Phospholipases PLBD1 and PLBD2
Introduction
A surprising number of orphan human enzymes (unknown substrate) still exist ten years after the completion of the human genome project. PLBD1 and PLBD2 are semi-orphans in the sense of being probable phospholipases of B class but with uncertain physiological substrates and thus functionalities. This is especially important in the case of PLBD2 which localizes to the lysosome, as its absence could plausibly lead to a serious yet unrecognized lysosomal storage disease.
No bioinformatic algorithm or experimental protocol leads with any certainty to determination of function. The gene pair here has seven targeted publications but cases exist where protein function remains unknown after ten thousand papers (eg PRNP).
PLBD1 and PLBD2 constitute a small gene family (sequence homology class) within vertebrates though one that occurs expanded in some early diverging eukaryotes. However, the Pfam clan (NTN: N-terminal nucleophile aminohydrolases) may have, among its ten family members, additional representatives in humans diverged beyond recognizability in primary sequence. These establish the great antiquity of the fold and certain of its features but are not likely to shed additional light on phospholipases specifically.
PLBD2 presents a special difficulty in that a sequence of post-translational steps are apparently necessary for its activation. Without these, potential substrates can hardly be assayed. These steps include removal of the signal peptide, mannosylation appropriate to the lysosome targeting receptor, and self-catalytic proteolytic activation (into 28k and 42k fragments which remain associated) to expose the substrate binding site as this becomes appropriate.
Because PLBD1 and PLBD2 are full length paralogs, the bioinformatic approach below considers both on an equal footing. PLBD1 has been more amenable to activation whereas PLBD2 has a high-resolution structural determination. Thus comparative genomics allows for annotation transfer, first from PLBD2 to a structural model for PLBD1 (already provided by the SwissModel pipeline), then perhaps transfer of PLBD1 experimental successes to PLBD2.
However the gene duplication event occurred some 650 million years ago and the two genes are quite diverged today. It's not known whether substrates have diverged or merely cell type of expression. (Increased gene dosage is seldom an explanation.) Yet certain core features remain conserved, including the fold, active site residues, signature motifs, certain glycosylation sites and even the fragmentation pattern, implying these are essential functional features under long-range strong selective pressure for their maintenance.
Disulfides are only separately conserved within each paralog but this fortuitously provides a reliable signature for assigning deeply diverged proteins from early eukaryotes to their orthology class. As the respective functions become known, we can hope to better understand how the gene duplication event contributed advantageously to increasing evolutionary complexity, leading to persistence of both enzymes in most species over immense time spans.
Conservation at critical sites
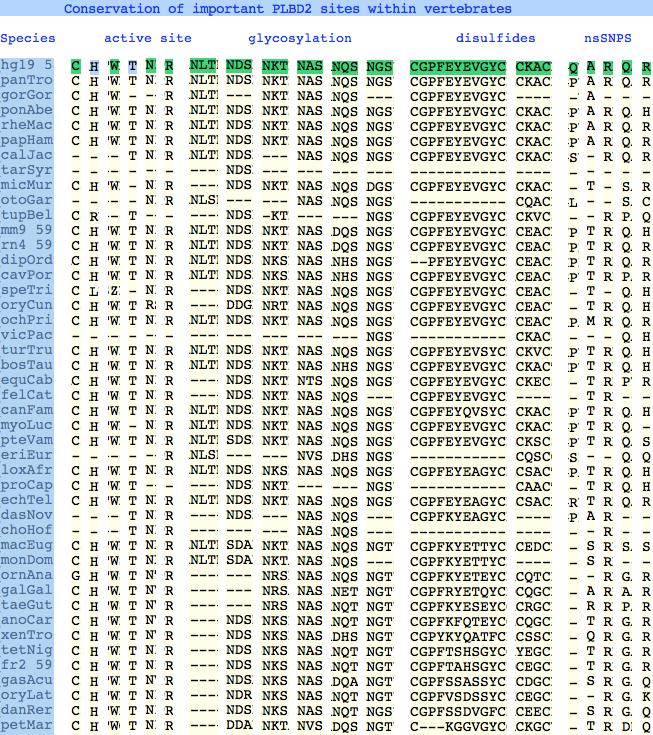
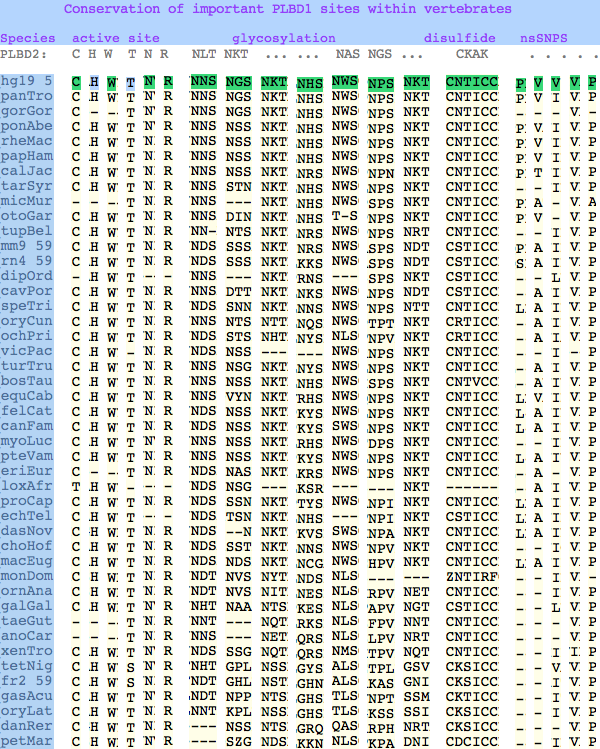
The six residues of PLBD2 associated with the active site are completely conserved within vertebrates to within genomic sequencing error. These same six residues are also completely conserved within PLBD1. Indeed 3 of the residues are conserved in the broader NTN hydrolase clan.
This is perhaps unsurprising since the active site was established a couple billion years earlier in the bacterial ancestor. However if PLBD2 and PLBD1 have different substrates, this establishes that these six residues are insufficient to distinguish the two active sites. Note H266 and T330 do not contribute their side chain, leaving them and W269 to separate phospholipases from the other NTN hydrolases.
The glycosylation sites are surprisingly conserved both within and between PLBD2 and PLBD1. Some of the motifs may be either recently acquired within later vertebrates or spurious glycosylation motifs with N and D both acceptable (or similar small amino acids) in the first slot of the NxS/T motif. Glycosylation is important in correct targeting of lysosomal proteins, more so than in generic endoplasmic reticulum proteins where motifs are often poorly conserved (as in sulfatases).
PLBD2 has two established disulfides. Strict sequence conservation of these throughout vertebrates (indeed, throughout metazoa) suggests both play an important role in protein structure and stability.
In PLBD1 however, the first disulfide is not a possibility and while an opportunity exists for a disulfide homologous to the the second disulfide of PLBD2, indels cloud the alignment and spacing would have to be different. There is additionally ambiguity given C...CC as to the cysteines involved. Indeed a second distal disulfide may occur utilizing C...CC.............C which has no counterpart in PLBD2. While cysteines can be conserved for many reasons other than disulfide (as in the nucleophile cysteine here), suitably proximity and side chain orientation in the SwissModel of PLBD1 would argue for disulfide. Comparative genomics suggests that C2 and C4 may form an ancient disulfide whereas C1 and C3 might represent a deuterostome innovation.
homSap CNTICCREDLNSPNPSPGGC human PLBD1 braFlo CSAICCRKDLAKVGAKPDGC Branchiostoma floridae strPur SKSICMRGDLM-TSPMPNGC Strongylocentrotus purpuratus XM_001192029 nemVec MNAICSRGDLIADGPRASGC Nematostella vectensis XM_001638165 monBre YNAICSRGDLESDSPSPGGC Monosiga brevicollis XM_001745398 SwissModel coordinates for PLBD1 show the 2nd and 4th sulfur atoms separated by 2.03 angstroms: ATOM 3552 SG CYS 471 49.680 -13.769 -12.461 ATOM 3579 SG CYS 475 49.273 -14.310 -4.881 ATOM 3585 SG CYS 476 51.067 -9.716 -9.172 ATOM 3678 SG CYS 490 50.737 -13.198 -5.75
The known human SNPs of PLBD2 are in some cases quite radical substitutions in terms of both physical qualities of the substituted amino acid and the degree of observed phylogenetic conservation at that site. These likely result in unstable and/or inactive enzyme. Both enzymes are autosomal so compensation might occur in the recessive state, or alternately, PLBD2 and PLBD1 could fill for each other to some extent. In either case, lysosomal storage disease might not be clinically observable.
Here Q54P may actually be a mutation in the reference sequence individual (with the SNP representing wildtype) as proline is quite well conserved throughout mammals. In A204V, valine is quite a bulky substituent for a site normally restricted to small amino acids; R354C is definitely a serious mutation, no doubt attributable to a CpG hotspot; Q521K appears milder as does R524C.
The known human SNPs of PLBD1 can be analyzed similarly. P26Q and V30L may be inconsequential as they occur in the rather unconstrained primary sequence of the N-terminus; V265I occurs at an ILV reduced alphabet; V377A and P534A are much more serious despite the aliphatic nature of alanine and likely give rise to dysfunctional protein.
