Selenoprotein evolution: SECIS
Introduction to selenocysteine insertion into selenoproteins
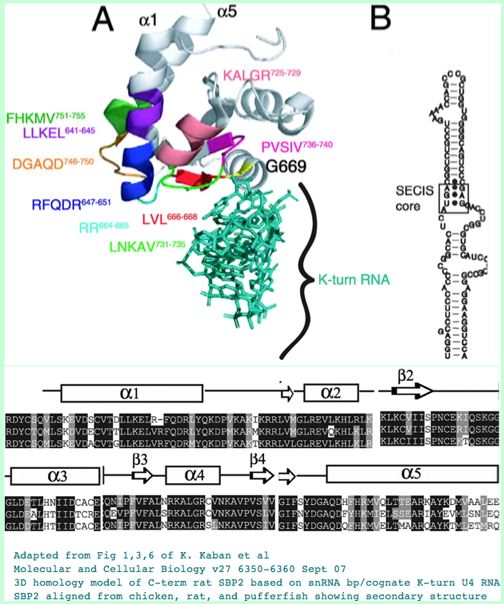
The SECIS element in 3' UTR of selenoproteins allows the insertion of selenocysteine for the appropriate TGA codon at time of translation. Auxillary gene products are needed for this to happen, including a selenocysteine-specific translation elongation factor (eEFSec), Sec-tRNA[Ser]Sec, ribosomal protein L30, and a binding protein (SBP2).
The SBP2 protein has three domains, including a conserved L7Ae RNA binding motif for SECIS RNA binding, a not completely overlapping ribosome binding site and a redox-sensitive cysteine-rich domain. SBP2 shuttles between the nucleus and the cytoplasm via nuclear localization motif SELEQNESSKKNKKKKEKSKSSYEVLPVQE and nuclear export signal motifs, the latter using the CRM1 pathway.
The kink-turn tertiary structure in the SECIS element is at the core of binding recognition. A mutation in that of SEPN or in SB2 lead to rigid spine muscular dystrophy or abnormal thyroid hormone function, the latter through inadequate production of DIO2 needed to activate thyroid hormone T4.
A strong paralog of BP2 was noticed and briely studied earlier. That protein, KIAA0256, also has 17 coding exons with near-identical intron locations and phases and full-length alignability, establishing KIAA0256 and BP2 as resulting from an ancestral gene duplication. The percent identity overall is only 42% but reaches 72% (human to human) in the 3 exons defining the L7Ae kink-turn binding motif.
Since both genes can be recovered from chondrichthyes, lamprey, and amphioxus, the duplication event preceded the emergence of chordates. The roots of the selenoproteome and the insertion mechanism lie much deeper in pre-metazoa so it follows that the parent gene must have assumed the key SECIS recognition roles (as did both descendent genes, at least initially).
We could ask if KIAA0256 still plays a role in SECIS in contemporary organisms. One possibility is that some of the 30-odd selenoproteins preferentially use KIAA0256 over BP2, correlating to the two observed structural classes of SECIS elements (which needs to be revisited in the genomics era). Another possibility is differential use during development or in various cell types. It's worth noting that KIAA0256 is considerably more conserved across its length than BP2, which has only a few exons conserved back to fish) suggesting the latter has a more limited role.
A literature review establishes that DIO, GPX, and SEPP1 were generally taken as proxies for all selenoprotein SECIS elements in terms of establishing a role for BP2. Yet the timing of expansion of these and specialization of the former to thyroid during the period of gene duplication may have tainted their representativeness. Possibly many other selenoproteins utilize KIAA0256 at least to some extent. That would account for the greater conservation of KIAA0256 and the difficulty finding SECIS elements in expected position for certain selenoproteins, ie their elements may be adapted to a different receptor and so have low Cove scores to the extent that tool is based on the BP2 fold paradigm.
Ribosomal protein L30 appears also to bind the SECIS kink-turn and play a role in selenocysteine insertion despite its meagre length of 114 amino acids. Likely many ribosomal proteins, L30 has remarkable sequence conservation, here still 94% at human-lamprey.
