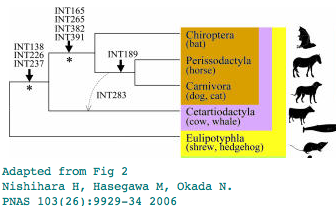
Pegasoferae?
Can rare genomic events establish Pegasoferae?
Pegasoferae is a novel proposal for the phylogenetic ordering within Laurasiatheres, grouping bats, perissodactyls, and carnivores to the exclusion of the other hoofed mammalian group artiodactyls. Bats have been placed in many previous locations, notably in the Euarchonta wing (with primates). While that particular idea is clearly refuted by many lines of evidence, the proper placement of bats remains under discussion.
Rare genomic events may be more useful for this than maximal likelihood because the orders of Laurasiatheres may have diverged relatively rapidly. Retroposon events are so numerous per million years however that they may be able to resolve branching at these tight nodes. However they suffer from homoplasy in that separate insertion events from a given parental element can look very similar and because deletions over time (no selection for their retention) can cause their disappearance and so confusion with lineages that never had the insertion.
Qualifying retroposons need to be situated between two well-conserved flanking markers because orthology is otherwise difficult to decisively establish in intergenic regions. These markers ideally are no more than 1500bp apart to allow tiling of traces for species without assemblies (eg vicugna, pig, dolphin, macrobat in Laurasiatheres) and spanning PCR runs. Higher sampling density greatly enhances the ability to correctly infer the sequence of events.
Short coding indels in coding exons can also be phylogenetically informative. Here if the exon is otherwise quite conserved, the risk of homoplasy (recurrent events at the same or indistinguishably similar position) is fairly low. These events are inherantly rare first because conserved regions of a protein may not admit indels structurally (ie are inactivated) and second because the window of relevancy for a given tree topology issue may only be a small fraction of elapsed evolutionary time (eg 1 million year stem on a 85 myr branch).
Coding indels can exhibit the usual problems of lineage-sorting: two co-existing alleles at the time of speciation that resolve differently in descendent lineages. Insertions, while a third as common as deletions and so less likely to have arise multiple times, are more subject to subsequent confusing reversion; deletions are less likely to revert to ancestral length for lack of genetic mechanism. It goes without saying that indels from repetitive regions or in dna of anomalous composition are wholly unsuitable for taxonomic purposes.
Analysis of L1MA9 retroposon INT189
The phlogenetic distribution of the L1MA9 retroposon INT189 has been taken as evidence for bats being the immediate outgroup of horse + dog. That interpretation can be revisited using newly available genomes. Yet only two sequences representing perissodactyl and carnivore are at GenBank as cat assembly has a gap in the critical region. But other new data in 3 bats and 4 cetartiodactyls and 2 shrew/hedgehog confirm the lack of L1MA9 near the distal exon.
The trouble is a second L1MA9 element lies upstream of the MER58A middle marker. This is lacking in both carnivores. Evidently it was deleted in stem carnivore -- otherwise it would be providing evidence for carnivores being outgroup to cows + bats + horse. In short this single intron is providing 'support' for two contradictory topologies.
The sizes of many bat genomes have been experimentally determined: the 30-genus average of 2.6 gbp is about 500,000,000 bp less than human. Since bats in essence have the same 20,000 coding genes as other mammals, that discrepancy has to arise from less intronic and intergenic dna. Possibly bats had fewer active retroposing elements. Far more likely, bats they have an average number and the discrepancy arises from a faster rate of deletions than insertions.
Thus for taxonomically informative (ancestral laurasiathere) retroposons, many millions of deletion events have occured. Since the L1MA9 elements here are only 100bp or so, it would come as no surprise if a high percentage of the older relevent ones have experienced partial (or full) deletions making them unrecognizable with RepeatMasker.
Thus presence of a retroposon in a given orthologous position bat can be informative but absence is not so informative. INT189 is an absence. That one event isn't insufficient anyway to establish branching order. So bat/horse/carnivore tree topology remains unresolved. If horse is the outgroup to carnivore + bat -- and cow outgroup to all of these -- then hoofed animals are parsimoniously ancestral (rather than arising twice by convergent evolution) and bat and carnivore lost hooves (a bit unreasonable as dog and bats retain the ancestral 5 digits).