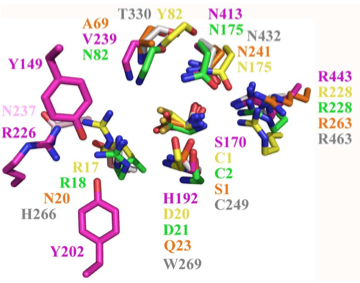
Structural superposition of active sites from five NTN hydrolases showing conserved side chains (*) and relevant main chains (....) (adapted from Fig 6 of Lakomek et al. BMC Struct Biol.2009;9:56:) * (*) * * PLBD2 phospholipase B-like gray 3FGR C244 H261 W264 T325 N427 R458 human numbering PLBD1 phospholipase B-like .... pred C228 H245 W248 T303 N402 R433 human numbering SwissModel Cephalosporin acylase pink 1OQZ S170 .... H192 .... N413 R443 Conjugated bile acid hydrolase green 2BJF C2 .... D21 .... N175 R228 Penicillin V acylase yellow 3PVA C1 .... D20 .... N175 R228 Penicillin G acylase orange 1K5S S1 .... Q23 .... N241 R263 Human SNPs resulting in amino acid substitutions: PLBD2: PLBD1: Q54P rs7965471 P26Q rs1141509 A204V rs12231990 V30L rs12296104 R354C rs56935204 V265I rs7957558 Q521K rs17852787 V377A rs2287541 R524C rs12425042 P534A rs1600
Intron evolution
PLBD1 and PLBD2, being full length paralogs, clearly indicate an early gene duplication and subsequent divergence to the current low percent identity. Segmental duplications preserve any introns present at the time of the event and these generally persist in both position and phase into living species.
However PLBD1 and PLBD2 -- despite having similar numbers of introns -- exhibit very little in common in terms of location as the diagram below shows. One possibility is that a second copy arose as a retroprocessed gene (a mechanism erasing existing introns) and was subsequently intronated at random positions. This is unlikely here given that 10-11 relatively rare events would be needed.
The remaining possibility is that the gene duplication took place prior to the main era in early eukaryotes during which the bulk of introns were established. This fits the current state of high divergence despite fairly slow rates of evolution during metazoan times.
The last five amino acids of each PLBD1 exon are colored below. Then using an alignment of PLBD1 to PLBD2, the colors are mapped to the homologous five residues within PLBD2. There they fall on the ends of exons only when these correspond to those of PLBD1. The outcome here -- despite uncertainties in alignment gapping -- shows intron positions do not correspond with the exception of the terminal intron (which also is phase 0).
While this merely compares human PLBD1 and PLBD2, the collected reference sequences (intronated against their respective genome assemblies) confirm that introns in both genes are deeply conserved.
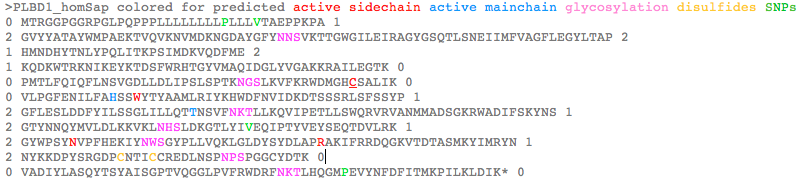
PLBD1 introns do not correspond well to those of PLBD2: >PLBD1_homSap Homo sapiens (human) first and last introns are not mappable 0 MTRGGPGGRPGLPQPPPLLLLLLLLPLLLVTAEPPKPA 1 2 GVYYATAYWMPAEKTVQVKNVMDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQTLSNEIIMFVAGFLEGYLTAP 2 1 HMNDHYTNLYPQLITKPSIMDKVQDFME 2 1 KQDKWTRKNIKEYKTDSFWRHTGYVMAQIDGLYVGAKKRAILEGTK 0 0 PMTLFQIQFLNSVGDLLDLIPSLSPTKNGSLKVFKRWDMGHCSALIK 0 0 VLPGFENILFAHSSWYTYAAMLRIYKHWDFNVIDKDTSSSRLSFSSYP 1 2 GFLESLDDFYILSSGLILLQTTNSVFNKTLLKQVIPETLLSWQRVRVANMMADSGKRWADIFSKYNS 1 2 GTYNNQYMVLDLKKVKLNHSLDKGTLYIVEQIPTYVEYSEQTDVLRK 1 2 GYWPSYNVPFHEKIYNWSGYPLLVQKLGLDYSYDLAPRAKIFRRDQGKVTDTASMKYIMRYN 1 2 NYKKDPYSRGDPCNTICCREDLNSPNPSPGGCYDTK 0 0 VADIYLASQYTSYAISGPTVQGGLPVFRWDRFNKTLHQGMPEVYNFDFITMKPILKLDIK* 0 >PLBD2_homSap Homo sapiens (human) 0 MVGQMYCYPGSHLARALTRALALALVLALLVGPFLSGLAGAIPAPGGRWARDGQVPPASRSRSVLLDVSAGQLLMVDGRHPDAVAWANLTNAIRETG 2 1 WAFLELGTSGQYNDSLQAYAAGVVEAAVSEE 0 0 LIYMHWMNTVVNYCGPFEYEVGYCERLKSFLEANLEWMQEEMESNPDSPYWHQ 0 0 VRLTLLQLKGLEDSYEGRVSFPAGKFTIKPLGFL 2 1 LLQLSGDLEDLELALNKTKIKPSLGSGSCSALIKLLPGQSDLLVAHNTWNNYQHMLRVIKKYWLQFREGPW 1 2 GDYPLVPGNKLVFSSYPGTIFSCDDFYILGSGL 0 0 VTLETTIGNKNPALWKYVRPRGCVLEWVRNIVANRLASDGATWADIFKRFNSGT 2 1 YNNQWMIVDYKAFIPGGPSPGSRVLTILEQIP 2 1 GMVVVADKTSELYQKTYWASYNIP 2 1 SFETVFNASGLQALVAQYGDWFSYDGSPRAQIFRRNQSLVQDMDSMVRLMR 2 1 YNDFLHDPLSLCKACNPQPNGENAISARSDLNPANGSYPFQALRQRSHGGIDVK 0 0 VTSMSLARILSLLAASGPTWDQVPPFQWSTSPFSGLLHMGQPDLWKFAPVKVSWD* 0
Signal peptide compositional anomaly
The first exon of both PLBD1 and PLBD2 are ill-behaved in alignments. The explanation can be see in their compositional distortion (very high GC content) that specialized masking tools such as seg and gnu recognize. Such dna manifests itself at the protein level by high levels of the amino acids, such as GPL that use those codons in the three reading frames.
Such regions are prone to repeated expansions and contractions via replication slippage. Not only do we expect such alleles in human but also that inter-species comparisons will be difficult and alignments problematic (as homology by definition is lost even if the sequences still align).
This matters very little to the mature protein since this region is trimmed off during maturation but the question still arises as to how signal peptide variations continue to be recognized efficiently by the signal receptor complex. Indeed a class of mutations could exist in which the signal peptide cannot be processed correctly and the protein never reaches the lysosomal compartment, in effect a knockout mutation.
This compositional anomaly may have caused vertebrate-wide sequencing problems. Many assemblies had difficulty sequencing back to the initial methionine and alignment programs also fell short. A set of reliable sequences could only be obtained after careful hand-curation and only then from fewer species than usual in comparative genomics.
Even then the set of first exons raises more questions than it answers as it seems to be evolving quite chaotically in fish. Mammals also exhibit a peculiar conserved insertion as placentals diverged from marsupials. And using SignalP 3.0 separately on each sequence, it emerges that the marsupial signal peptide and those of earlier diverging species are much shorter. That isn't a problem per se because signal peptide lengths are quite variable.
PLBD1 also exhibits a shift in the location of the signal peptide cleavage site over evolutionary time, crossing the boundary into exon 2 in some clades. (Since the exon break is extremely conserved, this conclusion is independent of alignment gapping.) Here again this would be functionally irrelevant since the co-processing of nascent chain takes place well after mRNA splicing. However this does provide an interesting case of homologous residues not being functionally homologous and so evolving under the different functional constraints.
PLBD1: ATGAcccgcggcggtccgggcgggcgcccggggctgccacagccgccaccgcttctgctgctgctgctgctgctgccgctgttgTTAGTCACCGCGGAGCCGCCGAAACCTGCAG
MTRxxxxxxxxxxxxxxxxxxxxxxxxxxVTAEPPKPA
MTRGGPGGRPGLPQPPPLLLLLLLLPLLLVTAEPPKPA
PLBD2: ATGGTGGGCCAGATGTACTGCTACCCCGGCAGCCACCTGGCCCGGGCGCTGACGCGGGCGCTGGCGCTGGCCCTGGTGCTGGCCCTGCTGGTCGGGCCGTTCCTGAGCGGCCTGGCGGGGGCGATCCCAGCGCCGGGGGGCCGCT...
MVGQMYCYPGSHxxxxxxxxxxxxxxxxxxxGPFLSGLAGAIPAPGGR...
MVGQMYCYPGSHLARALTRALALALVLALLVGPFLSGLAGAIPAPGGR...
Phylogenetic variation in first exon signal peptide of PLBD2:
<------ signal peptide -----------------> <---- start of 3FGW 3FGR 3FGT------------->
>PLBD2_homSap MVGQMYCYPGSHLARALTRALALALVLALLVGPFLSGLAGAIPAPGGRWARDGQVPPASRSRSVLLDVSAGQLLMVDGRHPDAVAWANLTNAIRETG
>PLBD2_panTro MVGQMYCSPGSHLARALTRALALALVLALLVGPFLSGLAGAIPAPGGRWARDGPVPPASRSRSVLLDVSAGQLLMVDGRHPDAVAWANLTNAIRETG
>PLBD2_ponAbe MVGQMYGSSGSHLA----RALALALVLALLVGPFLSGLAGAIPAPGGRWARDGPVTPASRSRSVLLDASAGQLLLVDGRHPDAVAWANLTNAIRETG
>PLBD2_rheMac MVGQMYCSSGSPLARALTRALALALVLALLVGLFLSGLAGAIPAPGGRWAHDGPVTPASRSRSVLLHAATGQLLLVDGRQPDAVAWANLTNSIHETG
>PLBD2_papHam MVGQMYCSSGSPLARALTRALALALVLALLVGLFLSGLAGAIPAPGGRWAHDGPVTPASRSRSVLLDAATGQLLLVDGRHPDAVAWANLTNAIRETG
>PLBD2_calJac MVGKMYSSPSSRLAQALTRALALALVLALLAGLFLSGLSGAIPAPGGRWARDGSVPSGSGSRSVVLDAAAGQLLLVDGRHPDAVAWANLTNAIHETG
>PLBD2_otoGar MvGPMYGSPGGRLARALTRALALALVLaLLIGLFLSCLAGAiPPPGSGRARDGLITPASRSSSVLLDATTDQLRLVDGRHPDAVAWANLSNAIHETG
>PLBD2_musMus MAAPVDGSSGGWAARALRRALALTSLLASLTGLLLSGPAGALPTLGPGWQRQNPDPPVSRTRSLLLDAASGQLRLEDGFHPDAVAWANLTNAIRETG
>PLBD2_ratNor MAAPMDRTHGGRAARALRRALA----LASLAGLLLSGLAGALPTLGPGWRRQNPEPPASRTRSLLLDAASGQLRLEYGFHPDAVAWANLTNAIRETG
>PLBD2_dipOrd MAAPPYGSRGGRPAGSLSRALV----LAVLVGLSPSGPAGAVPSPGDRWGRHKPEPPVSRSRSVLVDAASGQLRLVDGLHPGAVAWANLTNAIRETG
>PLBD2_cavPor MAAPTYVSLDGRPVRARALALA--PALCLLVGLSLGRLAGAVPAPGPRGARDGPVPAA--CRSVLLDAASGQLRLVDGLQPGAVAWANLTNAIPETG
>PLBD2_oryCun MVAPRDGCAGGRLARALALALL--------TGLLLGGLAGAAPAPGGGEQRDPPSPPASCCRSALLDAATGQLRLVDGRHPDAVAWANLTNAIHETG
>PLBD2_ochPri MAATRDSSAGCRLARVLTRALAL---LALPTGLFLSGPAGAIPVRGDGEERGRPAPSGSRCRSVLVDAESGQLRLVDGRHPAAVAWANLTNAIHETG
>PLBD2_turTru MVDPMYGCPGGRLARALTRALALALVLALLVGLFLSGLTGAIPTPRGHRGPGRPVPPASRCRSVLLDPEtGQLRLVDGRHPDAVAWANLTNAIRETG
>PLBD2_bosTau MVAPMYGSPGGRLARAVTRALALALVLALLVGLFLSGLTGAIPTPRGQRGRGMPVPPASRCRSLLLDPETGQLRLVDGRHPDAVAWANLTNAIRETG
>PLBD2_oviAri MVAPMYGSPGGRLARAVTRALALALVLALLVGLFLSGLTGAIPTPRGQRGRGMPVPPASRCRSLLLDPETGQLSLVDGRHPDAVAWANLTNAIRETG
>PLBD2_susScr MVAPMYGSPGGRLARALTRALALALVLALLVGLFLSGLTSAIPTPKGYRGSGRSVPPASRSRSVLLDTETGQLRLVDGRHPDAVAWANLTNAIHENG
>PLBD2_ursAme MAAPMYGSPGGRLARALTRALALALVLALLVGLFLSGLTGAIPISGRQWGPNGPVPPDSRSRSVLLDAETGQLRLVDGRHPEAVAWANLTNAIRETG
>PLBD2_musPut GS-GGRLARALTRALALALVLALLVGLFLSGLTGAIPISGRQWGPKGPVPPDSRSRSVLLDAETGQLRLVDGRHPDAVAWANLTNAIRETG
>PLBD2_canFam ...................................SGLTGATPVSGRRWGPSGPVPPASRSRSVRLDPQTGQFQLVDGRNPDAVAWANLTNAIRDTG
>PLBD2_myoLuc MVAPPSRSPGGRLTPALSRAPALAPGLALLAGLFLSGWTGAIPTPRDPWGPNGPVPPASRSRSVVLDARTGQLQLVDGRQPDAVAWANLTNAIHETG
>PLBD2_pteVam MVAPMDRSPGGRLAGALTRTLELTLVLAPLAGLFLSGRTSAIQTPGSRWGSEGPVSPASRSRSVLLDPQTGQLRLVDGRHPDAVAWANLTNAIHETG
>PLBD2_eriEur MVAPMCGSPGGRPARALTRALALAPALALLVGLFLSSLAGAIPPPEDNWGRNGSFPPVSRCRSVLLDSETGQLRLVDGRHPDAVAWANLSNAIHETG
>PLBD2_loxAfr MVAPVYGSPGGRLARALTQALAVALVLALLVGLFLSGLTGAISLTGHRWGPDGPAPPASRSRSVLLDTATGQLRLVDGRHPDAVAWANLTNAIRETG
>PLBD2_echTel MVATEYGSPGGRLARALTRAPALALMLALLVGLFLSGLTGAISPAGGRREPNGRVPPASSSRSALLDPATGQLRLADGRHPEAVAWANLTNAIHETG
>PLBD2_macEug mVATMYQ--GGCLALGLALGLGLVLVLSLP--------------------QPSLPPPPSRTRSVVMDSATGQLNVVEGWEAGAIAWANLTNAIAETG
>PLBD2_monDom MVATMCQ--GSSLALGLALALGLALGLR-------------------PPQPSLPPPAPSRSCSVVLDEASGQLKVVEGAQAGAVAWANLTNAIGETG
>PLBD2_anoCar MAPAWLLRFFGLALLLARSPARR------------------------PPPFPDPAAVPTRSCSVVLEPGSAALKLVNGWAPGAVAWANLTEGIRQNG
>PLBD2_galGal MAVVRALLVAAAVAAWVPGVASGP-------------------------------TPPPRSASVLLEPGSGRLRVLPGRQPAAVAWAELTDHIQAVG
>PLBD2_melGaL MAVVRALLVAAAVAAWVPGVASGP-------------------------------TPPPRSASVLLEPGSGRLRVLPGRQPAAIAWAELTDHIQAVG
>PLBD2_xenTro MGAQLLLIFMLFSLGAAQQAV---------------------------------------VSVLFDPATGNITTVEEKRVVGAVAWAELKDSILENG
>PLBD2_xenLae MAPWQLFIFSLFCVGAAQQQA--------------------------------------VVSVLFDPATGNITTVAEKKVAGAAAWAELTDSIQENG
>PLBD2_oryLat MAFRQNKTVCAKMSTFMKSLLVLGLFWGCGRAEI---------------------------RSAVIDKGSGKLTVVEGYHEGFVAWANFTNDIETSG
>PLBD2_dicLab MASRLNKTSAVGGFSKVLNVLAVLSGLCLLFASVGAE-----------------------IRTAVIDKQTGQLSVVDGYREGFVAWANFTDDIKTSG
>PLBD2_hipHip masrlnktDGVQDKQDVFCGEFSSASVAFYVLCLTCVRAEI--------------------KSAVIDGQSGELSVVDGFQKDFVAWANFTDDIQTSG
>PLBD2_parOli MASRINKMGVEDKQDVSCVEFCVRAEI----------------------------------KSAVIDAQSGDLCVRDGFHQDLVAWANFTDDIQTSG
>PLBD2_gasAcu MASRQNTTVTLRHFKAVLSALFVMCACVQAEI-----------------------------RSAVIDKQTGKLSVVEGYREGFVAWSNFTDDINTSG
>PLBD2_oreNil MACRRNGADRVRSFTEVLGLLKMFLLLFCLFAVRAEI-----------------------SRTAVIDKQTGQLSVIEGYQEDFVAWANFTNDIETSG
>PLBD2_sebCau MASRHNKMFAVGRFKVALSVLSTLCFMCASVGAEV--------------------------RTAVVNKQTGQLSVVEGYREDFVAWSNFTDDIKTSG
>PLBD2_osmMor MAFRLLRLSTTLHLAVFLHVLFLSCSSIKAEI-----------------------------STIVLDEKTGQLTILEGYRDDYVAWANFTDDIEHSG
>PLBD2_onyTsh MADRRTQMSLTTEKMFMFSCVFYLSWTSVRAEI----------------------------PSKILDKQTGQLSLEEGFRDDYVAWANFTDDIKNSg
>PLBD2_salSal madrrtqMSVTTEKMFMFLCVFYLSWTSVGAEI----------------------------HSAVLDKQTGQLSLEEGFRDDFVAWANFTDDIKNSG
>PLBD2_danRer MAHLQLLVSAVCVLLSVCQAQI---------------------------------------YSAIYEEETAQLLLIEGARTHSVAEANFTDHINTTG
