This review focuses on mutations in mitochondrial genes that encode proteins. These mutations are involved in a broad spectrum of human diseases, including a variety of multisystem disorders as well as more tissue-specific diseases such as isolated myopathy and Leber hereditary optic neuropathy (LHON). Because the mitochondrial genome contains a large number of apparently neutral polymorphisms that have little pathogenic significance, along with secondary homoplasmic mutations that do not have primary disease-causing effect, the pathogenic role of all newly discovered mutations must be rigorously established. A scoring system has been applied to evaluate the pathogenicity of the mutations in mtDNA protein-encoding genes and to review the predominant clinical features and the molecular characteristics of mutations in each mtDNA-encoded respiratory chain complex.
S297P homoplasmic in all tissues tested, undetectable in mother PMID: 19563916
Eur J Hum Genet. 2004 Mar;12(3):220-4.
The deleterious G15498A mutation in mitochondrial DNA-encoded cytochrome b may remain clinically silent in homoplasmic carriers.
We report on a patient with severe growth retardation and IgF1 deficiency, in which a mitochondrial abnormality was suspected. An isolated mitochondrial respiratory chain complex III deficiency was found in blood lymphocytes and skin fibroblasts. Sequence analysis of the cytochrome b, which is the only mitochondrial DNA-encoded subunit of complex III, revealed a homoplasmic G15498A mutation, resulting in the substitution of a highly conserved amino acid (glycine 251 into an aspartic acid). The mutation was found to be homoplasmic in all tissues examined from the mother and her brother (lymphocytes, fibroblasts, hair roots and buccal cells). Complex III deficiency was also demonstrated in these cells. Nevertheless, the mother and the brother were asymptomatic. This mutation had been considered as a cardiomyopathy-generating mutation in a previously reported case, and its pathogenicity has been demonstrated recently in yeast. However, it seems not to fulfil the classical criteria for pathogenicity of a mitochondrial DNA mutation, especially the heteroplasmic status, and to be clinically silent, albeit present, in nonaffected relatives. We suggest that other factors are contributing to the clinical variability expression of the G15498A mtDNA mutation.
Mitochondrial DNA mutations cause disease in >1 in 5000 of the population and approximately 1 in 200 of the population are asymptomatic carriers of a pathogenic mtDNA mutation. Many patients with these pathogenic mtDNA mutations present with a progressive, disabling neurological syndrome that leads to major disability and premature death. There is currently no effective treatment for mitochondrial disorders, placing great emphasis on preventing the transmission of these diseases. An empiric approach can be used to guide genetic counseling for common mtDNA mutations, but many families transmit rare or unique molecular defects. There is therefore a pressing need to develop techniques to prevent transmission based on a solid understanding of the biological mechanisms. Several recent studies have cast new light on the genetics and cell biology of mtDNA inheritance, but these studies have also raised new controversies.
Recommendations for yak conservation genomics
When yak and bison mitochondrial genomes are sequenced and polymorphisms are reported to GenBank, what exactly does that mean? Presumably it reflects an overwhelmingly dominant value of whatever heteroplasmy existed in the tissue sample used to sequence the dna.
The key bison study used white blood cells as dna source, rather than muscle/skin in the yak data. One might imagine this fraction of whole blood is quite heterogeneous in terms of stem cell origin -- five different, diverse leukocyte types exist -- but these all derive from a single hematopoietic stem cell type in bone marrow. Consequently, no other cell types of these bison was sampled. Thus we do not know whether the observed polymorphisms there are heritable (apart from those observed in multiple animals).
Mitochondrial disease in yak is not straightforward to analyze:
"Heteroplasmy is the presence of a mixture of more than one type of an organellar genome within a cell or individual. It is a factor for the severity of mitochondrial diseases, since every eukaryotic cell contains many hundreds of mitochondria with hundreds of copies of mtDNA, it is possible and indeed very frequent for mutations to affect only some of the copies, while the remaining ones are unaffected.
Bovine oocyte mitochondrial issues have been studied for decades, with novel explanations how germline mutations might propagate:
GS Michaels 1982: Restriction endonuclease analysis and direct nucleotide sequencing of bovine mitochondrial DNA have revealed a high apparent rate of sequence divergence between maternally related individuals. Oocytes had 260,000 dna genomic copies per cell, whereas primary bovine tissue culture cells contained only 2,600 copies. These experiments demonstrate directly the amplification of mitochondrial DNA in mammalian oocytes and are consistent with models which could generate mitochondrial DNA polymorphisms by unequal amplification of mitochondrial genomes within an animal.
