Bison: nuclear genomics
Bison conservation genomics: introduction
The main nuclear genome of bison, like the mitochondrial genome, will have significant conservation management issues because the consequences of nineteenth and twentieth century bottlenecks (and consequent inbreeding) are still with us today.
Most conservation herds are derived from a tiny founding populations and maintained for many decades at far too low a population level, with surplus animals removed episodically without the slightest consideration of population genetic impacts. Other management practices such elimination of predators, winter feeding, gender imbalance, culling of unruly bulls, and trophy hunts also interfere with natural selection (survival of the fittest).
The founding individuals of a given herd -- even previously wild animals experiencing millenia of natural selection -- still have a substantial genetic load . Autosomal recessives form an important component of that load and are the primary focus here. These are gene mutations found in one of the two copies of non-sex chromosomes that are more or less masked by compensation by the properly functioning copy.
When the founding population is small, the gene frequency of an autosomal recessive mutation is necessarily high. As inbreeding is unavoidable, offspring can inherit two bad copies of the gene. In this homozygous state, no compensation can occur and the disease associated with the mutation is fully manifested. Note populations often harbor mutations at different sites in the same gene. Here the affected offspring can be a compound heterozygote -- two bad copies but at different sites in the same protein.
Looking just at same-site autosomal recessives, the two variables are the frequency q of the bad allele in the population and the coefficient of inbreeding f. The latter simply tallies the percentage of identical alleles by inherited descent (autozygosity). There is an assumption here that would not be valid for the YNP bison nineteenth century bottleneck, namely that the surviving parental animals were not already inbred.
This can be translated into millions of DNA base pairs lacking heterozygosity assuming a bison nuclear genome size of three billion. Then for f at 1/4, there 716,000,000 base pairs of inbreeding derived homozygosity, enough for 5,000 protein coding genes. This DNA will be somewhat broken up into blocks by recombination. For f at 1/8, these numbers are 358,000,000 bp and 2,500 genes. Other coefficients of inbreeding are quickly computed:
- Father/daughter, mother/son or brother/sister → 25%
- Grandfather/granddaughter or grandmother/grandson → 12.5%
- Half-brother/half-sister → 12.5%
- Uncle/niece or aunt/nephew → 12.5%
- Great-grandfather/great-granddaughter or great-grandmother/great-grandson → 6.25%
- Half-uncle/niece or half-aunt/nephew → 6.25%
- First cousins → 6.25%
- First cousins once removed or half-first cousins → 3.1%
- Second cousins or first cousins twice removed → 1.6%
- Second cousins once removed or half-second cousins → 0.78%
The frequency of an autosomal recessive disorder in the offspring of a consanguineous mating is then qf + qq (1-f). Inserting various coefficients of inbreeding and realistic values of deleterious alleles, it quickly emerges that almost all autosomal recessive disease in bison arises from inbreeding. Very rarely does it arise in the offspring of remotely related animals.
Example: suppose a bison bull is dominant for a few years. If it breeds with a calf it previously sired, f is 1/4. The odds that recessive disease did not result from inbreeding only exceed 50-50 when q exceeds 1/3. However q = 0.1 is the largest q known in human disease (hemochromatosis). Cystic fibrosis is another extreme case but there q is only 0.02. For a bison disease allele at that frequency, with a disease observed in the offspring, the odds are overwhelming (94%) that it came from inbreeding, not mating of unrelated bison representative of the whole populations. The odds are still high for the grandparent and first cousin situations (88% and 77%) and are higher still for lower q that are more typical. In summary, autosomal recessive disease in bison can be brought back to natural levels simply by avoidance of inbreeding.
Extensive whole genome sequencing in humans has established that each individual human carries 275 loss-of-function variants and 75 variants previously implicated in inherited disease (both classes typically heterozygous and differing from person to person), additionally varying from the reference human proteome of 9,000,000 amino acids at 10,500 other sites (0.12%). The deleterious alleles include 200 in-frame indels, 90 premature stop codons, 45 splice-site-disrupting variants and 235 deletions shifting reading frame.