Summary of the phylogenetic distribution of the L1MA9 retroposon INT189: >PGM2_canFam Canis familiaris (dog) abseny -MER58 182-265 23% -L1MA9 6069-6302 27% >PGM2_felCat Felis catus (cat) genomic del absent -MER58 no data >PGM2_equCab Equus caballus (horse) -L1MA9 6172-6264 26% -MER58A 1-145 23% -L1MA9 6050-6302 23% >PGM2_myoLuc Myotis lucifugus (microbat) -L1MA9 6174-6264 20% -MER58A 38-157 26% >PGM2_pteVam Pteropus vampyrus (macrobat) -L1MA9 6161-6291 25% -MER58A 35-145 29% >PGM2_pipAbr Pipistrellus abramus (microbat) -L1MA9 6180-6301 252% -MER58A 38-157 24% >PGM2_bosTau Bos taurus (cow) -L1MA9 6155-6263 28% -MER58A 7-157 21% >PGM2_turTru Tursiops truncatus (dolphin) -L1MA9 6155-6265 29% -MER58A 37-148 21% >PGM2_susScr Sus scrofa (pig) cdna + tiled -L1MA9 6159-6264 27% -MER58 212-271 28% >PGM2_vicVic Vicugna vicugna (vicugna) tiled -L1MA9 6162-6310 24% -MER58A 35-157 20% >PGM2_ateAlb Atelerix albiventris (hedgehog) ... ... >PGM2_sorAra Sorex araneus (shrew) ... ... >PGM2_canFam Canis familiaris (dog) -MER58 182-265 23% -L1MA9 6069-6302 27% VISAELASFLATKNLSLSQQLKAIYVE YGYHITKASYFICHDQGTIKKLFENLRNY GTCATCAGCGCCGAGTTGGCTAGCTTTCTAGCAACCAAGAATTTGTCTTTGTCTCAGCAGCTAAAGGCCATTTATGTTGAGTACGTTTCTATTAACTCTG TTTAATTGAAATAATACTTTTTAAAAGTTTTATTATGTTTTTATGTGTGACACTAATATTCTAACCCTCTTACTTTGGGTGAGGGTTCTTCTGAAAACTA AAGGATCACTTTTTCTTTTAATGCTTAACTATTCAATACTAATTATCACTTATGACTGTGTTAATCCTTAACAAATGAGAACATCAGTTGCAGAAATAGC TAATTGAGGAGGGTGATTCCCTGATGTCAGAAAGGACAAAGGTTTTCGTGAAACATCTATTACGTGTTTAGAgccactagtcaagtctgcctttgtagtg caaaagcagctgatggcaagacgtacaggaatgggtgtggtgtggctgcaatgaaaTGAAACTTTCACCTCCCAAGATAGGCCGAAGGCCAGGCAGCAGT TTGGCAATACCTGGGGTCAATAGTTATACCTCTTTTTTATGCTAAATTATTCCTTTGAAGCTAGTCATTGTTATCGTTTCATTTAGCTTAAAATATACTG ATTGCTACATGTTCTGTATACACCACGTGAGATTATTTGTTCCTCATTTTGCATATTTGTACTTTTtttattgagatgtaattgacattaatgtcaggta taataacataatgattcgatatttatatattattacaaagtgatcaccatagtaagtcgagttaacatccacaccacatataatcacaaatattcattct tgtgatgatagcttttatgatctgtggtcttagcaactttcaaatatacagtacaatactagtagatacagtcaccaagttatatatATATAATTTTATT TCTTTTGATAGATATGGCTACCATATTACCAAAGCTTCCTATTTTATCTGCCATGATCAAGGCACCATTAAAAAATTGTTTGAAAACCTTAGAAACTAC >PGM2_felCat Felis catus (cat) genomic del incomplete coverage -MER58 ASFLATKNLsLSQQLKAIYGE YGYRITKASYFICHDQGTIKQLFENLRNY GCTAGCTTTCTAGCAACCAAGAATTTGTTTGTCTCAGCAGCTAAAGGCCATCTACGGCGAGTAAGTGTCTTCTAACCTGGTAAAGAAGTAATAG TGTTAAATATTTTCTTATGGTTCTACGTGTGAGATATTAATATTCTTTCTAATGCTCTTTGGTTGTGAATTCTATTTCTTTTTCTTTTTTTAATGTTTAT TTATTTTTGAGAGAGAGAGAGAGAGAGATGGAGTATGAGCAGGGGAGGGGCAGAGAGAGAGGGAGATACAGAATCCAAAGCAGGCTCCAGGCTCTGAGCT GTCAGCACAGAGCTCCACACGGGGCTTAAACTCACAAACCATGAGATCATGACCTGAGCTGAAGTCAGACACTCAACCGTTTGAGCCACCCACGTGCCCC ATGAATTCTATTTCTTATGAAACTAAATAATCATCTTTTCTTTTGATACTTAACCATGTAATGGTAATTATCATTCACGATTGCACGAATCCTTAACAAA TGAGGGCATCAGTTGCAGAAATAGCTAATTGAAGAATGTGATTTTAAGTGTGTGATGTCAAAAAAGATTAAAGGTGTTCATGAAATCTCTATTAAGTTTT TAGAGCAATGACCCAGGTCTGCCTTTATAAAGTGCAAAAGCAGCCCGTGGCAACACGTTGCAGTAAGACTCTTACTTACAAATACAGGCTAAAGGCCAGG CAGCAGTTTGGCAATCCCCAGGGTTAATTGTTGTACCTCTTTTTTATGCTAAATTATTCCTTTGAAGGTACTCATGGCTATTTGTTTCATTTGGTTTAAA ATATACTGGTTGACAAATGTACACTGTGTGGAATTATGTGTTCCTCATTTTGCATATTTGTATTTCCTTAACTGAGATATAACTGACATTAGTTTCAGGT ATGCGATACAGTGTTTCAATATCTGTATATATTACAAAATGATCATCACAGTACATCTAGTAACAGTCGCACCACACTTAATACAAAAGT TCCaTATGGCTACCGTATTACCAAAGCTTCATATTTTATTTGCCATGATCAAGGCACCATTAAACAATTATTTGAAAACCTTAGAAACTAT >PGM2_equCab Equus caballus (horse) -L1MA9 6172-6264 24% -MER58A 1-145 23% -L1MA9 6172-6264 26%-L1MA9 6050-6302 23% VISAELASFLATKNLSLSQQLKAIYVE YGYHITKASYFICYDQDTIKKLFENLRNY GTCATAAGCGCAGAGTTGGCTAGCTTTCTAGCAACCAAGAATTTGTCTTTGTCTCAGCAGCTAAAGGCCATCTATGTTGAGTAAGTTTCTATTAACTCTC TTTAACTGAGGTAATTTTTTTTATTAGtttcaaatgtacaacataatgattcaatgtatgtatatattttgaaatgatcgccacaataagtctggctaac ctgtatcaccgacatagGGCTCTTTTTAAATGTTTTATGTTCTTTTGCATGAAACAGTAATATTCTTTTGAATGCTCTTACTTTAGCTATGAATTGTTCC TTATGAAAACTAAGTAAGAGATCACTTTTTCCTTTCGATACTTAACCACTTAGTAGTATTACCCTTTGTGATTGCATTAATCCTTAACAAATGAGAACAT TAGTCACGGAAATGGTGAAGTGAAGAATGTAATTTTCAGTGTCTGAGGTCAAAAAAGATTAAATGTGTTCATGAAACATCTATTTAGTCTTTAACTTCat tgctcagctctgcctttgtagtgcagaaacagccggggacaatacataatgtaatgggtgtggggtggctgtgttccagtagatcttttacttaaaaata caggccgaaggccaggcagcagtttggcaatccctgGGGGAGATTATTGTACCTTTTTTTAATGTTAAATTATCCCTTTGAAGTTAGTCATGGTTATTTC ATTTAGTTTAGAATATAATGGTTAATACATAGTGTATGTACACCATGTGGAATTATTTTTTCCCATTTTGCATTTCTTCTtttgttgagatataattaac atagaacattatattagcttcaggtgtacagtgtaattatttgataattgtatatattgcagattgatcaccaccataagactagttaacatccatcacc acacatagttataaatttttttcttgtgatgagaacttttaaggtctattctcttagcaaccttcaaatatacaatacagtattattaattctagtcacc gtgctgtgtattatatcctcatgacccattTTATTATTTTGTTTCGAAAGGTATGGCTACCATATTACCAAAGCTTCATATTTTATCTGCTATGATCAAG ACACCATTAAAAAATTGTTTGAAAACCTTAGAAACTAC >PGM2_myoLuc Myotis lucifugus (microbat) -L1MA9 6174-6264 20% -MER58A 38-157 26% VISAELASFLATKNLSLSQQLKAIYVE YGYHITKASYFICHDQGTIKKLFENLRNY AAPE01636299 GTCATAAGCGCAGAGCTGGCTAGCTTTCTTGCAACCAAAAATTTGTCTCTGTCTCAGCAGCTAAAGGCCATCTACGTTGAGTAAGTTTCTATTGATTATTG AATTGAAGTAATATAGTTTGATTAGTTTCATGTGTACAATGTAATGATTCAATATGTGTATATATTGGGACATGGTTGCCACAATAAGTCGTTAACATAC