For yak, since oocytes are not used, the observed polymorphisms are not necessarily heritable even for female individuals (male mitochondria are not passed on). However in the case of yak polymorphisms I118T (domestic) and I192T (wild), multiple individuals (5, 2 respectively) sampled carried the same rare change, strongly implying (unless these are mutational hotspots) that these are entrenched in the germline and so inherited. Oocyte heteroplasmy however is also heritable so wildtype may still persist. The other polymorphisms may be mere somatic mutations that attained abundance in the sampled tissue but are still complemented by residual wildtype. This would have to be pursued in additional tissues or more definitively by sequencing offspring, perhaps not feasible in wild yak.
In summary, even deleterious polymorphisms may have limited effects, depending on stem cell origin and compensation by the wildtype component of heteroplasmy. On the other hand, should a bad alleles exert a negative dominant effect even as the minority allele in the mitochondria in which it resides (eg tainting oligomeric proteins), it would still have a deleterious phenotype even though it never comes to 100% frequency in any particular cell type. Somatic mutations in bison and yak may have limited impacts if onset of disease is delayed to late adulthood as in human. For conservation genomics, we are primarily concerned with heritable mitochondrial mutations, though enhanced levels of somatic mutations (due say to a faulty POLG dna polymerase) are also a concern.
In domestic yak, animals bearing I118T should not be encouraged to reproduce. To be on the safe side, higher frequencies of the other deleterious alleles are also undesirable, even though not quite proven to be heritable.
In wild yak, I192T is the primary cause of concern. It should be avoided if captive breeding comes into play. A017T, D214N, and V329M are not deleterious mutations but rather natural and possibly adaptive parts of yak diversity whose continuation should be encouraged.
These preliminary recommendations are based solely on CYTB. Since only rare recombination occurs in mitochondria (that could bring good alleles on different genes together) and no paternal contribution can dilute out undesirable heteroplasmy, it is unclear how these recommendations can be implemented, much less reconciled from those emerging from independent considerations of the other 12 mitochondrial genes.
Nuclear proteins that raise mitochondrial mutation rates
The genetic stability of mtDNA in every mammal (indeed every eukaryote) depends critically on the accuracy of dna replication. The consequences of any mutation in this machinery would be greatly amplified (like the broomsticks in the Sorcerer's Apprentice) by subsequent somatic errors created in replicating mitochondrial genomes. It is essential to consider these genes given the apparent elevated rate of mitochondrial polymorphism reported for bison and yak.
The nuclear encoded, mitochondrially functioning dna polymerase POLG on chr 15, the catalytic subunit The catalytic subunit (dna polymerase itself, 3’-5’ exonuclease for proofreading, 5’deoxyribosephosphate lyase for base excision repair), deserves special mention in regards to the extraordinary observed rates of yak and bison coding polymorphisms. Some 90 distinct [human disease alleles are known along the 1239 residue protein, causing progressive external ophthalmoplegia, sensory and ataxic neuropathy, Alpers syndrome, and male infertility (see PEOA1, SANDO, AHS, MNGIE at OMIM). POLG also contains a polyglutamine tract near its N-terminus of length 13 in human that may be subject to polymorphic replication slippage.
POLG is accompanied by an accessory dimer of POLG2. Now receiving considerable attention, two mitochondrial disease alleles have been found, G416A and G451E (causing adPEO). A helicase (PEO1 or twinkle) causing an adult-onset progressive external ophthalmoplegia PEO and topoisomerase TOP1MT are other nuclear encoded proteins critical to mitochondrial dna replication. The latter binds a specific site in the D loop control region. These too have been implicated in rare mitochondrial diseases.
These enzymes, especially POLG, needs extensive sequencing in bison and yak (indeed every once-bottlenecked endangered species). That might done economically on a population scale with whole-exome chips rather than sequencing whole genomes. The POLG gene itself is difficult to study in isolation, being comprised of 23 exons spread out over 18490 bp.