>PLBD2_calMil MCVGVRGQGLGLGLPLLLVLAAVGVSPSARGHL---------------------------LRSVVLDEHSGRLRVVGGLNPHSIAWANLTDRIRATG
>PLBD2_braFlo MAACRNIFCGRMLSCLLLFSFVFSAV-----------------------------SDGSKLASVRYDEAAKTYQITDKLDPSAAAWANFTDRISSTG
>PLBD2_acyPis MLSIRCILLSLLFVWALQCSATQK------------------------------NQTLLAVKTDNNRITIQPKHYSVKDKEIIIGKGKFIDRINSTG
>PLBD2_triAdh MAQCGKFLIYFSIFIITLATLCSCQS-------------------------------------GSVIYKDGLYTFSKGINKRAASYGTFTDKIASSG
The two paralogs do not align at all in the signal region and only poorly thereafter:
PLBD2_homSap: 67 DVSAGQLLMVDGRHPDAVAWANLTNAIRETGWAFLEL--GTSGQ-YNDSLQAYAAGVVEAAVSEE 128
Q+ V ++ DA + N N+++ TGW LE+ G Q ++ + + AG +E ++
PLBD1_homSap: 50 AEKTVQVKNVMDKNGDAYGFYN--NSVKTTGWGILEIRAGYGSQTLSNEIIMFVAGFLEGYLTAP 112
Phylogenetic variation signal peptide location in first two exons of PLBD1:
<------ signal peptide ---------> <-------------------------- second exon ---------------------------------->
>PLBD1_homSap MTRGGPGGRPGLPQPPPLLLLLLLLPLLLVTAEPPKPA:GVYYATAYWMPAEKTVQVKN-VMDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQTLSNEIIMFVAGFLEGYLTAP
>PLBD1_panTro MTRGGPGGRPGLPQPPPLLLLLLLLPLLLVTAEPPKPA:GVYYATAYWMPAEKTVQVKN-VMDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQTLSNEIIMFVAGFLEGYLTAP
>PLBD1_ponAbe MTRGGPGGRPGLPPPPPLLLLLLLPPLLLVAAEPANSA:GVYYATAYWMPTEKTVQVKN-VMDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQALSNEIIMFVAGFLEGYLTAP
>PLBD1_rheMac MTRGGPGGCPGLPPPLPLLLRLLLPPLLLVTAESPNPA:GVYYATAYWMPAEMTVEVKN-IMDKNGDAYGFYNNSVETTGWGILEIRAGYGSQALSNEIIMFVAGFLEGYLTAP
>PLBD1_papHam MTRGGPGGCPGLPPQLPLLLRLLLPPLLLVTAESPNPA:GVYYATAYWMPAEMTVEVKN-IMDKNGDAYGFYNNSVETTGWGILEIRAGYGSQALSNEIIMFVAGFLEGYLTAP
>PLBD1_calJac MTRGGPGGRLGLPPPPLLLLLLLLLPPLPTTAEPPTPA:GISYATAYWMPAEKTVQVKN-VMDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQALSNEIIMFVAGFLEGYLTAL
>PLBD1_otoGar MANRTLDRRLGLPPPPLLLLLLLPPPPLLVTAARKNPP:GVYYATAYWKPAEKTVEVKK-VIDKNGDAYGFYNNSMNATGWGILEIRAGYGSQALSNEMTMFVAGVLEGYLTAP
>PLBD1_musMus MCHRSPGRSLRPPSPLLLLLPLLLQPP-WAAALPASPT:GVHCATAYWSPESKKVEIKT-VLDKNGDAYGYYNDSIKTTGWGILEIRAGYGSQVLSNEIIMFLAGYLEGYLTAL
>PLBD1_ratNor MCHRSHGRSLRPPSPLLLLLPLLLQSP-WAAAPLRSSA:GVHYATAYWLPDTKAVEIKM-VLDKKGDAYGFYNDSIQTTGWGVLEIKAGYGSQILSNEIIMFLAGYLEGYLTAL
>PLBD1_cavPor MALCGPGCSPGLPPSPLLLLPLLL----LAAAWSPSPP:GIHYATAYWIPDTKTVEVKD-ILDKDGDAYGYYNNSMEATGWGILEIKAGYGSQELTNEIIMFVAGFLEGYLTAL
>PLBD1_speTri MSRRSLGCGRW-PPPPLQLLPLLLLLLPLAAAQP----:EVYYATAYWIPSEKSIKVKH-VMDKSGDAYGYYNDSMETTGWSILEIRAGYGSQALSNEIIMFVAGFLEGYLTAP
>PLBD1_oryCun MALWLPPLLFPLL---------------LAAAEPPSPE:GVSYATAYWMDAEKKVQVRN-VLDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQALSNEIIMFVAGFLEGYLTAP
>PLBD1_turTru MSRRSPDGSLGLLSPPALLLLLL------AAVVPSGLA:GVYYATAYWMPTEKRIQVQN-VLDRNGDAYGFYNNSVKTTGWGILEIRAGYGSRSLSNEIVMFAAGFLEGYLTAP
>PLBD1_bosTau MSRHSQDERLGLPQPPALLPLLLLL----AVAVPLSQA:GVYYATAYWMPTEKTIQVKN-VLDRKGDAYGFYNNSVKTTGWGILEIKAGYGSQSLSNEIIMFAAGFLEGYLTAP
>PLBD1_oviAri MPRHRRDERLGLPPPPARLPLLLLLL---AAAVPLSQA:GVYYATAYWMPTEKRIQVKN-VLDRKGDAYGFYNNSVKTTGWGILEIKAGYGSQSLSNEIIMFAAGFLEGYLTAP
>PLBD1_susScr MSRRSRDGRLGLPAPPAPL-LLLLLL---AAAVPPSLA:GVYYATAYWMPTEKRMLVKN-VLDRNGDAYGFYNDSMKTTGWGILEIRAGYGSQSLSNNIIMFAAGYLEGYLTAP
>PLBD1_equCab MARHRPDGRLGLPAPPAPPLPPLLLLLLV-AAVSPSQA:VVYSATAYWMPAEKTVQVKN-VMDRNGDAYGFYNNSVKTTGWGILEIRAGYGSQTLSNDITMFVAGFLEGYLTAL
>PLBD1_felCat MARRSRDGRPGLSAPPTPPLLPLLLL---AAAVSPSLA:EVHYATVYWMPAEKTIQVKN-VLDRNGDAYGFYNDSVKTTGWGVLEIRAGYGSQALSNEIIMFVAGFLEGYLTAP
>PLBD1_canFam MPRRARDARLEPCPPLLPLLLLLL-----AAAVPQGRA:EVYYATAYWIPDEKTIQVKN-VLDRNGDAYGFYNDSVKTTGWGILEIRAGYGSQILSNEITMFVAGFLEGYLTAP
>PLBD1_pteVam MSRRSLDGRLGLPATSAPPLLLLLLL---AAAVPPSLA:evyYATAYWMPAEKTVNVKN-LLDKNGDAYGFYNNSMNTTGWGILEIKAGYGSQTLSNDIIMFVAGYLEGYLTAP
>PLBD1_eriEur MSRRSRDGRLGLLLSPPLLLLLLLL-----AAAPPSLQ:EIYYATAYWMPEEEEIQVKN-VLDKNGDAYGFYNDSMLTTGWGILEIKAGYGSHQLSNDVVMFVAGFLEGYLTAP
>PLBD1_sorAra MARGGGDGPPALLPLPLLSLLLALL----AAAVPPSLA:EVHYATAYWMPDEQRVEIKT-TLDKKGDAYGYYNDSVLTTGWGILEIRAGYGSQDLTDEITMFVAGALEGYLTAP
>PLBD1_loxAfr MSSRSRGRHHGPAPQLPQLLLLLLLLLLVAAAAPPSLA:EVHYATVYWMSSEKTMQVKD-VLDKKGDAYGYYNDSVLTTGWGVLEIKAGYGSQALSNDIIMFAAGYLEGYLTAL
>PLBD1_proCap MCSRSV--PCRLSPPLSPPLSLPLLLLLLAAAAPPSLA:EVHYATVYWMSSEKTMQVKD-TLDKNGDAYGFYNDSMQTTGWGVLEIKAGYGSQGLSNDVIMYAAGYLEGYLTAp
>PLBD1_echTel MSTHSRGGR--PAPPLSPSLSLTPLLLL-AALVAPSLA:EIHYATAYWMSSEKTIQIKD-VLDKSGDAYGFYNDSVNATGWGILEIRAGYGSQNLSNDIIMFAAGFLEGYLTAP
>PLBD1_choHof MSRSCQAERLGPVPRRRLLLLLL-----VASAAPPSVA:EVFYATAYWIPSEKKIVVKD-ILDQNGDAYGFYNDSMKTTGWGILEIKAGYGSHIPSNEIIMFTAGFLEGYLTAE
>PLBD1_triVul MSRRSRDGRLGLPAPPAPLLLLLLL----AAAVPPSLA:GVYYATAYWMPTEKRMLVKN-VLDRNGDAYGFYNDSMKTTGWGILEIRAGYGSQSLSNNIIMFAAGYLEGYLTAP
>PLBD1_monDom MTRFSCFGRLQLW--PLQVLLLLLL----TFGAPVTQA:GIHYATVYWNSSTSSAEVKD-SLDPDGDAYGFYNDTIQTTGWGILEIRAGYGANSLTDEIIMFVAGFLEGYLTAQ
>PLBD1_ornAna MSRTCRGGRSGPPQPAPTPAGLLLLLL--TVASPLLQS:HVRYATAYWESATQTVRVKD-VLDWDGDAYGFYNHTVQTTGWGTLEIRAGYGAQALSDEVVMFVAGFLEGYLTAP
>PLBD1_taeGut MARAGGGVCRCCCWALVLLWAAAGGRA-----------:ELRYATVYWNRAEKILQVKN-TLDRSGDAYGFYNNSLQTTGWGVLEIRAGYGSQTLSNEDIMYVAGFLEGYLTAP
>PLBD1_galGal MARLGGGALCCCWGLVLLWAVAGGRA------------:EMRYATLYWNKAQKILQVKN-ILDRSGDAYGFYNNTVQTTGWGVLEIKAGYGHQTLSNEDIMYAAGFLEGYLTAP