L30 has weak homology and secondary structure parallels to the pertinent region of BP2 and KIAA0256. These latter proteins might be viewed as domain composites that have accreted the kink-turn binding domain long ago from this undoubtedly ancient ribosomal protein. This opens a window on the origin of selenocysteine insertion in early eukaryotes.
Key residues and regions in SECISBP2 and KIAA0256
>SECISBP2_homSap Homo sapiens (human) full length 0 MASEGPREPESE 0 0 GIKLSADVKPFVPRFAGLNVAWLESSEACVFPSSAATYYPFVQEPPVTE 2 1 QKIYTEDMAFGASTFPPQYLSSEITLHPYAYSPYTLDSTQNVYSVPGSQYLYNQPSCYRGFQTVKHRNENTCPLPQEMKALFK 0 0 KKTYDEKKTYDQQKFDSERADGTISSEIKSARGSHHLSIYAENSLKS 1 2 DGYHKRTDRKSRIIAKNVSTSKPEFEFTTLDFPELQGAENNMSEIQKQPKWGPVHSVSTDISLLREVVKPAAVLSK 0 0 GEIVVKNNPNESVTANAATNSPSCTR 1 2 ELSWTPMGYVVRQTLSTELSAAPKNVTSMINLKTIASSADPKNVSIPSSEALSSDPSYNKEKHIIHPTQK 0 0 SKASQGSDLEQNEASRKNKKKKEKSTSKYEVLTVQEPPRIE 0 0 DAEEFPNLAVASERRDRIETPKFQSKQQPQ 0 0 DNFKNNVKKSQLPVQLDLGGMLTALEKKQHSQHAKQSSKPVVVS 1 2 VGAVPVLSKECASGERGRRMSQMKTPHNPLDSSAPLMKKGKQREIPKAKKPTSLKK 0 0 IILKERQERKQRLQENAVSPAFTSDDTQDGESGGDDQFPEQAELS 1 2 GPEGMDELISTPSVEDKSEEPPGTELQRDTEASHLAPNHTTFPKIHSRRFRD 2 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTVAARQAYKTMLENVQQELVGEPRPQAPPSLPTQGPSCPAEDGPPALKEKEEPHY 1 2 IEIWKKHLEAYSGCTLELEESLEASTSQMMNLNL* 0 407–525 domain required for U insertion but not SECIS binding (399–516 in rat) 540 R540Q allele of SBP2 decreases GPX1 and DIO2 650–752 L7Ae motif kink-turn binding motif 676 invariant glycine (669 in rat) >KIAA0256_homSap Homo sapiens (human) length=1056 0 MDRAPTEQ 0 0 NVKLSAEVEPFIPQKKSPDTFMIPMALPNDNGSVSGVEPTPIPSYLITCYPFVQENQSNR 2 1 QFPLYNNDIRWQQPNPNPTGPYFAYPIISAQPPVSTEYTYYQLMPAPCAQVMGFYHPFPTPYSNTFQAANTVNAITTECTERPSQLGQVFPLSSHRSRNSNRGSVVPK 0 0 QQLLQQHIKSKRPLVKNVATQKETNAAGPDSRSKIVLLVDASQQT 1 2 DFPSDIANKSLSETTATMLWKSKGRRRRASHPTAESSSEQGASEADIDSDSGYCSPKHSNNQPAAGALRNPDSGTMN 0 0 HVESSMCA 1 2 GGVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ 0 0 VGFRCRGHSTSSERRQNLQKRPDNKHLSSSQSHRSDPNSESLYFE 0 0 DEDGFQELNENGNAKDENIQQKLSSKV 0 0 LDDLPENSPINIVQTPIPITTSVPKRAKSQKKKALAAALATAQEYSEISMEQKKLQ 0 0 EALSKAAGKKNKTPVQLDLGDMLAALEKQQQAMKARQITNTRPLSYT 1 2 VVTAASFHTKDSTNRKPLTKSQPCLTSFNSVDIASSKAKKGKEKEIAKLKRPTALKK 0 0 VILKEREEKKGRLTVDHNLLGSEEPTEMHLDFIDDLPQEIVSQE 1 2 DTGLSMPSDTSLSPASQNSPYCMTPVSQGSPASSGIGSPMASSTITKIHSKRFRE 2 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 2 ETNWRNMVETSDGLEASENEKEVSCKHSTSEKPSKLPFDTPPIGKQPSLVATGSTTSATSAGKSTASDKEEVKPDDLEWASQQSTETGSLDGSCRDLLNSSITSTTSTLVP GMLEEEEDEDEEEEEDYTHEPISVEVQLNSRIESWVSETQRTMETLQLGKTLNGSEEDNVEQSGEEEAEAPEVLEPGMDSEAWTADQQASPGQQKSSNCSSLNKEHSDSNYTTQTT* 0