In bison the overall genetic load will surely be worse in view of extreme bottlenecks, small herd size history and unavoidable inbreeding. Offspring with deleterious nuclear genes in the homozygous state will be more abundant than in humans who are inbred too but not nearly to the same extent.
Measuring incest in bison
Little bison nuclear genome data is currently available but that situation is changing rapidly with ongoing whole genome sequencing projects not only for bison but also of closely related species such as yak, water buffalo, domestic cow and fossil steppe and plains bison that can help establish a baseline of normality for current conservation herd bison.
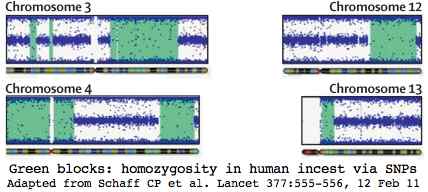
Humans however are already intensively studied. Here incest studies in human have transferable implications to bison herds with limited a number of bulls or a single bull maintaining breeding dominance across generations. The graphic at left shows how a human SNP chip detected incest in a 3-year-old boy with multiple medical issues without access to parental dna.
The green blocks show 668 million base pairs of DNA homozygosity out of the 716 Mbp expected for parent-child incest (coefficient of inbreeding 1/4, human genome size 3.000 Mbp). This represents a quarter of the genes, so approximately 5,000 of which 62 would be expected to have carried deleterious mutations. Some 31 on average would now be homozygous deleterious in the child.
Humans exiting Africa experienced significant bottlenecks then and during subsequent glaciations and climate change as well as founding population migrations. Inbreeding was unavoidable at times. Cousin marriage remains very common today in human populations, with some long been closed to outsiders and now rife with autosomal recessive disease. Thus there is considerable applicability of human data to the bison situation.
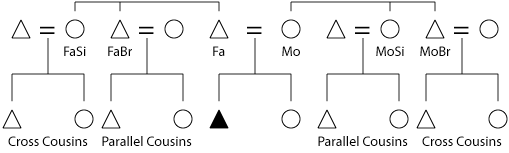
The diagram below shows genealogical terminology relative to inbreeding. It is important to track gender because X-linked mutations manifest themselves readily in males (because the X chromosome there is single copy). Additionally, the mitochondrial genome is maternally inherited and the two genomes need to be co-managed. The Y chromosome is also of interest because its non-recombining portion in bison-cow hybrid herds would still be intact (initial crosses always used a bison bull).
Incest is a crime in nearly all human societies but management-driven incest in bison is not. The SNP chip here had 620,901 markers, representing 12x the resolution available for the comparable cattle chip applied to bison. Thus the bison chip would give clear results but not the sharp resolution because the median marker spacing would slip to 32.4 kbp and the average spacing to 56.4 kbp. For matings between bison related at the second degree (uncle-niece, double first cousins), the inbreeding coefficient is 1/8 and expected level of homozygosity 358 Mb. Here the calf would carry roughly 15 deleterious mutations.
Bison are routinely corralled and tested for previous exposure to brucellosis. The blood samples taken also serve for DNA sampling, where a tiny volume placed on special filter paper would be stable for years at room temperature. These FTA cards provide DNA suitable for readout on the widely used bovine SNP Illumina beadchip. Thus it is fast and cheap to determine the extent of inbreeding at Yellowstone National Park even though cattle introgression (the original use of the chip in bison) is not the issue there. Inbreeding and long-ago introgression are just opposite extremes.
Actual opportunistic measurement of corralled, radio-collared, or naturally expired animals is vastly preferable to academic exercises in theoretic population modeling. There is no real interest in maintenance of neutral allele frequencies measured by microsatellites or junk DNA SNPs but rather in consequential frequencies of deleterious alleles in specific genes that are the legacy of the initial bottleneck, as well as adaptive alleles. The genes and alleles of interest have only been determined so far for bison mitochondria.
Real bison herds are impossibly complex. The females are strongly matrilocal. Although distinct herds may persist for decades because of physical barriers between valleys, bulls and sometimes family groups of cows may wander between them. The herd sizes and composition fluctuate dramatically from year to year depending primarily on the severity of culls, its targeting to extended family groups, snowfall in winter, and susceptibility to predation and disease.