No sequencing of yak or bison POLG has been done yet but that of cow, sheep and pig etc are readily retrieved from their respective genome projects. The Bos taurus POLG protein is 90% identical to human; it has not been specifically studied.
Numts: excluding mitochondrial pseudogenes
Mitochondrial research has been plagued by numt pseudogene alleles mistakenly obtained from the nuclear genome by primer cross-over. Here rna transcribed from mitochondrial genes somehow exits the mitochondria, enters the cell nucleus, gets reverse-transcribed into dna, and then gets heritably integrated into the nuclear junk genome (where it generally is not transcribed and rapidly accrues the mutation pattern of a pseudogene), sometimes becoming fixed across the entire population and even diagnostic of it.
This seemingly implausible sequence of events is surprisingly common. Counts for any species with assembled genome can quickly be conducted by Blat at the UCSC genome browser, though very old events would be missed. Querying cow genome for CYTB nuclear pseudogenes shows 19 nuclear genome matches to cytochrome b, ranging from quite strong to barely significant.
The best match occurs on cow chr28:34924178-34924995. Not quite full length 3', it contains 7 internal stop codons, 52 addition missense mutations (that characteristically do not follow site conservation patterns), and various indels and frameshifts. Not particularly recent, its date of formation could be bracketed by examining sheep and pig genomes for an orthologous numt at syntenic location (+psCYTB +SFTPD (or CGN1, bovine conglutinin).
It's not clear pig contains the orthologous pseudogene at chr14:34281258-34281963 because this feature is not immediately syntenic to porcine SFTPD at chr14:85511174-85522800. If so, the most recent CYTB pseudogene in cow predates the divergence of cow and pig. It then will be found in both yak and bison genomes unless lost through large-scale deletion. Note pig has a much more recent CYTB at chr2:104178282-104179415.
The sheep genome is not currently in a satisfactory state of assembly. This is far more likely than pig to contain a demonstrably syntenic CYTB pseudogene. No sheep pseudogenes are posted at GenBank nr nor locatable by tblastn against wgs or hgts databases. Although 31 CYTB pseudogenes from 11 pecoran species are available, these species all lack genome projects. However upon blastn of the cow chr28 feature, Kobus kob (AF052940) and Capra hircus (GU120393) have very strong matches.
Recent numt pseudogenes can capture ancestral values that prevailed in the mitochondria at the time of formation. Unlike bone fossils, this dna has steadily accrued changes up to the present, but the benefit is nuclear pseudogenes evolve up to 12 times slower than the mitochondrial parental gene. Thus it might represent an atypical heteroplasmic allele existing at that time, be affected by lineage sorting, or reflect a parallel nuclear mutation and so not really settle the issue of ancestral value. A joint tree (1, 2) containing both mitochondrial CYTBs and nuclear pseudogenes (as outgroups) considered in chamois (Rupicapra) has many complexities because genes evolve so differantly in the two compartments (for example the pseudogene might have arisen from a heteroplasmic variant that existed at the time).
The yak study specifically considered whether numts could explain divergent, low-frequency mtDNA haplotypes, but ruled out all but the very most recent on the basis of the separate confirmatory D-loop haplotype phylogeny and great similarity to other haplotypes without unusual sequence features.
Bioinformatic tips and tricks
New sequencing technologies have greatly affected the amount of mammalian mitochondrial genomic data available at GenBank. Five years ago, it was acceptable to publish population-level D loop sequences accompanied by a few fragmentary coding reads; today, a publication might offer 60-70 entire mitochondrial genomes. This favors evolutionary study of mitochondrial proteins over comparative genomics of nuclear genome products because the latter is still restricted to around 50 species (Dec 2010) almost all incompletely sequenced.
Many long-standing issues such as introgression, historic bottlenecks, population mixing, accrual of deleterious coding variants, hard polytomies, and lineage sorting during speciation can now be approached and resolved, especially with the increasing sequencing of end-Pleistocene frozen dna. This may allow more enlightened management of endangered species such as bison where populations reached rock bottom -- recovering numbers is not enough if genomic integrity is still at risk.
However, the flood of data raises significant issues in extraction of significant information: it is not instructive to align the tens of thousands of sequences available for each of 13 mitochondrial proteins -- that give a an intractable array of 3789 amino acids by 12500 sequences, enough to fill 20 x 100 = 2000 screens on the largest possible computer monitor. That data must be distilled down somehow to take-away information.