Note the Irish proband with compound mutation K438stop/IVS8ds+29G/A (paternal allele inactive, maternal allele a splice donor mutation leading to early truncation) has been incorrectly described as a SECISBP2 knockout; in fact 48% production of wildtype maternal allele still occurs. Additionally, KIAA0256 may be able to partially compensate for reduction in level or loss of SECISBP2. Knockout mice for tRNA(Sec), unable to make any selenoproteins, die in utero.
These same authors, in Figure 1B published twice identically among rare papers to ever consider KIAA0256 aligned full length SECISBP2 to an intermediate region of the KIAA0256 gene that happened to begin with a methionine, namely residues 422-849 of the 1056 residue protein (which includes the motif-bearing residues 632-829 of exons 14-16). Furthermore the m,odel KIAA0256 transcript was an exon-skipping minor splice variant:
KIAA0256 originally arose in a GenBank submission package from a large-scale mRNA project at the Kaluza Institute. It skips over highly conserved exon 8 (which does not alter downstream reading frame since it resides in a series of consecutive phase 00 spliced exons). Modelling on this oddity, NCBI later confused the record by mixing in its own predicted genes (from genome assemblies without significant transcript programs) and labelling them mRNAs.
While baboon also has experimental transcripts skipping this exon (FC178616, FC178616), mammalian transcripts almost always retain the exon, for example human (AK307480), macaque (CJ457866), mouse (AK145135), rat (CK602552), dog (CO708934), horse (CX604216), cow (CK846448), sheep (EE864720), and even chicken (DR417186).
Transcription and its processing in mammals is a noisy process and artefacts abound. While exon-skipping in some cases may have physiological significance, lacking significant comparative genomics support, the null hypothesis is that they do not. Here the full length gene must contain exon 8 since it is quite conserved in amino acid sequence throughout tetrapods and assuredly ancestral. As all SECIS binding experiments to date used protein lacking exon 8; consequently we know nothing about the SECIS binding properties of intact protein.
KIAA0256 and SECISBP2 actually align well over their entire lengths and have 17 near perfectly comparable exons (once per trillion odds for coincidence). There is no reason to believe that such an odd fragment studied on a few SECIS sequences could accurately represent binding properties of full length protein in regards to the 25-odd orthology class of SECIS elements. However this became accepted folklore with the selenocysteine research community and the properties of full length KIAA0256 have never been determined.