ATTACCACATGTGGCAATGTATTTTAAGTGTATTATGTTCTTGCGTATGAGATGCTAATGTTCTTTCCAAAGCTCTGACTTTAGTTATGAATTCTATTTC TTAAGAAAACGAAACGAGATTATCTTTTCCTTTTGATACTTACCATTTGTGATAGCACTAATCTTTACTAAATGAGAACATGACACAGAATGTGATTTTA AGTGTCTGATGCCAAAAAAGATTAAATGTGTTCATGAAACGTCTATTTAGTCTTTATAGCAGTTTCTCAACTCTTGCCTTTCTGATGCAAAAGGAGCCAG ACACAGTACATAATGCAATGGGCGTGGTATGGCTGTTCCAGTATAATTTTACTTACAAGTATAGGCTGAAGGCAAGGTAGCAGCTTGGTGAGCCCTCGGG TAAATTGTTGCACCTCCTTTTAATGCTAAATGATTGCTTTGAAGCTAGTCATGGTCATTTGTCTCATTACGTATTTGAGAATGTGCTGGTTGGTGCCCGT TCTGTATATGCTATGCATAATTATTTGTTCCTCATTTTGCATGTATTTGTATTTGTTTTGATAGGTATGGCTACCATATTACCAAAGCTTCATATTTTAT CTGCCATGATCAAGGAACCATTAAGAAATTATTTGAGAACCTTAGAAACTAT >PGM2_pteVam Pteropus vampyrus (macrobat) -L1MA9 6161-6291 25% -MER58A 35-145 29% VISAELASFLATKNLSLSQQLKAIYVE YGYHITKASYFICHDQGTIKKLFENLRNY GTCATAAGCGCGGAGTTGGCTAGCTTTTTAGCAACCAAGAATTTGTCTTTGTCTCAGCAGCTAAAGGCCATCTATGTTGAGTAAGTTTCTATTGACTCTA CATAACTGAAATAATATTTTTTATTAGTTTCAGGTGTACAGCACAGTGATTCGGTATATGTATATATTATGACATGATTGCTATAAGTCTATTGCATGCA TCAGTCTATTACTACATGCATCACCACACGTAGTAATATTTTTAAATGTATTATGTACTTGTGCACAAGATACTAATATTCTTTCCAATGCTCTTACTTT AGTTATGAATTCTATTTCTTATAAAAACCAAATAAGAAATTACCTTTTCCCTTTGATACTTAGCCATTTAATAGTAATTACCATTTGTGATGACAGTAAC CTTTACCAGATGAGACATTAGCCACAGAAACAGCTAAAGAATATGATTTTAAGTGTCCGATGTCAAAAGATTAAATGTGTTTATGAAACATCCTATTTAG TCTTTTTATAGCATTATTCAGCTGTGCCTTTGTAGTACAAAAGCAGCCAGACCCGATGCATATGTAATGGGTGCAGCGTGGCTACATTTCTGTAAAATTT TTACTTACAAATATAGGCTGAAGGCCAGGCAACAGTTTGGTGATCCCCTGAGTAAATTGTTATACTTCTTTCTTAATGCTGAACTATTCCTTTGAAGCTA GTCATGGTCATTTGTTTCATTAAGCGTTTTAGAATGTACTGGTTGATACATGTTCTGTGTACACTATGCAGAATGATTTGTTCCTTATTTTGCATGTGTT TGTATTTATTTTGATAGGTATGGCTACCATATTACCAAAGCTTCATATTTCATCTGCCATGATCAAGGCACCATCAAAAAATTATTTGAAAACCTTAGAAACTAT >PGM2_pipAbr Pipistrellus abramus (microbat) -L1MA9 6180-6301 25% -MER58A 38-157 24% AB258957 AIYVE YGYHITKASYFICHDQGTIKKLFENLRNY GGCCATCTATGTCGAGTAAGTTTCTATTGATTATTGAATTAAAGTAATATAATTTGATTAGATTCATGCGTACAGTGTAATGATTCAATACATGTATATA ATGGGACATGGTTGCCACAATAAGTCGTTAACATACATCACCACCTGTGGCAATATATTTTAGGTGTATTATGTTCTTTAGTATGAGACACTAGTACTAA TATTCTTTCCAAGGCTCTGACTTTAGTTATGAATTCTATTTCTTAAGAAAATGAAACGAGATTATCTTTTCCTTTGGATACTTACCATTTGTGATTGCAC TAATCTTGATTAAACGAGAACATTACACAGAATGTGATTTTAAGTGTCTGATGCCAAAAAAGATTACATGTGTTCATGAAACATCTATTTAGTCTTTATA GCAATTTCTCAACTCTTGCCTTTCTGGTGCAAAAGCAGCCTGACACAATACATAATGTAATCGGCGAGGGATGGCTGGTCCAATAAAACTGTACTTACCA ATGTAGGCTGAAGGCAAGGTAGCAGCGTGGTGTTCCCTCAGAATTATTTGTTCCTCATTTTGCACGTATTATTTGTTTTGATAGGTATGGCTACCATATT ACCAAAGCTTCATATTTTATCTGCCATGATCAAGGCACCATTAAGAAATTATTTGAAAACCTAAGAAACTACGATGGGAAGAATAATTAT >PGM2_bosTau Bos taurus (cow) -L1MA9 6155-6263 28% -MER58A 7-157 21% VITAELASFLATKNLSLSQQLKAIYVE YGYHITRASYFICHDQETIKQLFENLRNY AB258958 [L1_Carn7] GTCATAACTGCAGAGTTGGCCAGCTTTCTAGCAACCAAGAATTTGTCTTTGTCTCAGCAGCTAAAAGCCATCTATGTTGAGTAAGTTTCTATTGACTATT TAATTGAAGTAATTTTTTTTTATCAGttcaggtatacaacacagtgattcagtgtatgtctatattgtgaaatgatcacagtggatacaattaacatgca tccccacacaggaatattttttaatgtTTTACTCTCTTCTTGTGCACCCGATACTCATATTCTTTCTGATGCTCTTGCTTTAGTTATGAATTCTATTTCG TATGAAAACTAAATAAGAGATCACCTTTTCCTTTTGCTACTTAAGCAGTTAATAGTAATTACCATTCATGATGACGTTAATCCTTAATAAATGAGAACGT TAGCTGCAGAAATGGCTAAGGGAAGAATGTGATTTTTTAAATGTCCAGTGTTGAAAAAGACTAAATGTGTTCATTAAACATCTATTTAgtctttgtagca attacttatttctgcctttctagtgcaaaagcaaccagacacaaggtaatgggcatgacgtggctgtattccaatgataaaacttttacttacaaacaga gactgagggccACACAGCAGGGCAGTGATTCCTGGTGTAGATTGTTGGACCTCTTTATTTAATGCTGAATTACTCCTTTGAAATTAGTCATGGTTGTTTG TTTTAGAATATACTGTTTGATAGATACATGTTCAGTGTACACTGTGCCCAATTATTTGTCCCTCATTTGCATGTAACCATGTTTGTATTGATAGGTATGG CTACCATATCACCAGAGCTTCGTATTTTATCTGCCATGATCAAGAAACTATTAAACAATTATTTGAAAACCTTAGAAACTAT >PGM2_turTru Tursiops truncatus (dolphin) -L1MA9 6155-6265 29% -MER58A 37-148 21% FISAEVGSFLAQNCLVSAAKAIYV YGYHITKASYFICHDQGTIKKLFENLRNY TTCATAAGTGCAGAGGTTGGCAGCTTTCTAGCACAGAATTGTCTTGTTTCAGCAGCTAAAGCCATCTATGTTGAGTAAGTTCTTCTATGACTGTTAAATG AGTAATGTTTTTTTTCATTTCAGTTGTGCAACACAATGATTCAATGTATATCTATTATTGTGAAATGATTGCAACAAATACAGTTTACATGTATCCCCAC ATGTAGTAATATTTTTTAATGTTTTACTCCGTTCTTATGCATGAGATACTAATATTCTTTCTGATGTCCTTACTTTGGCTATGAATTCTATTGCCTATAA AAACTAAATAAGGGATCACCTTTTCCTTTCGATATTTAACTACTTAATAGTAGTTACCCCTTCATGATGACATTGATTCTTAACAAATGAGAACATTAGT TGCAGAAATGGCTAAGGGAAGAATGTGATTTTTAAGTGTCCAATGTCAAAAAAGACACATGTGTTCACAAAACATGTTTAGCCTTTAAAGCAATTATTCA CCAGTGTCTTTGTAGTGCAAAAGCAGCCAGACACAATACATAAGGTAATGGGCATGGCATGGCTACGTTCCAATAGAGAAACTTTTACTTAGAAATACAG GCTGAGGGCCACAGAGCAGTTCAGCGATCCCTGGGGTAGATTGTTGGACCTCTTTTATAAAATTGGACCTCTTTTTTTTTTTTTTTTTTTTTGGCGGGGG GTACGTGGACCTCTCACTGTTGTGGCCTCTCCCGTTGCAGAGCACAGGCTCCAGACGCGCAGGCTCAGTGGCCATGGCTCGCGGGCCCAGCCGCTCCACG GCATGTGGGATCTTCCCAGACCGGGGCACGAACCCGTGTCCCCTGCGTCGGCAGGCGGACTCTCAACCACTGCGCCACCAGGGAAGCCCTGAACCTCTTT TTTAATGCTGAATTATTCCTTTGAAATTAGTCGCGGTTATTTGTTTTAGAATATACTGGTTGATACATGTTCAGTGTACACTGTGCAGAATTATTTGTTC CTCGTTTTGCATGTAATTGTGTTTGTATTGATAG GTATGGCTACCATATCACCAAGGCTTCGTATTTTATCTGCCACGATCAAGGCACTATTAAAAAATTATTTGAAAACCTTAGAAACTAC >PGM2_susScr Sus scrofa (pig) cdna + tiled -L1MA9 6159-6264 27% MER58 212-271 