One wonders what management purpose the obscure theoretical constructs of population ecology can serve in real world conservation genomics, given real bison herds have constantly shifting and essentially unmeasurable hereditary allele parameters in 20,000 genes, with two weakly related bison differing at more than 10,500 amino acid sites.
Microsatellites are worthless for conservation genomics
(to be continued)
Results to date from the bovine SNP chip
(to be continued)
The bison prion gene marker M17T
The prion gene PRNP is one of the most intensively sequenced of all mammalian autosomal genes. Bison has three independent sequences though none are geographically sourced at their GenBank entry or accompanying publications. The source of AY769958 is WGFD WBb0401, suggesting Wyoming Game and Fish Department, Wyoming Bison bison 0401 but that sequence is not further discussed (or even used) in full text.
The other 274 bison sequenced came from four unnamed American federal herds and one private herd. M17T being the only variant observed, only two GenBank entries were made. Sample codes CR227All1 and CR227All2 have never been used again in GenBank submissions (this requires Entrez advanced search as these are not accession numbers). The first portion CR227 has not been used again either; CR does not correspond to author initials or a bison refuge acryonym. The frequency of M17T was not reported. It is not present in WGFD WBb0401.
The bison PRNP gene overall is identical to those of its outgroups, yak, domestic cow, and water buffalo as befits a slowly evolving protein (upper quartile). However an interesting bison allele has been identified, namely M17T. This lies well within the signal peptide cleaved during maturation so would not appear in the final GPI-anchored protein on the exterior of the cytoplasmic membrane unless abnormally processed.
Consequently this allele cannot influence the species barrier of bison to prion disease. The species barrier itself is difficult to predict but that of bison will be identical to cow. That risk comes from two main sources: germline or somatic mutation in an individual or transmission from deer, elk or moose affected with chronic wasting disease (scrapie that has previously crossed the cervid species barrier). Bison are not mixed with cattle because of the brucellosis issue and sheep allotments are seldom in proximity, so gransmission from a transmission from public lands mad cow or scrapie sheep is implausible.
Inherited prion disease is autosomal dominant with high penetrence so does not revolve around inbreeding or high background frequency of disease alleles in a population (though late onset can bring about large pedigrees). It represents toxic gain of function, not loss of normal protein function which remains unknown despite 12,051 scientific studies as of 17 Feb 2011.
The greatest single risk for inherited prion disease is amplification of the octapeptide repeat region PHGGGWGQ by replication slippage. Here bison have six octapeptide repeats, in the normal range but nonetheless a risk factor for disease expansion. Note the number of bison in conservation herds is very small relative to the one-per-million incidence of repeat expansion observed in human. Domestic cattle may have four to seven repeats with the 7x repeat in Brown Swiss a borderline concern given the situation in human.
CpG hotspot mutations dominate the point mutation spectrum in mammals. The most common outcome for a CpG mutation is the purine transition CpA. Any of 12 amino acid substitutions can arise. When CpG resolves as the pyrimidine transition TpG, 8 non-synonymous outcomes are possible in addition to internal stop codons (which would not cause prion disease). These 20 point mutations are shown at bottom and need individual assessment for disease-causing potential.
A known CpG point mutation in domestic cattle E211K (corresponding to homologous E200K human pathogenic allele) causes genetically based mad cow disease, as specifically predicted eleven years earlier by the author of the present article. That constitutes a proven risk factor for bison which has CpG in the identical position (bottom).
Again, the number of bison in conservation herds is dwarfed by the 103,000,000 million cattle for which E211K has been observed (heterozygously) in one source cow its calf, and by presumption some degree of ancestors, though prevalence is low. Although artificial insemination is practised on a massive scale, it has not been determined by haplotyping whether E211K arose from the sire. The affected ten year old cow was a Bos indicus x Bos taurus hybrid.
The signal peptide allele M17T is neither deleterious to normal function nor causative for prion disease (being discarded). It does not prevent proper maturation cleavage according to thoroughly vetted bioinformatic prediction tools such as SignalP. From the phylogenetic standpoint, methionine and threonine constitute the reduced alphabet at position 17. Alanine is less common but also tolerated.