>PLBD1_melGal MARLGGGPLCCCWGLVLLWAVAGGRA------------:EMRYATLYWNKAQKILQVKN-ILDRSGDAYGFYNNTVQKTGWGVLEIKAGYGHQTLSNEDIMYAAGFLEGYLTAP
>PLBD1_sisCat MIRFGNPSSSDTRRQRCRSWYWGGLLLLWAVAETRA--:DIHYATVYWLEAEKSFQIKD-VLDKNGDAYGYYNDTIQSTGWGILEIKAGYGNQPISNEILMYAAGFLEGYLTAS
>PLBD1_ambMex MGGLRQLLPLCALLLLQPLGAR----------------:AIRYATVYWTD-RKTVLVKE-VLDKGGDAYGFYNDTIQSTGWGVLEIRAGYAPTSRTNEEIMFAAGYLEGYLTAL
>PLBD1_takRub MFLLTSTCAFVLLTLPATSSTADG--------------:GTAAATVYWDPQHKTVLLKEGVLEQEGDAYGYFNDTLSSTGWSVLEIRAGYGTTPETDEVIFFLAGYLEGFLTAQ
>PLBD1_danRer MPDFSFCVLFLIGFLFSSRSD-----------------:KLK-ATVYWDATHKSAVLKQGVLDPAGASYGYYDNVLLSTGWGVLEVRAGYGDTTQTDDITMFTAGYLEGFLTAP
>PLBD1_ictPun MTEFMVCVCMFLCAVIAVRTDS----------------:VHK-ATAYWDPDSKTVLLKDGVLEDTGDAYGFYNDSFSETGWGVMEVRAGYGQTPRADERTFFLAGYLEGFLTAR
>PLBD1_perFla MEKQSIKLCVLLSTLAASVQTY----------------:QLQEATVYWDGAQKSVILKEGVMETEGGAYGYFNDTLLLSGWGVLEICAGHGGITQEDETTFFLAGYLEGYLTAG
>PLBD1_gasAcu MFLEKTLYVLLLCSVSTTSSAD----------------:KMTAATVYWDPQHKVVLLKEGVLEKEGDAYGYLNDTLSSTGWSVLEIRAGYGETPETDEVTFFLAGYLEGFLTAQ
>PLBD1_oryLat MKLEVFLLLHVIATFASSQ-------------------:KLTAATVYWDAQHKLVLLKEGVLETEGDAYGYLNNTLSTSGWSILEIRAGYGKTPEDDEITFFLAGYLEGFLTAQ
>PLBD1_pimPro MDTNSICVLLLLCSVSTTSSAD----------------:KMTAATVYWDPQHKVVLLKEGVLEKEGDAYGYLNDTLSSTGWSVLEIRAGYGETPETDEVTFFLAGYLEGFLTAQ
>PLBD1_dicLab MPLVTRLYVFLLFTVVTSFASAD---------------:KMTAATVYWDPLHKLVKLKEGVLETEGDAYGYLNDTLSSSGWSILEIRAGYGKTPETDELTFFLAGYLEGYLTAQ
>PLBD1_salSal MKRVCLLFFFYVAASFASAD------------------:EMKAATVYWDATHKTVQLKEGVIEKEGDAYGYLNDTLSQTGWSVLEIRAGYGETLEHDEVTYFLAGYLEGFLTAP
Difference alignment of exon 1 from placental mammals:
PLBD2_homSap MVGQMYCYPGSHLARALTRALALALVLALLVGPFLSGLAGAIPAPGGRWARDGQVPPASRSRSVLLDVSAGQLLMVDGRHPDAVAWANLTNAIRETG
PLBD2_panTro .......S.............................................P...........................................
PLBD2_ponAbe ......GSS.....----...................................P.T...........A......L......................
PLBD2_rheMac .......SS..P....................L.................H..P.T..........HAAT....L....Q...........S.H...
PLBD2_papHam .......SS..P....................L.................H..P.T...........AAT....L......................
PLBD2_calJac ...K..SS.S.R..Q...............A.L.....S..............S..SG.G....V..AA.....L..................H...
PLBD2_otoGar ...P..GS..GR..................I.L...C......P..SGR....LIT.....S.....ATTD..RL..............S...H...
PLBD2_musMus .AAPVDGSS.GWA....R.....TSL..S.T.LL...P...L.TL.PG.Q.QNPD..V..T..L...AAS...RLE..F..................
PLBD2_ratNor .AAP.DRTH.GRA....R.....----.S.A.LL.......L.TL.PG.R.QNPE.....T..L...AAS...RLEY.F..................
PLBD2_dipOrd .AAPP.GSR.GRP.GS.S...V.----.V...LSP..P...V.S..D..G.HKPE..V.......V.AAS...RL...L..G...............
PLBD2_cavPor .AAPT.VSLDGRPV..--......PA.C....LS.GR....V....P.G....P..A.--C......AAS...RL...LQ.G...........P...
PLBD2_oryCun ..APRDGCA.GR.....A--------....T.LL.G.....A.....GEQ..PPS....CC..A...AAT...RL..................H...
PLBD2_ochPri .AATRDSSA.CR...V.......---...PT.L....P.....VR.DGEE.GRPA.SG..C....V.AES...RL......A...........H...
PLBD2_turTru ..DP..GC..GR....................L.....T....T.R.HRGPGRP......C......PET...RL......................
PLBD2_bosTau ..AP..GS..GR....V...............L.....T....T.R.QRG.GMP......C..L...PET...RL......................
PLBD2_oviAri ..AP..GS..GR....V...............L.....T....T.R.QRG.GMP......C..L...PET...SL......................
PLBD2_susScr ..AP..GS..GR....................L.....TS...T.K.YRGSGRS.............TET...RL..................H.N.
PLBD2_ursAme .AAP..GS..GR....................L.....T....IS.RQ.GPN.P...D.........AET...RL......E...............
PLBD2_myoLuc ..APPSRS..GR.TP..S..P...PG....A.L....WT....T.RDP.GPN.P..........V..ART...QL....Q.............H...
PLBD2_pteVam ..AP.DRS..GR..G....T.E.T....P.A.L....RTS..QT..S..GSE.P.S...........PQT...RL..................H...
PLBD2_eriEur ..AP.CGS..GRP...........PA......L...S......P.EDN.G.N.SF..V..C......SET...RL..............S...H...
PLBD2_loxAfr ..APV.GS..GR......Q...V.........L.....T...SLT.H..GP..PA............TAT...RL......................
PLBD2_echTel ..ATE.GS..GR........P....M......L.....T...SPA...REPN.R.....S...A...PAT...RLA.....E...........H...
Consensus MVAPMYGSPGGRLARALTRALALALVLALLVGLFLSGLAGAIPaPGGRWGRDGPVPPASRSRSVLLDAATGQLRLVDGRHPDAVAWANLTNAIRETG
Alignment of first two exons of PLBD1 from vertebrates showing onset of conservation:
Understanding conserved residues in PLBD1 and PLBD2
Although the gene duplication creating these paralogs took place in early unicellular eukaryotes with PLBD1 quite diverged in primary sequence from PLBD2 today, it is nonetheless instructive to compare individual residues and residue patches that are still conserved, given the folds have diverged rather little. Here we wish to exploit the situation that more is known about the maturation and substrates of PLBD1 whereas excellent crystallographic structures exist for PLBD2 and certain of its ancient homologs.