28% VISAELASFLATKNLSLSQQLNAIYVE YGYHVTKGTYFICHDQGNVKKLFENLRNY GTCATAAGCGCAGAGTTGGCCAGCTTTCTAGCAACCAAGAATTTGTCTTTGTCTCAGCAGCTAAATGCCATCTATGTTGA GTAAGTTTCTATTGACTGCATTTAATTGAAGTAATTTTTTTAATCAGTTTCGGGTGTACGACATAATGATTCAGTGTATATGTATTGTGAAATGATCCCAA TGAGTACAGCTAACATGCATCCCACACGTAATAATATTTTTTTTTCTTTCTTTTTCTTTTTTTAGGGCTACTCCTGTGGCATATGGAAGTTCCCAGGCTA AGGGTCGAATAGGATCCATAGCCGCTAGCCTAAGCCACAGCCACAGCAGCACGGAATTCGAGCCACATCTTTGACCTCCGCTACAGCTCATGGCAATGCC AGATCCTTAACCCACTGAGCAAAGCCAGGGATCAAACCCAACATCTCATGGATCCTAGTCGGGTTTGTTAACCCTTGAGCTGCAAAGGGAACTCCCATAA TAATCCTTTTAAATGTTTTACTCTGTTCTGATGCATGAGACTAATATTCTTTCTGATACTCTCATTTTAGCTATAAAGTTGATTTCTTATGAAAACTCAG TAAGAGATCACTCTTTCCTTTTGATATTTAACCCCTTAATAGTAATTACCATTCATGATGACATTAATCCATAACAGATGAGAACAGTAGTTGCAGAAATGGGTAAT GGAAGAATGTGATTTCAACTAAATGTCCAATATCAAAAAAGACTAAGTGTGTTCATGAAACATCTATTTACTATTTATAGCAGTTATTCAGCTCTGCCTT TGTAGTGGTAAAGTGGTCAGACACAATACTTAAGGTAAAAGTTTCCAGTTATGAAACTTTTACTTACAAATATGGGCTGAGACTGGGCAATAGTTCAGTG ATTCCTTGGGGTAGATTCTTGGACCTCTTTTTTTAAATGTTGGACCTCTTTTTTAATGCTAAGTTATTCCTTTGAAATTAGTCTTGCTTATTTGTGTCAT TTGTATTGAAGTATACTGGTGAATTACATGTTCTGTGTATGCTGTGTGGAATTATTTGTTCCTCATTTTGCATGTAATTGTATTTGTATTGATAGG TATGGCTACCATGTTACCAAAGGTACATATTTTATCTGCCATGATCAAGGCAATGTTAAAAAATTATTTGAAAACCTTAGAAACTACGATGGGAAGAATAATTAT >PGM2_vicVic Vicugna vicugna (vicugna) tiled -L1MA9 6162-6310 24% -MER58A 35-157 20% VITAELASFLATKNLSLSQQLKAIYVE YGYHITKASYFICHDQGTVKKLFENLRNY GTCATAACTGCAGAGTTGGCTAGCTTTCTAGCAACCAAGAATTTGTCTTTGTCTCAGCAGCTAAAGGCCATTTACGTTGAGTAAGTTTCTATTAATGCTG TTTAATTGGAGTAAGCTTTTTATCCATTTCAGATGTACCACATTATGACTCAGTATACGTCTACATTGTGAAATGATCACAATTAGTAAAGTTAACGTGT ATCATCACACATAGTAATATTTTATAATGCTTTACTCTGTTCTTGTGCATGGGACACTAATGTTCTTTCTGATGCTCTTTCTTTAGTTATGAATTCTGTT TCTTATGAGAACTAGATAAGAGATCATCTTTTCCTTTTGATACCTAATCACTTAATAGTAATTACCATTCATGATGACATTAATCCTTACAAATGAGAAA ATTAGTTGCAGAAATGGCTAATGGAAGAATGCGATTTTAAGTGTCTAATGTCAAAAAAGACTAAATGTGTTCATGAAACATCTGTTTAGTCTTTATAGCA ATTACTCAACTCTACCTTTGTAGTGCAGAAGCAGCCAGACTCAACACATAAGGTAATGATGTGGCTGTTCCACTAATAAAACTTTTACTCAAAAACACTG GCTGAGGGCCAGGCAACAGTTCAGCAATCCCTGGGGTAGATAGTTGGACCTCTTTTTTTTAATTCTAAATTATTCCTTTGAAACTCATCATGGTTATTTG TGTCATTTATTTTAGAGTATACTGGTTGATGACATGTTCAGTGTACACTGTGCAGAATTCTTTGTTCCTTGTTTGCATGTAATTGTATTTGTATTGATAG GTATGGCTACCATATTACCAAAGCTTCATATTTCATCTGTCACGATCAAGGCACTGTTAAAAAATTATTTGAAAACCTTAGAAACTAC >PGM2_ateAlb Atelerix albiventris (African hedgehog) No repetitive sequences [VISAELASFLATKNLSLSQQLKAIYVE] YGYHITKASYFICHDQVTIKKLFENLRNY AB258952 TTAATGTGTTTGTTAAACATCTATTTATTCTTTACAGCTATCACTCAACTCTGACTTTGTAATACAAAATAGCCACACTTAGTCCATGAGGTCATGGACC TGATGTGACTGCCCCAATAAAACTTATACCTACAGATATAATCAAAATAAGATAAAATGGATGCTATCAATACTTAAGAATATTGGCTAAGTAAAAACAA AGAACTAGTTTAGAAACCTACAGGGGGTTATTGTTCTTCCTTTTTTTCATGCTATATTATTCCTTTGAAGCCAGTCATAGTTATTAGTCTCATTAACTTT ATAATATACTGGTTATATATGTTCTGTGTATACTAGGTAAAGTTATTTCTACCTAATTTTGCATACGTTTTATTTGTTTGCTAGGTATGGCTACCATATA ACCAAAGCTTCATATTTTATCTGCCATGATCAAGTCACCATTAAAAAATTATTTGAAAACCTTAGAAATTAT >PGM2_sorAra Sorex araneus (shrew) +SOR1_SINE +SOR1_SINE VISAELASFLATRNLSLSQQLKAIYVE YGYHITKASYFICHDQSIIKKLFENLRNY AALT01183695 AALT01470682 GTCATTAGCGCGGAGCTGGCCAGCTTTCTCGCCACCAGGAACCTGAGTTTGTCCCAGCAGCTAAAGGCCATCTATGTGGAGTAAGTTCCCTACTGACTGT GCTTAATCAAAATAACCCGTATTTTTGGATCCATTTTTAACGGTTTATTATGCTCTTGTGTGTGTGATACTGATAGTCTCTCTAATGTCCTCACTTCAGT TATAAATCCTATTTCTTAAAAACATGAAGTTAAGGGGCTGAACCGATAGCACAGCGATAGCAAGGTTTGCCTTGCATGTGACCGATCTGGGTTCGATTCC CAGCATCCCATTTGGTCCCCTGAGCACTGCCAGGAGTAATTCCTGAGTGCATGAGTCAGGAGTAATCCCTGTGCATCGCTGGGTGTCACCAAAAAAAAAA AAACCATGAAGTTAAAAAATCACCTTTTGGGGGGGGTCGGAGAGATAGTGCAGCAGTGGGTAGGGAGCTTGAGTCATTCATGGGTCACCCAGCTTCAATC CCTGGCACGCCCTGTGGCCTCCCAAGTCCCGCCAGGAGTGATCCCTGAGCTCAGAACCATAAGCAAGCCCTGAGCACCATTGGTGTGGCCCCAGAATAAA TAAATTAGAGATAGAAATCACTTTTTCATGCTTAACTACTTAATAATACTTATGATTGCCATACTCCCTAATGAATGAGATCTAATCGCAGAACTAGTTA TTAGTTAAAAGTGTGAATTTAAATGTGTAGTGTCAAAAAAATG ACCAAGATAACCAGCTTATAACTTAGACTTATAAATGACTGCTTATCAATATATCT AAGGCCAGACAGCAGTTTACTTTAGCAGTTCCTAGGATAGGTTATTGTTCCTCTTTTTTTTTTCCCTTTATTTTTTGCCCCCTAGAGATCTACCTTTTAA AAAAATATTTTTTTAATTGAATCACAATGAGATACACAGTTACAAATTGTTTCTGATTTGATTTCAGTCAGACAATGTTCAAATATCTGTCCCTTTCACA GTGTACATTTCCCACCACCAGTGTCCCCACTTTCCTTCCTTGTTCCTCTTTTTTCATGCTCAGTGATTCCTTTGAAGCGAATTATGGTCATTCGCTTCAC TTGCTTAAAAGCAAATGAATCAGCGGCCGATTGATGTCCTGTGTTCGAAAGACAGAATTCTTTGTTCCTTATTTTGCGTGTATTTGTATTGATAGGTATG GCTACCATATAACCAAAGCTTCGTATTTTATCTGTCACGATCAAAGCATCATTAAAAAGTTGTTTGAAAACCTTAGAAATTAT
Analysis of L1MA9 retroposon INT391
Extended validation of this L1MA9 insertion, which occurs in a short intron of the gene ACSL5, is feasible in May 2008 because of newly available genomes. elements. The basic technique consists of first establishing the comparative genomics of the two outside coding exons. These are needed to reliably probe contig assemblies and trace archives with tblastn and blastn respectively. A complete intron can often be obtained by tiling out to the center from the two ends. It is imperative to avoid paralogous exons in doing so.