These amino acids are neighbors in the genetic code related by single basepair transitions (threonine taken as central). Note however that amino acids such as branched chain aliphatics also related by simple common mutations but not observed. This raises the question of why the PRNP signal peptide is so conserved relative to other signal peptides, for example PLBD2.
Some 4500 genes have signal peptides in mammals, all interfacing with the same signal receptor processor (SRP). Many-to-one protein interactions cannot co-evolve (as claimed for speciation by Ernst Mayr) because if the SRP changed to accommodate a change in PRNP, that change throws off its adaptive fit to the other 4499 proteins it must continue to recognize.
This conservation suggests that the signal peptide of PRNP might not always be cleaved or influence protein processing in some other way, resulting in a mix of ultimate cellular destinations or different membrane topologies. Indeed two papers provided evidence for alternative C- or N-terminal insertion of retained single-pass membrane retention in the endoplasmic reticulum lumen.
However no significance to this was ever found and the research track was abandoned. Strong selection on the prion protein overall remains a mystery in view of minimal impacts of knockout mutations and implausible compensation by an immensely diverged tandem paralog PRND whose signal peptide does not exhibit such striking conservation.
Regardless, M17T is implausibly functionally significant because of its phylogenetic incoherence. Note indels in signal peptides are also unusual and non-recurrent; the one at position 3 is a striking synapomorphy of euarchontoglires but with no known functional significance.
Thus the primary interest in M17T is as a neutral nuclear gene marker in bison. However it is not represented in the bovine SNP chip because the polymorphism has not been observed among a thousand bovine PRNP genes sequenced in several dozen widely varying breeds. It does not appear that M17T in bison reflects lineage sorting of a persistent polymorphism because of its absence both in intensively sampled cattle and other close-in pecoran ruminants.
Allele frequency in bison might be fairly high in some herds because of bison expansion from bottlenecks. Its correlation with mitochrondrial haplotypes or nuclear microsatellites might actually be known since bison dna samples are hard to come by and could have been used for multiple purposes. PRNP is not on chromosome X or Y in any mammal to date with assembled genome so its inheritance could not correlate with maternal inheritance of mitochondrial dna . Microsatellites could be mapped into cow genome to determine their proximity to the PRNP gene.
Although M17T does not occur in cattle, its presence in bison -- even in the homozygous state -- is not proof of the absence of past introgression because subsequent back-crossing with bison could have quickly eliminated it.
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Bison bison CR227All1 AY769958
MVKSHIGSWILVLFVATWSDVGLCKKRPKPG Bison bison CR227All2
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Bison bonasus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Bos taurus
MVKRHIGSWILVLFVVMWSDVGLCKKRPKPG Bubalus bubalis
MVKSHIGSWILVLFVVMWSDVGLCKKRPKPG Syncerus caffer