Localization of conserved residues within compared secondary structures: s = beta sheet, h = alpha helix
38 PTGVHCATAYWSPESKKVEIKTVLDKNGDAYGYYNDSIKTTGWGILEIRAGYGSQVLSNEIIMFLAGYLEGYLTALHMYDHFTNLYPQLIKN----PSIV PLBD1
61 PPVSRTRSLLLDAASGQLRLEDGFHPDAVAWANLTNAIRETGWAYLDLST---NGRYNDSLQAYAAGVVEASVSEELIYMHWMNTVVNYCGPFEYEVGYC PLBD2
ssssssssss sssssss ssssssssss ssssssssss hhhhhhhhhhhhhh hhhhhhhhh hh
ssssssssss sssssss ssssssssss sssssssss s hhhhhhhhhhhhhh hhhhhhhhhh hhhh
134 KKVQDFMEKQEMWTRQNIKAQKDDPFWRHTGYVVTQLDGLYLGAQKRASEE-KIKPMTMFQIQFLNAVGDLLDLIPSLSPTKSSSMMKFKIWEMGHCSAL PLBD1
158 EKLKNFLEANLEWMQREMELNPDSPYWHQVRLTLLQLKGLEDSYEGRLTFPTGRFTIKPLGFLLLQISGDLEDLEPALNKTN----------GSGSCSAL PLBD2
hhhhhhhhhhhhhhhhhhhh hhhhhhhhhhhhhhhhhhhhh hhhhhhhhh sss
hhhhhhhhhhhhhhhhhhhh hhhhhhhhhhhhhhhhhhhhh hhhhhhhhh sss
233 IKVLPGFENIYFAHSSWYTYAAMLRIYKHWDFNIKD------KYTLSKRLSFSSYPGFLESLDDFYILSSGLILLQTTNSVYNKTLLKQVVPK-TLLAWQ PLBD1
253 IKLLPGGHDLLVAHNTWNSYQNMLRIIKKYRLQFREGPQEEYPLVAGNNLVFSSYPGTIFSGDDFYILGSGLVTLETTIGNKNPALWKYVQPQGCVLEWI PLBD2
sssss sssssssssssss ssssssss sssssss ssss sssssssss hhhh hh
sssss sssssssssssss ssssssss sss sss sssssss ssss sssssssss hhhh hh
326 RVRVANMMAEGGKEWAQIFSKHNSGTYNNQYMVLDLKKVTINRSL-DKGTLYIVEQIPTYVEYSDQTNV-LRKGYWASYNIPFHKTIYNWSGYPLLVHKL PLBD1
353 RNVVANRLALDGATWADVFKRFNSGTYNNQWMIVDYKAFLPNGPSPGSRVLTILEQIPGMVVVADKTAELYKTTYWASYNIPYFETVFNASGLQALVAQY PLBD2
hhhhhhhh hhhhhhhh sssssssss ssssssss sssssss hh sssss hhhhhh hhhhhhh
hhhhhhhh hhhhhhhh sssssssss ssssssss ssssssshhhhhhhsssss hhhhhh hhhhhh
424 GLDYSYDLAPRAKIFRRDQGNVTDMASMKYIMRYNNYKEDPYSKGDPC-------STICCREDLNGAS---------PSPGGCYDTKVADIFLASQYKAYAISGPTVQDGLPPFNWNRF--NETLHRGMPEVFDFNFVTMK -
453 GDWFSYTKNPRAKIFQRDQSLVEDMDAMVRLMRYNDFLHDPLSLCEACNPKPNAENAISARSDLNPANGSYPFQALHQRAHGGIDVKVTSFTLAKYMSMLAASGPTW-DQCPPFQWSKSPFHSMLHMGQPDLWMFSPIRVPWD
hhhhhhh hhhhhhhhh sss ssssssssss hhhhh ssssss sss sss sss
hhhhhhhh hhhhhhhhh sss sss sss ssssssssss hhhhh ssssss sss sss sss
Suppose a phylogenetically broad set of curated PLBD1 and PLBD2 are aligned together. After careful consideration of gap placement, a restricted number of residues will prove very deeply conserved in both proteins throughout eukaryotes. Of these, some are universal localizational, modificational, structural, or catalytic features basic to the entire NTN clan of 12 protein families and so not particular to phospholipases.
This class of residues must be found by structural alignment of crystallographic structures, as primary sequences are too diverged for these to be accurately located by ClustalW or similar methods. Since the fold of PLBD2 was originally recognized by the fold comparison (via Dali) to all of PDB, these are known already: the autocatalytic cysteine residue at the N-terminus of the 40 kDa fragment and the three active site residues noted above.
An additional 6%-14% of structurally equivalent amino acids (themselves only half of the chain) are identical as amino acids, with IMPC (inosine monophosphate cyclohydrolase) being the highest but PVA (penicillin acylase V) and CBAH (conjugated bile acid hydrolase) also significant. An actual primary sequence multiple alignments is needed to produced a list of super-invariant amino acids (plus those with narrow reduced alphabets, while eliminating accidental matches) within PLBD1 and PLBD2. They may largely lie within the beta strands of the core αββα sandwich as they are better conserved than alpha helices within NTN hydralases but it is fair to say that this fold class is not understood until an explanation can be given for each of the universally conserved residues.
1oqz CA cephalosporin acylase 3pva PVA penicillin V acylase 1k5s PGA penicillin G acylase 2bjf CBAH choloylglycine hydrolase 2ntm IMP cyclohydrolase 1ryp --- a chain among 28 of yeast proteasome Dali report on 3fgr vs 1oqzB: AyaA is a candidate super-invariant region but only second A occurs in both PLBD1 and PLBD2 DSSP leeeeeeeeelllleeeeeelllllLLEEEEEEEEhhhhleEEEEEEELllllhhhHHHHH Query vsrtrsllldaasgqlrledgfhpdAVAWANLTNAiretgwAYLDLSTNgryndslQAYAA 61 ident A A Sbjct ---------------pqapiaaykpRSNEILWDGY------GVPHIYGV-------DAPSA 33 DSSP ---------------llllllllllLLLEEEEELL------LLEEEELL-------LHHHH DSSP HHHHHHHHHHHHHHHHHhhLLLL--LLLL------LLLH-HHHH--HHHHHHHHHHHHH Query GVVEASVSEELIYMHWMntVVNY--CGPF------EYEV-GYCE--KLKNFLEANLEWM 109 ident Sbjct FYGYGWAQARSHGDNIL-rLYGEarGKGAeywgpdYEQTtVWLLtnGVPERAQQWYAQQ 91 DSSP HHHHHHHHHHHHHHHHH-hHHHHhlLLHHhhhlhhHHHHhHHHHhlLHHHHHHHHHHLL DSSP hhhhhhllllhhhhhHHHHHHHHHHHHHHHHL---------llllllllllLLLLlLHHHHL-HHHH-HHHHhhlLLL Query qremelnpdspywhqVRLTLLQLKGLEDSYEG---------rltfptgrftIKPLgFLLLQI-SGDL-EDLEpalNKT ident L Sbjct ---------------SPDFRANLDAFAAGINAyaqqnpddispdvrqvlpvSGAD-VVAHAHrLMNFlYVAS---PGR DSSP ---------------LHHHHHHHHHHHHHHHHhhhhlhhhllhhhhlllllLHHH-HHHHAHrLMNFlYVAS---PGR
In effect, these residues must be 'subtracted off' the larger set of phylogenetically invariant residues between PLBD1 and PLBD2 because they have nothing to do with phospholipases per se. The remaining conserved residues however provide the defining specializations of class B phospholipases. These sites developed subsequent to the divergence of phospholipases from generic NTN hydralases but prior to the gene duplication giving rise to PLBD1 and PLBD2. They are evidently mission-critical, being retained in both paralogs in all surviving species up until the present day. Several active site residues fall into this category as described earlier.
(to be continued)
Comparative structural genomics
Another approach here is to structurally model a series of phylogenetically spaced primary sequences. This can be done for PLDB2 using the known mouse structures and for PLBDB1 using the refined SwissModel available for it (admittedly derived from mouse PLDB2).
Suppose 25 such models were obtained for each paralog for human, mouse, ... elephant, opossum, bird, frog etc. These could be compared and perhaps an ancestral progression obtained. Variation then could be suppressed assuming the substrate has remained the same for long evolutionary periods. Then the two resulting averaged structures could be subtracted, defining the stable differences between PLDB2 and PLDB2.
(to be continued)
Conserved in PLBD2 but not PLBD1: lysosomal specificity
(to be continued)
Conserved in PLBD1 but not PLBD2: neutrophil specificity
"The native protein needed modifications to acquire deacylation activity against phospholipids including phosphatidylcholine, phosphatidylinositol, phosphatidylethanolamine and lysophospholipids. Enzyme activity was associated with fragments derived from the 42 kDa fragment. The enzyme revealed a PLB nature by removing fatty acids from both the sn-1 and sn-2 positions of phospholipids. The enzyme is active at a broad pH range with an optimum of 7.4. Immunoblotting of neutrophil postnuclear supernatant using antibodies against the 42 kDa fragment detected a band at a molecular mass of 42 kDa, indicating a neutrophil origin of the novel PLB precursor."
(to be continued)
Conserved residues determining mannosylation
Many proteins exported to the endoplasmic reticulum are glycosylated but few are ultimately targeted for the lysozome. The key feature is said to be mannosylation recognizable by the lysomal mannose receptor. Yet it is unclear why the generic glycosylation NxT/S is sometimes mannosylated in a suitable way, rather than receiving some other oligosaccharide. This could be key to timing the targeting to the lysozyme during the evolutionary history of phospholipases.