In the case of INT391, dog, cat, horse, microbat, macrobat had the retroposon judging by location, fragment coordinates relative to the full length retroposon, and strand orientation relative to the coding exons (minus strand here). Cow, dolphin, pig, vicugna, and shrew did not have it. Since cetartiodacytl L1MA9s might have been interrupted by a later retroposon breaking the L1MA9 into two unrecognizable shorter pieces, it is necessary to remove other repeats and re-run RepeatMasker.
Despite more intensive phylogenetic sampling, INT391 continues to support Pegasoferae as Nishijimi et al originally stated. It should be noted that the MER-class retroposon, while not at issue here, exhibits the type of homoplasy that makes retroposons dicey as tree topology markers. Introns are often susceptible to multiple insertions of similar retroposons as well as to complicated patterns of micro deletions that prevent their recognition even if they aren't fully deleted.
Summary of the phylogenetic distribution of the L1MA9 retroposon INT391: >INT391_ACSL5_Peg_canFam +MER91 85-140 23%9 -L1MA9 6082-6298 24% span 762bp >INT391_ACSL5_Peg_felCat -MER91C 97-140 27% -L1MA9 6082-6301 22% >INT391_ACSL5_Peg_equCab -MER91B 62-128 26% -L1MA9 6082-6302 21% >INT391_ACSL5_Peg_myoLuc no MER -L1MA9 6079-6277 23% >INT391_ACSL5_Peg_pteVam -MER91B 8-62 24% -L1MA9 6060-6302 25% >INT391_ACSL5_Peg_bosTau -MER91C 55 -85 28% No L1MA9 >INT391_ACSL5_Peg_turTru -MER91 261-311 22% No L1MA9 >INT391_ACSL5_Peg_susScr -MER91 257-306 8% No L1MA9 >INT391_ACSL5_Peg_vicVic -MER91 284-337 24% No L1MA9 >INT391_ACSL5_Peg_sorAra no MER91 No L1MA9 Markup of exons and intronic retroposons of INT391 within ACSL5: blue: coding exons magenta: L1MA9 INT391 red: MER91 retroposon >INT391_ACSL5_Peg_canFam +MER91 85-140 23% -L1MA9 6082-6298 24% span 762bp GDPKGAMLTHQNIISNVSSFLKCME YTFKPTPEDVTISYLPLAHMFERIVQ ACAGGTGACCCTAAAGGAGCCATGCTGACCCATCAAAATATTATTTCAAATGTTTCTTCTTTCCTCAAATGTATGGAGGTCAGTGGTCAATTGTCAAGGA GGTCTTCATTAAAATGTAAATCTGTCATAAGATTTTAATCCTGATGTAAGAGGAGTCAGAGACTAACACAAAACAAAACAAAAACAAAACTCATGATAAA GGCCTGAAGAAGGGACAAATAGTGGTGTCTCTTTGTCCAGAGGACTGTGCATTTTCAAGCCTTGGCCTTTTAGAATCACTGCACATCTCTACACTCAGTG AAATTAAGGggcacctctcagagttatacagtgcaccacctgtacaactgggtgtggcagtcctgGGAAGGAGCAGTTTTTTTTAAATTAAAGAAAAAAT Tttgagatacaattaacataacactatattaatttcagatacacaacataatgatttcatatatatgttgcaaaatggttcccacaataaatctaacatc cattatcacacatagctatagtttctttttcttgtgatgagaatttttaagatctgctcacttactaacttgcagatatgcaatacagtattattaacta tagttaACGGGAGTTACTTTTAAGTCTCCTTCGGAAGAGAAAGTTGGCATTAACACAATGTCTCCTCCTTGTTCTAATCTACAGTATACTTTCAAGCCCA CCCCTGAAGATGTGACCATATCCTACCTGCCCTTGGCTCATATGTTTGAGAGGATTGTACAG >INT391_ACSL5_Peg_felCat -MER91C 97-140 27% -L1MA9 6082-6301 22% GDPKGAMLTHENIVANSSAFLKCME CIFKPTTEDVSISYLPLAHMFERIVQ GGTGACCCTAAAGGAGCCATGTTGACCCATGAAAATATTGTTGCAAACAGTTCTGCTTTTCTCAAATGTATGGAGGTCAGTGGTCAATTTAAAAAGAGGT AGTCATTAAAATGTAAATCCATCATAAGATTTTGATCTTGATGTCAGAGGAGGCAGAGACAAAAAACAAAACAAAACCAAAAGCCACGTTAAAGGCCTGA CAATGAATCAGTGTGGACAAATACTGGTGCATCTTTGTCCAGAGGACTGTGCATTTTCCAGCCTTGGTCTCTTAGAATCACTGCATGTATCTACACTCAG TGAAGTTAAGGAGCACCTTAACTTCagtcatacagtgcaaaacctgtgcaactatgtgtggcaatcctgGCAATTTCTTTAAAAGTAAAGAAAAAAAttt gttgagatattattgacgtattaatttcaggtgtacaacgtgattccatatatgtatgtactgcaaaatggtccctgtgataaattccaagtccatcaac acacataattttttttcttgtgatgagaacttttcagatctactcacttaacaactttcaaatctgcaacacagcattattaactgtagttaATAGGAGC TGCTTTTAAATCTCCTTTAGAATAGAAAGTTAGCACTAATCCAATGGTGTCTCTTTCTTGTTCTGGTCTATAGTGTATTTTCAAGCCCACCACTGAGGAT GTGTCCATTTCCTACCTCCCCTTGGCTCATATGTTTGAGAGGATTGTACAG >INT391_ACSL5_Peg_equCab -MER91B 62-128 26% -L1MA9 6082-6302 21% GDPKGAMITHQNITSNTAAFLRSME GTFEINLEDVTISYLPLAHMFERVVQ GGTGACCCCAAAGGAGCCATGATAACCCATCAAAATATTACTTCAAATACTGCTGCTTTTCTTAGATCTATGGAGGTCAGTGATCAATTGAAAAAGAGGA ATTCCTAATTAAATTTCAATTGAAAATTCCTAATTAAAATAGGAATCTGCCATAAGATTTTAATCTTGAAATTAGAGAAGGCATAGAGGAAAAAAATAGG TTTAAGGCCTAAGTATGCACACATATCAGTGCCTCTTTGTCCAGAGGACTGTGCATTTTCACGTCTTGGTCTTTTAGGATCACTGCAGAGCTCTACACTC TGTGCAgttaagggtacctcttacagttgtacagtacatcacctgcacaaccatatgtggcagttctgGGAAGGAGTAGttttttaaaaattaaaaaaat attttattgagatatgattgacatataacattatgctagtttcagatgtacaacataatgatttgaggtttgggtatattgcaaaatgatccccacaata agtctagttaacatccatcaccacgcatagttacaaattttttcttgtgatgaaaacgtttaagatctactctcttagcaaatttctaatatataataca gtattactaactagaattaATAGTAGTTTTTAAATCTCCTTCGAAGAGAAAGTTGGATTAATACAATGTTGTCTCCTCTTTGTTCCCTGATCTGTAGGGT ACTTTTGAGATCAACCTTGAGGATGTGACCATATCCTACCTCCCCTTGGCTCATATGTTTGAAAGGGTTGTACAG >INT391_ACSL5_Peg_myoLuc no MER -L1MA9 6079-6277 23% GDPKGAMLTHQNVVSNASAFLRCVE ESFAPTPEDVSISYLPLAHMFERVVQ AAPE01034117 GGTGACCCCAAAGGAGCCATGCTAACCCATCAAAATGTTGTTTCAAATGCTTCAGCTTTCCTCAGATGCGTGGAGGTTAGTGGTAGCTTGAAAAAGAGGT CTTCGTTAGAATGTGACTCTGTCATAAGATTTTAATCTTGAAGCTAGAGGAGGCAGAGAAGAAAAAAACCAAAACAGGTTAAGGGCCTGAGTGTGGACAA ACACATGTGCATCTTTGTGTGGAGGGCTGTGCATTTTCAAGCCGTGATCTTTGAGGATCCCTGCAGACCTCTACTCCAGCGCAGTCCAGGGCACCTCTCC CAGTTCTTCAGGGCACCCCCTGCATGACTGTATGGGGCACTCATGGAAGGAAATAGTTAAAAAAAAATTTAAATTTTAAATGAGATGTAACGATGCCTaa cattataatagtttcaggtgtgcaacataatgattcaatatttatatgtattgcaaaatgatcctcatagtaagtgtagttaatatccatcactgcacac agttacaaattctttgttcttgtgatcagaacttctaagatcaactctctcagcaactttcgaatatacaatagagtgttattaactatagttaacaAGG GTAGTTCTTAAATCTCTTTGGTAAAGAAGGTTGGCATTAATCCGATTTTGTCTCCTCCCCCTTCCCGATCTGTAGGAAAGCTTTGCACCCACCCCCGAGG ATGTGAGCATATCCTACCTCCCCTTGGCTCATATGTTTGAGAGGGTTGTACAG >INT391_ACSL5_Peg_pteVam -MER91B 8-62 24% -L1MA9 6060-6302 25% GEPKGAVLTHQNVISNAAAFLKLLEVS DSFQVTPKDVTISYLPLAHMFERIVQ ti|1386642117 ti|1371644127 GGTGAGCCCAAAGGGGCCGTGCTAACCCATCAAAATGTCATTTCAAATGCTGCTGCTTTTCTCAAACTTTTGGAGGTCAGTCGATCAAATGAAAAAGAAG TCCTGATCAAAATGTGAATTTGTCATAAGATTTTAATCTTGAAGTCAGAGGAGGCAGAGAGGGGGAAAAAAAACAGGTTAAGGGCCTGAATGTGGGCAAA TATTTGTGCATCTTTGTCTGGAGGACTGTGCATTTTCAAGCCTTGGTCTTTTAGGATCACTGCAGACCTTTGTACTCAGTTAAGGGCACCTCTTAGAGTG ATGCAGTGTACCGCCCGCACAACTGTATGTGGCCCACCTAGAAAGAAGTAGCTTAAATTTTTTAAAAATTTTAATTGAGATATAATTGATATCTAACATT GCCTTAGTTTCAGGTGTACAATGTAATGATTCAATATTTGTATATGTTGCTAAACGATCCTCAAAATAAGTCTAGCTAAGAAAGATCACCACACTTAGAT AAAAACTCTTTTTTTGTGTGTGACAAGAACTTTTAGCAACTTTCATTATTAACTGTCGTTAACAGGGTAGTTCTTAAATCTCCTTTGGAAGAGAAAGTTG GCATTAATCCAATGTCATTTCCTCTTTGTTCTTTATCTATAGGACAGCTTCCAGGTCACTCCCAAGGATGTGACCATATCCTACCTCCCCTTGGCTCATA TGTTTGAGAGGATTGTACAGGTGAGT >INT391_ACSL5_Peg_bosTau -tRNA-GluSine -MER91C 55-85 28% No L1MA9 GDPKGAMLTHANIVSNASGFLKCME GVFEPNPEDVCISYLPLAHMFERIVQ GGTGATCCCAAAGGAGCCATGTTAACCCATGCAAATATTGTTTCCAATGCTTCTGGTTTTCTCAAATGTATGGAGGTCAGTGGTCAATTGAAAACAAGGC CCTCATTAAAATGTAAATCTGTCGTAAGATTTTAATCTTAAAGTGAGAGGAGGCAGAGAGGGAAAAAACTGATTGAAGGCCTGAGTGTGGATGAATACCA GTACATCTTTGTCTGGAGTTTTGCCCTTTTATTTATTTATTAatatatatatatatatatatatatTTTTTAATCTGGACCATTTTTAAAGTTTTTATCG AATGTGTTATAGTATTGGTTCTGTTTTATGTTTTGATTTTTGGGGGGCTACAAGgtacatgggatctcagctccctgaccaggggtagaactcacaccct ctgcattggaaggtgaagtcttaaccactggacctctggggaagtccCATAGAGTTTTGCTGTGTTAGGGTCACTGCAGATCTCCACACTCAATGCAGTT AGAgcagcccttagatttacacagggcacatctgcacagctgtatgcagcagtcctAGAAAGAAGTGTTTAAATCCTCTTTGGAAGAGGAAATTGACATT AACCCATTGTTGTCTCTTTTCCATTTCCTGATCTCTAGGGTGTTTTTGAGCCCAATCCTGAGGACGTGTGTATATCCTACCTCCCCTTGGCTCATATGTT TGAAAGGATTGTACAG >INT391_ACSL5_Peg_turTru -MER91 261-311 22% No L1MA9 GDPKGAMLTHENIVSNAAAFLKCVE HTFEPSSEDVTISYLPLAHMFERVVQ GGTGACCCCAAAGGAGCCATGTTAACCCATGAAAATATCGTTTCAAATGCTGCTGCTTTTCTCAAATGTGTGGAGGTCAGTGGTCAATTGAAAAGGAGGC CCTCGTTAAAATGGGAATCTGTCATAAGATTTTAAAGTTAGAGGAGGCAGAGGGGGAAGAAACAGGTTGAAGGCCTGAGTGTGGACAAATACTGGTGCAT CTTTGTCTAGAGTTTTGCTCTTTTAGGGTCACTGCAGATCTCTGCACTCAGTGCAGTTAGGGCACCCCTTAGGGCACAGTGCACACCTGTACAACTGTAT GCAGCAGTCCTAGAAAGAAGAAGTGTTTAAATCTTCTTTGGAAGAGAAAGTTGGCATTAATCCACTGTTGTCTCCTTTCCATTTCCTGATCTATAGCATA CTTTTGAGCCCAGTTCTGAGGACGTGACCATATCCTACCTCCCCTTGGCTCATATGTTTGAGAGGGTTGTACAG >INT391_ACSL5_Peg_susScr -MER91 257-306 8% No L1MA9 GDPKGAMITHQNIVSNVASFLKRLE YTFQPTPEDVSISYLPLAHMFDRIVQ ti|2023263948 GGTGACCCCAAAGGAGCCATGATAACCCATCAAAATATTGTTTCAAATGTTGCTTCTTTTCTCAAACGTCTGGAGGTCAGTGGTCGACTGAAAAAGAAGC CCCTGTTGAAATGTGAATCTGTTATAAGATTTTAAAGTTAGAGGAGGCAGAGAGGAAAGAACCAGGTCAAAGCCCCAAGTATGGGAAAATACTAGTGCAT CTTTGGAGTTTTGCTCTTCTAGGGTCACTATAGATCTCTACACTCAGTGTAATTAGGGCACCCCCCAGAGTTGTGCAGTGCACACCTGCACAACTGTATG TGGCAGTACTAGAAAGTAGTGTTTAAATCTTCTTTGGAGGAAAAAGTTGGCATTAATCCATTGTTGTCTCCTTTCCCTTTCCTGATCTACAGTACACTTT TCAGCCCACCCCTGAGGACGTGTCCATATCCTACCTCCCCTTGGCTCATATGTTTGATAGGATCGTACAG >INT391_ACSL5_Peg_vicVic -MER91 284-337 24% No L1MA9 GDPKGAMITHENVVSNVAAFLKFME YSFEPTPEDVAISYLPLAHMFERVVQ ti|1970855441 GGTGACCCCAAAGGAGCCATGATAACCCATGAAAATGTTGTTTCAAATGTTGCTGCTTTTCTCAAATTTATGGAGGTCAGTGATCAACTGAAAAAGACAC CCTCGTTAAAATGTGAATCTGTCATAAGACTTTAATCTTCAGGTTAGAGGAGGCAGAGAGGGAAAATGACAGGTTTAAAGCCTGAGGGTTGACAAAGACT GGTGCATCTTTGTCTGGAGGACTGTGCGTTTCCAAGTTTTACTCTTAAGAATCACTGCCGGTCTCTCCACCCAGTGCAGTTAGGGCATCTCTTAGATTTG CGCAGTGCACACTTGTGCAACTGTATGTGGCGGTCCTAGAAAGAAGTAGTGCTTAAATCTTCTTTGGAAGAGAAAGTTGGCATTAATCGAATGTTGTCTT CCTCCCATTCCCTGATCTCTAGTATTCTTTCGAGCCCACCCCTGAGGATGTGGCCATATCCTACCTCCCCTTGGCTCATATGTTTGAGAGGGTTGTACAG >INT391_ACSL5_Peg_sorAra -SOR1SINE No L1MA9 WGPKGAKITHEILSSKAZAFLNSVE YAFEPTPEDVSISYLPLAHMFERVVQ AALT01576933 GGGCCTAAGTGGTGCTGAGGATGGAACCCAGGCCTTCTGCAGCTCCAACCCCCTGGGCCAGCTCTCCAGCTCTAAAGTGCCCCTAATGTAAGGGGAT