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Rangifer tarandus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Alces alces
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Capreolus capreolus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Kobus megaceros
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Connochaetes taurinus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Ammotragus lervia
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Hippotragus niger
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Capris hircus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Cervus elaphus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Cervus elaphus nelsoni
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Dama dama
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Odocoileus virginianus
MVKSHIGSWILVLFVAMWSDVALCKKRPKPG Oryx leucoryx
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Ovibos moschatus
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Ovis aries
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Ovis canadensis
MVKSHIGSWILVLFVAMWSDVALCKKRPKPG Tragelaphus strepsiceros
MVKSHIGGWILVLFVAAWSDIGLCKKRPKPG Sus scrofa
MVKSHMGSWILVLFVVTWSDVGLCKKRPKPG Camelus dromedarius
MVKSHMGSWILVLFVVTWSDMGLCKKRPKPG Vicugna vicugna
MVKSLVGGWILLLFVATWSDVGLCKKRPKPG Myotis lucifugus
MVKNYIGGWILVLFVATWSDVGLCKKRPKPG Pteropus vampyrus
MVKSHIANWILVLFVATWSDMGFCKKRPKPG Tursiops truncatus
MVKSHIGGWILLLFVATWSDVGLCKKRPKPG Canis lupus familiaris
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Felis catus
MVKSHIGSWLLVLFVATWSDIGFCKKRPKPG Mustela putorius
MVKSHIGSWLLVLFVATWSDIGFCKKRPKPG Mustela vison
MVKSHIGSWILVLFVAMWSDVGLCKKRPKPG Ailuropoda melanoleuca
MVKSHVGGWILVLFVATWSDVGLCKKRPKPG Equus caballus
MVRSHVGGWILVLFVATWSDVGLCKKRPKPG Diceros bicornis
MVKNHVGCWLLVLFVATWSEVGLCKKRPKPG Erinaceus europaeus
MVTGHLGCWLLVLFMATWSDVGLCKKRPKPG Sorex araneus
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Homo sapiens
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Pan troglodytes
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Gorilla gorilla
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Pongo pygmaeus
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Nomascus leucogenys
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Hylobates lar
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Symphalangus syndactylus
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Macaca arctoides
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Macaca fascicularis
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Macaca fuscata
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Macaca mulatta
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Macaca nemestrina
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Papio hamadryas
MA--NLGCWMLFLFVATWSDLGLCKKRPKPG Callithrix jacchus
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Cebus apella
MA--NLGCWMLVVFVATWSDLGLCKKRPKPG Cercopithecus aethiops
MA--NLGCWMLVVFVATWSDLGLCKKRPKPG Cercopithecus dianae
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Colobus guereza
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Presbytis francoisi
MA--NLGCWMLVLFVATWSDLGLCKKRPKPG Saimiri sciureus
MA--KLGYWLLVLFVATWSDVGLCKKRPKPG Tarsius syrichta
MA--NLGCWMLVVFVATWSDVGLCKKRPKPG Microcebus murinus
MA--RLGCWMLVLFVATWSDIGLCKKRPKPG Otolemur garnettii
ME--NLGCWMLILFVATWSDIGLCKKRPKPG Cynocephalus variegatus