(to be continued)
Reference sequences
PLBD1 reference sequences
>PLBD1_homSap Homo sapiens (human) FLJ22662 PMID: 19019078,20093120 0 MTRGGPGGRPGLPQPPPLLLLLLLLPLLLVTAEPPKPA 1 2 GVYYATAYWMPAEKTVQVKNVMDKNGDAYGFYNNSVKTTGWGILEIRAGYGSQTLSNEIIMFVAGFLEGYLTAP 2 1 HMNDHYTNLYPQLITKPSIMDKVQDFME 2 1 KQDKWTRKNIKEYKTDSFWRHTGYVMAQIDGLYVGAKKRAILEGTK 0 0 PMTLFQIQFLNSVGDLLDLIPSLSPTKNGSLKVFKRWDMGHCSALIK 0 0 VLPGFENILFAHSSWYTYAAMLRIYKHWDFNVIDKDTSSSRLSFSSYP 1 2 GFLESLDDFYILSSGLILLQTTNSVFNKTLLKQVIPETLLSWQRVRVANMMADSGKRWADIFSKYNS 1 2 GTYNNQYMVLDLKKVKLNHSLDKGTLYIVEQIPTYVEYSEQTDVLRK 1 2 GYWPSYNVPFHEKIYNWSGYPLLVQKLGLDYSYDLAPRAKIFRRDQGKVTDTASMKYIMRYN 1 2 NYKKDPYSRGDPCNTICCREDLNSPNPSPGGCYDTK 0 0 VADIYLASQYTSYAISGPTVQGGLPVFRWDRFNKTLHQGMPEVYNFDFITMKPILKLDIK* 0 >PLBD1_braFlo Branchiostoma floridae (lancelet) XM_002595538 0 MEGRACRSCRLHHLSAVFLLFLVTIAA 1 2 GAEIQATAYLQAQGKVQVKLGVLDKQNGDAVATYDDR 2 1 LTENGWGVLNVVSGFGPKKLSDNDIMYLAGYLEGVLTQE 2 1 RIYQHYLNLYGIFFMGKSEDLVGK 0 0 VKKFYTAQDTWVRAQVKQSTDPVMKHLSYILSQYDGLVKGYNDN 0 0 LFPHVSFFQKLDIFAFQLLNGNGDTFDIIPAVNPSSRPDFSNMSRVEIDDWVSAHSHCSALVK 0 0 VLGAYENVYMSHSSWFNYAATMRIYKHYNFNIANPATATRKMSFSSYP 1 2 GYLESLDDFYLMDSGLVMLQTTNNVFNGTLYDLVKPESILAWQRVRTANMLARNGDQWGAIMNVHNS 1 2 GTYNNQYMIIDLNLIELGKTIHDGALYVVEQIPGLVMSADQTDILRA 1 2 GYWPSYNIPFYEKVYNLSGYPEFAKSQGLDYTYQLAPRAKIFRRDAGKVKDMESMKAIMRYN 1 2 DYLHDPYSKGNPCSAICCRKDLAKVGAKPDGCYDTK 0 0 VSDYYLARNLTSFAINGPTLGTGLEPFSWSDKFKISHIGLPKVYNFSFVTMTPAEL* 0 >PLBD1_strPur Strongylocentrotus purpuratus (urchin) XM_001192029 0 MANKFRMFKILTAFLVLVLVNLST 1 2 GELLQGTVYKQEDGTFTVSSGIIDKQGVAYGSYNNTLFQTGWGELHLFAGYSTADNVALSDADRMYAAGILEGALTAK 2 1 QISQTLRNINVTFFSAESDPEIWRRVADFFETQDAWMKGMIIERADEDPFWEGVGLVLAQFEGLIKGYEMSQFSNAST 0 0 SNGFLAMQVLNSCGDLLDLKSAVMPSLIPDWDKLTKKEFLKFIRTSGHCSALVK 0 0 ICAALVKVGRFAPPFQSLLYSIS 0 0 SYFKSQAILKLNSPSCQLFGIE 1 2 GFLESLDDFYIMSSGLSMLQTTNNIFNKTLYKYVKPQSLLAWQRVRVANMMARSGKDWARIVARYNS 1 2 GTYNNQYMVIDRTKIKPNVAILDDALWVVEQVPTLVASGDQTNILRA 1 2 GYWPSYNVPFYEEIYNISGYPEYAYKGGADISYQLAPRAKIFRRDQGNVVDMESFKKIMRFN 1 2 DYKNDPYSEGDPSKSICMRGDLMTSPMPNGCYDTK0 0 VTNLAMAAKQTSFVINGPTRGDGSLPPFKWVAPFTGWSHVGLPTVYDFNFVEMCPKEL* 0 >PLBD1_nemVec Nematostella vectensis (anemone) XM_001638165 0 MTLIRNSVMITVTFVLILFVFGCHGSQKSATVYYNRGQG 2 1 YSLKFGVVDKLMGVAYGTFEDSLNTTG 2 1 WYELNIVSGTGIEPYNDDVIMHAAGYLEGALTAS 2 1 QINDNYANLYGVFFKSEDDPMVAKVEKFFIEQ 0 0 DIWMRKMIALKSSNSSFWRQMGNIIAQFD 1 2 GLVEGYQKYPATDK 0 0 ALGVFAFQMLNGVGDLLDLTKALMPERMADWDHMTEKEILEK 0 0 VAMDGHCSALIKVLPAYENVFASHVS 2 1 WFTYSAMLRVYKHYHLNLKDETT 1 2 AAQRMSFSSYPGFLESLDDFYIMDS 2 1 KLVMLQTTNNVFNKSLYEQVVPESLFSWQRVRLANLVASSGRQWADIVGQYNS 1 2 GTYNNQYMVLDLKLIQLNNTIQDNALWVVEQIPT 2 1 LVASGDQTAILRAGYWPSYNVPFYEL 0 0 VYNLSGYPDFVARHGVQFSHELAPRAKIFRRDQSM 0 0 VHDLDSMKHIMRYNDFQHDPYSQGNPMNAICSRGDLIADGPRASGCYDGK 0 0 VTDFTMAQSLISHAINGPTHE 0 0 QQVPFHWSQYQFKNKHEGQ 0 0 PDLFNFDFVEMKPKF* 0 >PLBD1_monBre Monosiga brevicollis (choanoflagellate) XM_001745398 MSSLNNGIPEPLLKFLAAQFNWTRSQVAANQDDVFWQQVGLIMA QYDGLRAGYGANVYDKHVLPEFAFQLLNGNGDFFDIIPKAVDVTKMSSREFHDWRMRN GRCSALIKLTGDFSDLFMSHSAWYIYQAMNRIYKHCASYNFQATITHAKKISFSSYPG YLESLDDFYLMSSGLVMLQTTNNVFNTDLQQYIQPESLQSWIRIRTATALAQTSEDWA ELAGRHNSGTYNNQYMVMDLNKFTPGQPLLDGTLYVAEQIPGTWEYADVTKMLSLGYW PSYNVPFFEKIYNLSGYPAVVKQHGTDDSYELAPRAKIFRRDQTTVVDLDSFKAIMRY NDYKNDPYAKGDPYNAICSRGDLESDSPSPGGCYDTKVTTYSMALKLQSQVINGPTTS HGLPPFSWSQFPNASHLGMPEVFNFTFETMDAGW*
Trimmed alignable sequences
(to be continued)
PLBD2 reference sequences
>PLBD2_homSap Homo sapiens (human) PMID: 19706171,19237744,17007843 0 MVGQMYCYPGSHLARALTRALALALVLALLVGPFLSGLAGAIPAPGGRWARDGQVPPASRSRSVLLDVSAGQLLMVDGRHPDAVAWANLTNAIRETG 2 1 WAFLELGTSGQYNDSLQAYAAGVVEAAVSEE 0 0 LIYMHWMNTVVNYCGPFEYEVGYCERLKSFLEANLEWMQEEMESNPDSPYWHQ 0 0 VRLTLLQLKGLEDSYEGRVSFPAGKFTIKPLGFL 2 1 LLQLSGDLEDLELALNKTKIKPSLGSGSCSALIKLLPGQSDLLVAHNTWNNYQHMLRVIKKYWLQFREGPW 1 2 GDYPLVPGNKLVFSSYPGTIFSCDDFYILGSGL 0 0 VTLETTIGNKNPALWKYVRPRGCVLEWVRNIVANRLASDGATWADIFKRFNSGT 2 1 YNNQWMIVDYKAFIPGGPSPGSRVLTILEQIP 2 1 GMVVVADKTSELYQKTYWASYNIP 2 1 SFETVFNASGLQALVAQYGDWFSYDGSPRAQIFRRNQSLVQDMDSMVRLMR 2 1 YNDFLHDPLSLCKACNPQPNGENAISARSDLNPANGSYPFQALRQRSHGGIDVK 0 0 VTSMSLARILSLLAASGPTWDQVPPFQWSTSPFSGLLHMGQPDLWKFAPVKVSWD* 0 >PLBD2_musMus Mus musculus (mouse) NM_023625 0 MAAPVDGSSGGWAARALRRALALTSLTTLALLASLTGLLLSGPAGALPTLGPGWQRQNPDPPVSRTRSLLLDAASGQLRLEDGFHPDAVAWANLTNAIRETG 2 1 WAYLDLSTNGRYNDSLQAYAAGVVEASVSEE 0 0 LIYMHWMNTVVNYCGPFEYEVGYCEKLKNFLEANLEWMQREMELNPDSPYWHQ 0 0 VRLTLLQLKGLEDSYEGRLTFPTGRFTIKPLGFL 2 1 LLQISGDLEDLEPALNKTNTKPSLGSGSCSALIKLLPGGHDLLVAHNTWNSYQNMLRIIKKYRLQFREGPQ 1 2 EEYPLVAGNNLVFSSYPGTIFSGDDFYILGSGL 