This section explains a practical desktop protocol for extracting the 'reduced phylogenetic alphabet' at each residue of the mitochondrial proteome. The method depends heavily on current capabilities of Blastp at NCBI and so may not be completely stable to changes made there over time.
First note that tBlastn cannot be used against the nr or wgs nucleotide databases at NCBI (or with Blat at UCSC) since the significantly different genetic code of mammalian mitochondria is no longer supported as a parameter option. Other oddities involve missing terminal nucleotides that are added before translation. However mitochondrial dna is usually translated sensibly at GenBank protein entries.
The vertebrate mitochondrial code:
TTT F Phe TCT S Ser TAT Y Tyr TGT C Cys
TTC F Phe TCC S Ser TAC Y Tyr TGC C Cys
TTA L Leu TCA S Ser TAA * Ter TGA W Trp
TTG L Leu TCG S Ser TAG * Ter TGG W Trp
CTT L Leu CCT P Pro CAT H His CGT R Arg
CTC L Leu CCC P Pro CAC H His CGC R Arg
CTA L Leu CCA P Pro CAA Q Gln CGA R Arg
CTG L Leu CCG P Pro CAG Q Gln CGG R Arg
ATT I Ile ACT T Thr AAT N Asn AGT S Ser
ATC I Ile i ACC T Thr AAC N Asn AGC S Ser
ATA M Met i ACA T Thr AAA K Lys AGA * Ter Bos can use ATA as initiation codon
ATG M Met i ACG T Thr AAG K Lys AGG * Ter
GTT V Val GCT A Ala GAT D Asp GGT G Gly
GTC V Val GCC A Ala GAC D Asp GGC G Gly
GTA V Val GCA A Ala GAA E Glu GGA G Gly
GTG V Val i GCG A Ala GAG E Glu GGG G Gly
AAs = FFLLSSSSYY**CCWWLLLLPPPPHHQQRRRRIIMMTTTTNNKKSS**VVVVAAAADDEEGGGG
Start = --------------------------------MMMM---------------M------------
Base1 = TTTTTTTTTTTTTTTTCCCCCCCCCCCCCCCCAAAAAAAAAAAAAAAAGGGGGGGGGGGGGGGG
Base2 = TTTTCCCCAAAAGGGGTTTTCCCCAAAAGGGGTTTTCCCCAAAAGGGGTTTTCCCCAAAAGGGG
Base3 = TCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAG
Blastp output at NCBI now has a very useful feature: clustering of identical individual sequences into single alignments, display of multiplicities, with all the accessions visible with an extra click. The only exception involves double-counting of SwissProt entries which, since SwissProt conducts no sequencing, always arise from another entry.
After collecting high resolution amino acid frequencies at a given site, it is necessary to determine the phylogenetic distribution of each variant (in practice just those of moderate occurrence). That is now very convenient to do provided the associated accessions have been saved:
Simply paste the blastp match list of protein accessions having the chosen amino acid variant into the Entrez text query box. Never mind if it only returns 20 out of your 157 input sequences -- it hasn't forgotten. It doesn't matter if the list has redundant entries (typically SwissProt and the protein giving rise to the SwissProt entry). After retrieval, set the "Find Related Data" to "Taxonomy" and wait for the options to load, then click "Find Items".
Miraculously, this returns a page that can be set to display a text phylogenetic tree your input sequences, the full set entered with all redundancy removed. That text tree has labelled higher taxonomic nodes and individual species deeper down. Final edits can be made quickly that capture the phylogenetic spread of the variant allele for interpretive purposes.
The two most common outcomes:
- all the species carrying the variant comprise a monophyletic clade. If the origin of the clade is fairly ancient, then the variation is a derived informative adaptive change relative to ancestral (synapomorphy). If the site is invariant in all members of the co-clade (meaning the ancestral state has persisted to all other extant species), then the site is a phyloSNP (definition and examples: 1 2 3 4).
- species carrying the variation are scattered incoherently across the mammalian phylogenetic tree. This means that the variation has arisen multiple times (all fairly recently) but has not persisted when it arose earlier, ie it is not a preferred allele for this protein at this site and gets replaced.