GCAGGAAATATGGCAGAGCTGAAGTCATGAACCCAGAAACAACAGGAGGAGGTGATGGGCTTTTCTTTGTAACTGCATCTGTGATTGTGGTCTTGTGGAA TGTCGCTGCACATTGCAAAGCCAAAGACGGGCTGTGTGCTTTATAAAGGGTCTTTCTCTCCACCTCTTGTCTCCTCCAGGTGACCCCAAAGGAGCCATGA TCACGCATGAAAATATTGTTTCAAACGCCTCTGCTTTCCTCAAGTGTGTGGAGGTCAGTGGATGTGGGAAAAGAGGTCCTAGCAAAAGGGTGGATGCCAC AAAGTTCAGAAGTGGAAGTTAGAGCAGCAGCAGGGCTGGAGGGTGGCGTTCAAAGGGCTGTGTGTGTGCAGATGCCCCGACAGCTTGGGACATCAGTGTT ATCATTATCATTATTATTATTACCATTTTGGTTTTTGGGGTACACTTGGGAATGGACAGGGGGCACTTCTGGCTTATGCACTCAGGAATTACTCCTGGTG GTGCTCAGGGAACCATGTGGGATGCTGGGAATCAAGCCACATGCAAGGCAAATGCCCTACCCACTGTGCTATTGCTCCAGTCTCATCAGTGTTTTAGGAA GCTGTGTATGTTGCTGCCTTGATATCCAGCACCTCTCTGCTCTCGGCGTGTAACAGCGCCCCTCAGAGCTCCACGGGGGGTCTAGCCTGCACACCCAGGT GTGGCCCTGCTGGAAATGCCTGGTCTTTAGGTCTTCTTTGTCTGGGGAAATTTGGCATTGATCGATGGTCTCTTTCCTCTGTGCCCTGATCTGTAGTATG CGTTCGAGCCCACGCCTGAGGATGTGAGCATCTCCTACCTCCCCTTGGCACACATGTTTGAGAGGGTCGTGCAG
Phylogenetically informative coding insertions and deletions
introduction
Analysis of a potentially informative indel in CHML
A loss of one amino acid can be observed within an exon of the gene CHML in carnivores, bats, and horse but not in other Laurasiatheres (or other placental mammals. This gene is similar enough to a second gene CHM that it too needed full-on phylogenetic annotation -- it has no indel in any species so must not be cross-annotated for CHML in species without assemblies. The key to that is including upstream amino acids where sufficient divergence between the two genes resides.
The background story here is a common one in gene dosage compensation. In the amniote ancestor and birds and lizards today there is but one multi-exonic gene CHM. In mammals, as the sex chromosomes underwent upheaval, the ortholog ended up on chrX which has implications for reduced levels of expression compared to an autosomal gene. That may have favored persistence of a retroprocessed (intronless) copy on chr1. In marsupials, the two copies remained quite similar. In placentals, the second gene CHML diverged extensively from the first while remaining well conserved within its orthology class.
The one-residue indel appears cleanly restricted to Pegasoferae. There are 5 Laurasiatheres with it and 5 without it, which needs further buttressing by PCR (adding more basal species has the effect of localizing the event farther back on the stem). The parental gene did not develop the indel in any bioinformatically accessible species.
Interestingly, the retrogene CHML landed within intron 1 of encephalopsin OPN3 with the same direction of transcription. This raises some questions about how CHML is translated (as it would apparently be spliced out). Encephalopsin has an interesting history in mammals involving multiple independent losses.
The region annotated below corresponds to a known domain called GDI (for GDP dissociation inhibitor: pfam00996). The 3D structure of this region is available in the parent gene in rat, PDB 1LTX so the structural location of the indel could be determined. Very likely it lies within a loop and has minimal functional impact.
CHML chr1:239,864,567-239,864,746 gene_genSpp AFRQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLNATKNFLQCLGRFGNTPF CHML_homSap Homo sapiens (human) AFRQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLNATKNFLQCLGRFGNTPF CHML_panTro Pan troglodytes (chimp) AFRQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLNATKNFLQCLGRFGNTPF CHML_ponPyg Pongo pygmaeus (orang_sumatran) AFRQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLNAIKNFLQCLGRFGNTPF CHML_macMul Macaca mulatta (rhesus) AFRQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLNATKNFLQCLGRFGNTPF CHML_calJac Callithrix jacchus (marmoset) AFEQCLFSEYLKTKKLTPNLQHFILHSIAMTSESSCTTIDGLKATKNFLQCLGRFGNTPF CHML_otoGar Otolemur garnettii (bushbaby) AFKQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTVDGLKATKNFLQCLGRFGDTPF CHML_micMur Microcebus murinus (mouse_lemur) AFEQCLFSEYLKTKKLTPNLRHFILHSIAMTSESSCSTLDGLKATKTFLQCLGRFGNTPF CHML_tupBel Tupaia belangeri (tree_shrew) DFKQCSFSDYLKTKKLTPNLQHFILHSIAMTSESSCTTLDGLQATKTFLQCLGRFGNTPF CHML_musMus Mus musculus (mouse) DFKQCSFSDYLKTKKLTPNLQHFILHSIAMSSDSSCTTLDGLQATKNFLRCLGRFGNTPF CHML_ratNor Rattus norvegicus (rat) DFQQCLFSEYLKTKRLTPNLQHFILHSIAMTSESSCTTLDGLKATKNFLQCLGRFGNTPF CHML_cavPor Cavia porcellus (guinea_pig) DFKQCSFSEYLKAKKLTPNLQHFVLHSIAMTSETSCTTLDGLKATKIFLQCLGRFGNTPF CHML_oryCun Oryctolagus cuniculus (rabbit) DFKQCSFSEYLKTKKLTPNLQHFILHSIAMTSESSCTTLDGLRATKNFLQCLGRFGNTPF CHML_ochPri Ochotona princeps (pika) AFVHCSFSDYLKTKKLTPNLQHFVLHSIAMT-ESSCTTIDGLKATKNFLRCLGRFGNTPF