MA--QLGCWLMVLFVATWSDVGLCKKRPKPG Tupaia belangeri
MA--NLGYWLLALFVTMWTDVGLCKKRPKPG Mus musculus
MA--NLGYWLLALFVTTCTDVGLCKKRPKPG Rattus norvegicus
MA--NLGYWLLALFVTTCTDVGLCKKRPKPG Rattus rattus
MA--NAGCWLLVLFVATWSDTGLCKKRPKPG Cavia porcellus
MA--NLGYWLLALFVTTWTDVGLCKKRPKPG Apodemus sylvaticus
MA--NLGCWLLVLFVATWSDLGLCKKRTKPG Dipodomys ordii
MA--NLSYWLLAFFVTTWTDVGLCKKRPKPG Clethrionomys glareolus
MA--NLSYWLLALFVATWTDVGLCKKRPKPG Cricetulus griseus
MA--NLSYWLLALFVATWTDVGLCKKRPKPG Cricetulus migratorius
MA--NLGYWLLALFVTMWTDVGLCKKRPKPG Meriones unguiculatus
MA--NLSYWLLALFVAMWTDVGLCKKRPKPG Mesocricetus auratus
MA--NLGYWLLALFVATWTDVGLCKKRPKPG Sigmodon fulviventer
MA--NLGYWLLALFVATWTDVGLCKKRPKPG Sigmodon hispiedis
MV--NPGCWLLVLFVATLSDVGLCKKRPKPG Spermophilus tridecemlineatus
MV--NPGYWLLVLFVATLSDVGLCKKRPKPG Sciurus vulgaris
MA--HLGYWMLLLFVATWSDVGLCKKRPKPG Oryctolagus cuniculus
MA--HLSYWLLVLFVAAWSDVGLCKKRPKPG Ochotona princeps
MVKSHLGCWIMVLFVATWSEVGLCKKRPKPG Cyclopes didactylus
MVRSRVGCWLLLLFVATWSELGLCKKRPKPG Dasypus novemcinctus
MVKGTVSCWLLVLVVAACSDMGLCKKRPKPG Echinops telfairi
MVKSSLGCWILVLFVATWSDMGLCKKRPKPG Elephas maximus
MVKSSLGCWILVLFVATWSDMGLCKKRPKPG Loxodonta africana
MVKSSLGCWMLVLFVATWSDVGLCKKRPKPG Procavia capensis
MMKSGLGCWILVLFVATWSDVGLCKKRPKPG Orycteropus afer
MVKSGLGCWILVLFVATWSDVGVCKKRPKPG Trichechus manatus
MAKIQLGYWILALFIVTWSELGLCKKPKTRPG Macropus eugenii
MGKIHLGYWFLALFIMTWSDLTLCKKPKPRPG Monodelphis domestica
MGKIQLGYWILVLFIVTWSDLGLCKKPKPRPG Trichosurus vulpecular
Effect of CpG hotspot mutation on bison prion protein
normal: 1 MVKSHIGSWILVLFVAMWSDVGLCKKRPKPGGGWNTGGSRYPGQGSPGGNRYPPQGGGGW 60
MVKSHIGSWILVLFVAMWSD+GLCKK+PKPGGGWNTGGS+YPGQGSPGGN YPPQGGGGW
CpG CpA: 1 MVKSHIGSWILVLFVAMWSDMGLCKKQPKPGGGWNTGGSQYPGQGSPGGNHYPPQGGGGW 60
normal: 61 GQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGGWGQGGTHGQWNKPSKPKTNM 120
GQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGGWGQGGTH QWNKPSKPKTNM
CpG CpA: 61 GQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGGWGQGGTHSQWNKPSKPKTNM 120
normal: 121 KHVAGAAAAGAVVGGLGGYMLGSAMSRPLIHFGSDYEDRYYRENMHRYPNQVYYRPVDQY 180
KHVAGAAAAGAVVGGLGGYMLGSAMSRPLIHFGSDYED YY ENMH YPNQVYYRPVDQY
CpG CpA: 121 KHVAGAAAAGAVVGGLGGYMLGSAMSRPLIHFGSDYEDHYYHENMHHYPNQVYYRPVDQY 180
normal: 181 SNQNNFVHDCVNITVKEHTVTTTTKGENFTETDIKMMERVVEQMCITQYQRESQAYYQRGASVIL 245
SNQNNFVHDCVNITVKEHTVTTTTKGENFT+TDIKMME+VVEQMCITQYQRESQAYYQ+GASVIL
CpG CpA: 181 SNQNNFVHDCVNITVKEHTVTTTTKGENFTKTDIKMMEQVVEQMCITQYQRESQAYYQQGASVIL 245
normal: 1 MVKSHIGSWILVLFVAMWSDVGLCKKRPKPGGGWNTGGSRYPGQGSPGGNRYPPQGGGGW 60
MVKSHIGSWILVLFVAMWSDVGLCKK PKPGGGWNTGGS YPGQGSPGGN YPPQGGGGW
CpG TpG: 1 MVKSHIGSWILVLFVAMWSDVGLCKK-PKPGGGWNTGGS-YPGQGSPGGNCYPPQGGGGW 58
normal: 61 GQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGGWGQGGTHGQWNKPSKPKTNM 120
GQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGGWGQGGTHGQWNKPSKPKTNM
CpG TpG: 59 GQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGGWGQGGTHGQWNKPSKPKTNM 118
normal: 121 KHVAGAAAAGAVVGGLGGYMLGSAMSRPLIHFGSDYEDRYYRENMHRYPNQVYYRPVDQY 180
KHVAGAAAAGAVVGGLGGYMLGSAMSRPLIHFGSDYED YY ENMH YPNQVYYRPVDQY
CpG TpG: 119 KHVAGAAAAGAVVGGLGGYMLGSAMSRPLIHFGSDYEDCYYCENMHCYPNQVYYRPVDQY 178
normal: 181 SNQNNFVHDCVNITVKEHTVTTTTKGENFTETDIKMMERVVEQMCITQYQRESQAYYQRGASVIL 245
SNQNNFVHDCVNITVKEHTVTTTTKGENFTETDIKMME VVEQMCITQYQRESQAYYQ GASVIL
CpG TpG: 179 SNQNNFVHDCVNITVKEHTVTTTTKGENFTETDIKMME-VVEQMCITQYQRESQAYYQ-GASVIL 245