0 0 VTLETTIGNKNPALWKYVQPQGCVLEWIRNVVANRLALDGATWADVFKRFNSGT 2 1 YNNQWMIVDYKAFLPNGPSPGSRVLTILEQIP 2 1 GMVVVADKTAELYKTTYWASYNIP 2 1 YFETVFNASGLQALVAQYGDWFSYTKNPRAKIFQRDQSLVEDMDAMVR 0 0 LMRYNDFLHDPLSLCEACNPKPNAENAISARSDLNPANGSYPFQALHQRAHGGIDVK 0 0 VTSFTLAKYMSMLAASGPTWDQCPPFQWSKSPFHSMLHMGQPDLWMFSPIRVPWD* 0 >PLBD2_braFlo Branchiostoma floridae (lancelet) XM_002612057 0 MAACRNIFCGRMLSCLLLFSFVFSAVSDGSKLASVRYDEAAKTYQITDKLDPSAAAWANFTDRISSTG 2 1 WSFLTVTTNEKYDDSVQAYAAGLVEGYLTRD LMYNHWLNTVGAAFCSSRSAFCKNLESFLKTNLAWMQEQIQASGDTDDYWHQ 0 0 VKLTLQQLSGLDDGYNDDPRQPSLDINPFGFL 2 1 IFQIGGDMEDLQEALKDKDSHRVLGSGSCSALVKLLPGNADLLVAHDTWDTFQSMLRIIKKYQFPFKLGGKK 1 2 GEDKIPGHTVSFSSYPGVIYSGDDFYITSASL 0 0 VAQETTIGNSNPALWKYVQPQGQVLEWLRNIVANRLANKAMDWATIFKKYNSGT 2 1 YNNQWMIVDYKTFTPNKDLPEKGLLVVLEQLP 2 1 GMVMMDDVTSVLAKQAYWPSYNSP 2 1 YFEKIFNTSGLPAMVEKYGDWFSYEHTPRANIFRRDHGKVTDISSMIKLMR 2 1 YNDFQNDPLSKCDCTPPYSAENAISARSDLNPANGTYPFSALQHRCHGGTDMK 0 0 MTSYSMHESHQMMAVSGPTHDQQQPFQWSTSDYDKQFYHLGHPDLFNFDPIHVIWFDQSDN* 0 >PLBD2_droMel Drosophila melanogaster (fruitfly) U57314 retinal lamina neuron ancestor (lama) PMID: 16077094,8892229 0 MERPEYDGTYCATALWTKQVGFQIENWKQQNDLVNIPTGVGRICYKDSVYENGW 0 0 AQIEVETQRTYPDWVQAYAAGMLEGSLTWRNIYNQWSN 2 1 TISSSCERDESTQKFCGWLRDLLTTNYHRLKRQTEKAENDHYWHQLHLFITQLEGLETGYKRGASRARSDLEEEIPFSD FLLMNAAADIQDLKIYYENYELQNSTEHTEEPRTDQPKNFFLPSATMLTKIVQEEESPQVLQLLFGHSTAGSYSSMLRIQK RYKFHYHFSSKLRSNTVPGVDITFTGYPGILGSTDDFYTIKGRHLHAIVGGVGIKNENLQLWKTVDPKKMVPLVARVMAANRI SQNRQTWASAMSRHPFTGAKQWITVDLNKMKVQDNLYNVLEGDDKHDDAPVVLNEKDRTAIQQRHDQLRDMVWIAEQLPGMMTKK DVTQGFLVPGNTSWLANGVPYFKNVLELSGVNYSEDQQLTVADEEELTSLASVDKYLRTHGFRGDLLGSQESIAYGNIDLKLFS YNARLGISDFHAFAGPVFLRFQHTQPRTLEDEGQDGGVPPAASMGDERLSVSIEDADSLAEMELITERRSVRNDMRAIAMRKIGSGP FKWSEMSPVEEGGGHEGHPDEWNFDKVSPKWAW* 0 >PLBD2_acyPis Acyrthosiphon pisum (aphid) XM_001948827 0 MLSIRCILLSLLFVWALQCSATQKNQTLLAVKTDNNRITIQPKHYSVKDKEIIIGKGKFIDRINSTG 2 1 WAYLEIRTSQKAKDEDQAYGAGYLEGTLTADLIYSYWFNTAKGYCTDRPNVCQQLKDYMTTNKNWIKSKLNESDPYWYQ 0 0 VGLYYKQLDGLYDGYMRGKSPSTPDLTWDDLY 2 1 WLNALDDLGDLSIALYPSDISNRVLGSGSCSALIKLMPDNKDILVSHATWSG 2 1 YETMLRIQKRYSLRFRKSKKSNKLIRGFDMSFSSFPGGIQSGDDFYLISSGLTTMETTIENYNDSLWSNVKPVGQ 0 0 VLEFVRAMVANRLADNPTDWANLFKLHNSGTYNNQWMILNYAAFQPGSPLPPRDVLHVLEQIPGHVMHDDFTGHLINRTYWASYNVPYFPFIFNVSGNYEMEQIYGSW 2 1 FSYSETPRARIFARDHVKIHCDKCMLHLMRSNNYTRDPESRCDCSPPYSAENAISSR 2 1 NDLNPANGTYPIRALGHRSHGATDVKVTSSQLFQQLQFKAIAGPTQGSNNSLGPFCWSKSDFNDKVSHLGHPDCFNFKPVLHQWSL* 0 >PLBD2_triAdh Trichoplax adhaerens (trichoplax) XM_002107718 introns largely conserved 0 MAQCGKFLIYFSIFIITLATLCSCQSGSVIYKDGLYTFSKGINKRAASYGTFTDKIASSG 2 1 WTYLDVHTNPQDDDFITAYAAGYVEGILTAKY IYMHWKNTVGDYCKQKSIYCQKLKSFIMKNNQWMATQIKHRPHSIYWYH 0 0 INLTLIQQKGLRDGYHKAMPHKPIDEFSFL 2 1 LIELSGDLESLETALKDEDTHHVLGSGSCSAFIKVLPDNRDLYFAHDTWTGYQTMLRIYKYYELNFSMLPKTN 1 2 VTVPGTRISFSSYPGTILSGDDYYLIGSGL 0 0 ATMETTNGNSNEKLWKYVTPSSVLEWIRTIIANRLTSSGNDWVKIFSKYNSGT 2 1 YNNQ 00 WMILDYKLFAPKRPLNPNTLWVLEQIP 2 1 GKIESADVTNVLKKQGYWASYNVP 2 1 YFSSIFNMSGNQEQAKKYGNWFTHDKCPRALIFKRDQHKVNSMESLMKLMR 2 1 YNDFKHDPLSRCNCTPPYSAENAISARSDLNPADGKYNIGALGHRCHGGTDSK STNYTMFHSGLKSYAIAGPTHEQQPPFRWSTAKFNMTKPLGHPDLFNFTRQLVSWD* 0 >PLBD2_monBre Monosiga brevicollis (choanoflagellate) introns all novel 0 MWSCGAAAAAVVAVVVLASPATATVARFVEQTDVQTTYASVFYVESDDSYVVKTENHPWDGDFEKDE 0 0 AVRIKYTPGYLVAGWDQLHVKSNSAMDDATVAYAAGYGEAQLTAEMIYNYAYNNGYDTFTPNDKLADYLAKNQAFMAASIASNRSDANGYWYHVDLILRQLQGVCDGYNSSD FAKSFPLPCESMLAINLMGDMEDLSDALASSDEWYTEDRFFRATHCSALVKLVGGASSPSDIYISQDTWSSLNSMTRIMKRYDLNFLQ 2 1 AKGADDRIAGSSIVFSSYPGSLYSGDDFYLTSAGMAVIETTIGNSNPELYQYIVPDTVLEWIRNIMANRLASNSQTWYEVYRQFNSGT 1 2 YNNMNMILDYKQFKPQEALQDELLTIVEQIP GTVTKTDVTGYLRNMTYWGS 1 2 YNVAFDQNIRELSGANQAEQLYGPW 2 1 FSYWNTSRALIFAREQKNVSSLEDLKRLMRLNQFKTDPL 2 1 YRGWTNCTPAYTAENVIATRGDLNDP 0 0 NGIYSLSSFGLRNHVATDSKISTFSTYDSNNLNVWAIS 2 1 GPTNGPPPNQPVFNWSTSYYKDTRHRGMPEAFDFDWVNFNWPF* 0 >PLBD2_dicDis Dictyostelium discoideum (slime_mold) AAFI02000019 AF411829 introns both novel 0 MRVIRSLLLLTIAIIGSVLSQSSIDDGYTVFYSQPDNYYVKPGTFSNGVAQAIFSNEMMTTGWSFMSISSSEGLYPNDIIAAGAGYLEGYISQEMIYQNWMNMYNNEYHNVIGSD VENWIQENLQYLQTMIDSAPSNDLYWQNVETVLTQITYMQRGYNQSVIDNGVDASQSLGITEFFLMNMDGDMIDLGPALNLTNGKQVTSPATATSPKQAFKEFMRRTGHCSALIKMTDDLSDLFSGHTTW 2 1 SSYYEMVRMFKVYNLKYLFNGQPPASKVTMFSGYPGTLSSIDDFYLLDTKIVVIETTNGLMNNNLYHLITSESVLSWIRVIVANRLATGGESWCQTFSLYNSGTYNNQ 0 0 WIIVDYNKFIKGYGALDGTLYILEQVPDYVEYGDQTAILRTGYWPSFNIPFYENIYGLTGFNETYAQFGNWFSYQASPRSMIFKRDANNIHSLTQFQAMLRYNNWQNDPFSQGNAGN QISSRFDLVTADDPNNQYLDPDAFGGIDSKVVSADMVAALLVNAQSGPSHDNETPFTWNSQWNQKYTYAGQPTTWNFDWMTMSLQSMKPASPSSDSSSDSTTFN* 0