CHML_canFam Canis familiaris (dog) AFMQCSFSEYLKTKKLTPNLQHFVLHSIAMT-ESSCTTIDGLKATKNFLQCLGRFGNTPF CHML_felCat Felis catus (cat) AFMQCSFSEYLKTKKLTPNLQHFVLHSIAMT-ESSCTTIDGLKATKNFLQCLGRFGNTPF CHML_equCab Equus caballus (horse) DFTQRPFSEYLKTQKLTPNLQHFILHSIAMT-EPSCLTVDGLKATKHFLQCLGRYGNTPF CHML_myoLuc Myotis lucifugus (microbat) DFTQCSFSEYLKTKKLTPNLQHFVLYSIAMT-ESSCTTVDGLKAAKNFLRCLGRFGNTPF CHML_pteVam Pteropus vampyrus (macrobat) AFTQCSFSEYLKTKNLTPSLQHFILHSIAMMSESSCTTVDGLKATKTFLQCLGRFGNTPF CHML_bosTau Bos taurus (cow) AFTQCSFSEYLKTKKLTPSLQHFVLHSIAMMSESSCTTIEGLKATKNFLQCLGKFGNTPF CHML_turTru Tursiops truncatus (dolphin) DFMQCSFSEYLKAKKLTPSLQHFVLHSIAMTSESSCTTIDGLKATKNFLQCLGRFGNTPF CHML_susScr Sus scrofa (pig) AFVQSSFSEYLKTKKLTPNLQHFVLHSIAMMSESPCTTIDGLKATKNFLQCLGRFGNTPF CHML_vicVic Vicugna vicugna (vicugna) AFVQSSFSEYLKTKKLTPNLQHYILHSISMTSESSCTTLDGLKATKKFLQCLGRFGNTPF CHML_eriEur Erinaceus europaeus (hedgehog) AFIQCSFSDYLKTKKLTPNLQHFILHSIAMTPEASCSTVDGLKATKIFLQCLGRFGNTPF CHML_sorAra Sorex araneus (shrew) TFKQCSFSEYLKTKRLTPNIHHFVLHSIAITSQSSCTIIDGLKATKTFLWCLGWFSKNPF CHML_dasNov Dasypus novemcinctus (armadillo) AFEQCSFSEYLKTKKLTPNLQHFILHSIAMTSQSSCTTLDGLKATKNFLQCLGRFGNTPF CHML_choHof Choloepus hoffmanni (sloth) AFKQCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLKATKNFLQCLGRFGNTPF CHML_loxAfr Loxodonta africana (elephant) AFKHCSFSEYLKTKKLTPNLQHFVLHSIAMTSESSCTTIDGLKATKTFLQCLGRFGNTPF CHML_proCap Procavia capensis (hyrax) ... CHML_echTel Echinops telfairi (tenrec) AYEESTFSEYLKTQKLTPILRHFVLHSIAMASETSTSTLDGLRGTKNFLQCLGRYGNTPF CHML_monDom Monodelphis domestica (opossum) ... CHML_ornAna Ornithorhynchus anatinus (platypus) AQKECTFSDYLKTQKLTPNLQHFILHSIAMVSEVNCCTIDGLKATQRFLQCLGRYGNTPF CHML_anoCar Anolis carolinensis (lizard) NYKNSTFAQFLKTRKLTPSLQHFILHSIAMVSEKDCNTLEGLQATRKFLQCLGRYGNTPF CHML_galGal Gallus gallus (chicken) CHM chrX:85,097,861-85,098,040 genSpp GYEEITFYEYLKTQKLTPNLQYIVMHSIAMTSETASSTIDGLKATKNFLHCLGRYGNTPF CHM_homSap Homo sapiens (human) GYEEITFYEYLKTQKLTPNLQYIVMHSIAMTSETASSTIDGLKATKNFLHCLGRYGNTPF CHM_panTro Pan troglodytes (chimp) GYEEITFYEYLKTQKLTPNLQYIVLHSIAMTSETTSSTMDGLKATKNFLHCLGRYGNTPF CHM_ponPyg Pongo pygmaeus (orang_sumatran) GYEDITFYEYLKTQKLTPNLQYIVLHSIAMTSETASSTIDGLKATRNFLHCLGRYGNTPF CHM_macMul Macaca mulatta (rhesus) GYEEITFCEYLKTQKLTPNLQYIVLHSIAMTSQTASSTIDGLKATKNFLHCLGRYGNTPF CHM_calJac Callithrix jacchus (marmoset) ... CHM_otoGar Otolemur garnettii (bushbaby) ... CHM_micMur Microcebus murinus (mouse_lemur) AYEEITFSEYLKTQKLTPNLQYFVLHSIAMTSELASSTLDGLKATKNFLRCLGRYGNTPF CHM_tupBel Tupaia belangeri (tree_shrew) AYEETTFSEYLKTQKLTPNLQYFVLHSIAMTSETTSSTVDGLKATKKFLQCLGRYGNTPF CHM_musMus Mus musculus (mouse) AYEGTTFSEYLKTQKLTPNLQYFVLHSIAMTSETTSCTVDGLKATKKFLQCLGRYGNTPF CHM_ratNor Rattus norvegicus (rat) AYETITFSEFLKTQKLTPNLQYFVLHSIAMTSETTSSTIDGLKATKNFLHCLGRYGNTPF CHM_cavPor Cavia porcellus (guinea_pig) GYEEIAFSEYLKTQKLTPNLQYFVLHSIAMTSETTTTTLDGLKATKNFLHCLGRYGNTPF CHM_oryCun Oryctolagus cuniculus (rabbit) ... CHM_ochPri Ochotona princeps (pika) AYEEITFSEYLKTQKLTPNLQYFVLHSIAMTSETASNTIDGLKATKNFLHCLGRYGNTPF CHM_canFam Canis familiaris (dog) ... CHM_felCat Felis catus (cat) AYEEITFSEYLKTQKLTPNLQYFVLHSIAMTSETASSTIDGLKATKNFLHCLGRYGNTPF CHM_equCab Equus caballus (horse) AYEDITFSEYLKTQKLTPNLQHFVLHSIAMTSKTTSSTIDGLKATRKFLHCLGRYGNTPF CHM_myoLuc Myotis lucifugus (microbat) AYEEITFSEYLKTQKLTPNLQYFVLHSIAMISERASSTIDGLKATKNFLCCLGRYGNTPF CHM_pteVam Pteropus vampyrus (macrobat) AYEEITFSEYLKTQKLTPNLQYFVLHSMAMTSETGSSTIDGLKATKNFLHSLGRYGNTPF CHM_bosTau Bos taurus (cow) AYEEITFSEYLKTQKLTPNLQYFVLHSIAMTSETASSTIDGLRATKNFLHCLGRYGNTPF CHM_turTru Tursiops truncatus (dolphin) ... CHM_susScr Sus scrofa (pig) AYEEISFSEYLKTQKLTPNLQYFVLHSIAMTSETASSTIDGLKATRNFLHCLGRYGNTPF CHM_vicVic Vicugna vicugna (vicugna) ... CHM_eriEur Erinaceus europaeus (hedgehog) AYEKMTFSEYLKTQNLTPNLQYFVLHSIAMASETGSSTIDGLKATKNFLQCLGRYGNTPF CHM_sorAra Sorex araneus (shrew) ... CHM_dasNov Dasypus novemcinctus (armadillo) ... CHM_choHof Choloepus hoffmanni (sloth) ... CHM_loxAfr Loxodonta africana (elephant) ... CHM_proCap Procavia capensis (hyrax) AYEEITFSEYLKMRKLTPNLQYFVLHSIAMTSETTSTTIDGLKATRNFLHCLGRYGNSPF CHM_echTel Echinops telfairi (tenrec) AYEDCTFSEYLKTQRLTPNLQHFVLHSIAMVSETSTSTLDGLRETRNFLQCLGRYGNTPF CHM_monDom Monodelphis domestica (opossum) ... CHM_ornAna Ornithorhynchus anatinus (platypus) AQKECTFSDYLKTQKLTPNLQHFILHSIAMVSEVNCCTIDGLKATQRFLQCLGRYGNTPF CHM_anoCar Anolis carolinensis (lizard) NYKNSTFAQFLKTRKLTPSLQHFILHSIAMVSEKDCNTLEGLQATRKFLQCLGRYGNTPF CHM_galGal Gallus gallus (chicken)
Analysis of indel 2
etcetc