Reference set of 27 vertebrate SECIS BP2 L7Ae motif exons
>SECISBP2_homSap Homo sapiens (human) full length 0 MASEGPREPESE 0 0 GIKLSADVKPFVPRFAGLNVAWLESSEACVFPSSAATYYPFVQEPPVTE 2 1 QKIYTEDMAFGASTFPPQYLSSEITLHPYAYSPYTLDSTQNVYSVPGSQYLYNQPSCYRGFQTVKHRNENTCPLPQEMKALFK 0 0 KKTYDEKKTYDQQKFDSERADGTISSEIKSARGSHHLSIYAENSLKS 1 2 DGYHKRTDRKSRIIAKNVSTSKPEFEFTTLDFPELQGAENNMSEIQKQPKWGPVHSVSTDISLLREVVKPAAVLSK 0 0 GEIVVKNNPNESVTANAATNSPSCTR 1 2 ELSWTPMGYVVRQTLSTELSAAPKNVTSMINLKTIASSADPKNVSIPSSEALSSDPSYNKEKHIIHPTQK 0 0 SKASQGSDLEQNEASRKNKKKKEKSTSKYEVLTVQEPPRIE 0 0 DAEEFPNLAVASERRDRIETPKFQSKQQPQ 0 0 DNFKNNVKKSQLPVQLDLGGMLTALEKKQHSQHAKQSSKPVVVS 1 2 VGAVPVLSKECASGERGRRMSQMKTPHNPLDSSAPLMKKGKQREIPKAKKPTSLKK 0 0 IILKERQERKQRLQENAVSPAFTSDDTQDGESGGDDQFPEQAELS 1 2 GPEGMDELISTPSVEDKSEEPPGTELQRDTEASHLAPNHTTFPKIHSRRFRD 2 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTVAARQAYKTMLENVQQELVGEPRPQAPPSLPTQGPSCPAEDGPPALKEKEEPHY 1 2 IEIWKKHLEAYSGCTLELEESLEASTSQMMNLNL* 0 >SECISBP2_panTro Pan troglodytes (chimp) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVQQELVGEPRPQAPPSLPTQGPSCPAEDGPPALTEKEEPHY 1 >SECISBP2_macMul Macaca mulatta (rhesus) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVQQELAGEPRPQAPPSPPTQGPSCPAEDGPPALTEKEEPHY 1 >SECISBP2_otoGar Otolemur garnettii (bushbaby) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKERRLVLGLREVLKHLKLKKLICVISPNCERQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVQRELAGEPGPQVPSSLPMEGPSCSVEDSPPAPTEKEEPHY 1 >SECISBP2_tupBel Tupaia belangeri (tree_shrew) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRVVLGLREVLKHLKLKKLKCVIISPIZEKIQSK 1 2 GGLDDTLHTIIAYACAQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMEARQAYRSMLESARQELAGEPGLQAPPQPPVQGPRASSEGSAPAPTGRQEPHC 1 >SECISBP2_musMus Mus musculus (mouse) 1 YCSQMLSKEVDACVTGLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKTQSK 1 2 GGLDDTLHTIIDCACEQNIPFVFALNRKALGRSLNKAVPVSIVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLETMRQEQAGEPGPQSPPSPPMQDPIPSTEEGTLPSTGEEPHY 1 >SECISBP2_ratNor Rattus norvegicus (rat) exons 1416 1 YCSQMLSKEVDACVTGLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKTQSK 1 2 GGLDDTLHTIIDCACEQNIPFVFALNRKALGRSLNKAVPVSIVGIFSYDGAQDQ 0 0 FHKMVELTMAARQAYKTMLETMRQEQAGEPGPQTPPSPPMQDPIQSTDEGTLASTGEEPHY 1 >SECISBP2_cavPor Cavia porcellus (guinea_pig) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCIIISP 1 2 GLDDTLHTIIDYACAQNIPFVFALNRKALGRSLNKTVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVRQELAGEPRPQMPPDPPSEGPSSSLEDTAPDPSAEEPHY 1 >SECISBP2_oryCun Oryctolagus cuniculus (rabbit) 1 YCSQMLSKEVDACVTDLFKELVRFHDLMYQDPVKATTKCQFELRVGKALDHLRLKKLKCIIVFPKHKKQS 1 2 TIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENMRHELAGEPGPPTPQPVQGPSCSAEDGPPAPTEGEVPHY 1 >SECISBP2_canFam Canis familiaris (dog) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHRMVELTMAARQAYKTMLENVRQELAGEPGTPALANPPMQGLGCSTQDSPPAPTEKEEPHY 1 >SECISBP2_felCat Felis catus (cat) 1 YCSQMLSKEVDACVTDLLRELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIGYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHRMVELTMAARQAYKTMLENARQELAGEPGPPAPGSPPPQPPAPAGRDEPRY >SECISBP2_equCab Equus caballus (horse) 1 YCSQILSKEVDACVTELLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPCVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTKAARQAYKAMLENVHQELAGEPGPQAPASPPAQGPSCSTEGAPPAPTGKEEPHY 1 >SECISBP2_bosTau Bos taurus (cow) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKAKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACDQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYRTMLENARQELPGELGPCAPVGPPSQGPGCPVEDSPLAPTEKEEPHY 1 >SECISBP2_eriEur Erinaceus europaeus (hedgehog) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCIIISPNCEKIQSK 1 2 GGLDETLHTIIDCACEQNIPFVFALNRKALGRSLNKGVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKALLENMRQELAEESGSPAPSSPPVQSPSEDGPPAPAEKEEPHY 1 >SECISBP2_dasNov Dasypus novemcinctus (armadillo) 1 YCSQVLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGELDDTLHTIIDYAASRHSICVALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKAMLENVRKELAGEPGPRSPPSPPALGPHSSAGDVHPTSAGKEEPHY 1 >SECISBP2_loxAfr Loxodonta africana (elephant) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALHRKALGRSLNKPVPVSVVGIFSYDRAQ 0 0 DQFHKMVELTMAARQEYKTMLESVRQELAEEPRAGSPPSPPTQGPGCSAEVPRPAPTEKEEPRY 1 >SECISBP2_monDom Monodelphis domestica (opossum) 1 YCSQMLSKEVDDCVMDLLKELVRFQDRMYQKDPVKAKTKRRLVMGLREVLKHLKLKKLKCVIISPNCEKSKSK 1 2 GGLDETLHTIIDYACEQNVPFVFALNRKALGRSVNKVVPVSVVGIFSYDGAQ 0 0 DQFHKMIALTMEARQAYKIMLSTLKEEPALETENPPSPSLPRPSESCPSELGQTDPTQEEEPNY 1 >SECISBP2_triVul Trichosurus vulpecula (possum) 1 YCSQMLSKEVDDCVMDLLKELVRFQDRMYQKDPVKAKTKRRLVMGLREVLKHLKLKKLKCVIISPNCEKSKSK 1 2 GGLDETLHTIIDYACEQNVPFVFALNRKALGRSVNKVVPVSVVGIFSYDGAQ 0 0 DQFRKMIELTMEARQAYKVMLATLKEGAEALQTENPLPTSLTPQGQGCSSELSKTTDPTKEEEPNY 1 >SECISBP2_galGal Gallus gallus (chicken) 1 YCSQVLSKEVDSCVTDLLKELVRFQDRLYQKDPVKAKIKRRLVMGLREVLKHLRLKKLKCVIISPNCEKIQSK 1 2 GGLDETLHNIIDCACEQNIPFVFALNRKALGRCVNKAVPVSVVGIFSYDGAQ 0 0 DHFHRMVQLTTEARKAYKDMVAALEEELKELSKPLNZKSCLSETGKTSSTKEDIPNY 1 >SECISBP2_anoCar Anolis carolinensis (lizard) 1 YCTQVLSKEVDSCVTDLLKELVRFQDRLYQKDPVKAKTKRRLVMGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDETLHLIIDSACEQNIPFVFALNRKALGRCLNKAVPVSVVGIFSYDGAQ 0 0 DYFHKMVELTMEARQAYKDMISALERELKKKTVRKKPLQSRPLDTVEASSTEEDVPDY 1 >SECISBP2_xenTro Xenopus tropicalis (frog) NM_001097262 1 YCSQVLSKDVDNCVMELLKELVRFQDRLFLKEPAKAKSKRRLVMGLREVLKHLKLQKLKCIIISPNCEKIQSK 1 2 GGLDDTLQTIISHACEQNVPFVFALNRKALGRCLNKAVPVSVVGVFSYDGAQ 0 0 DHFHKLCELTVQARQAYKDMIAAAQEQQSETEAGKNEEDPVAVNGQNKSDDMREESKAEEPDEPNY 1 >SECISBP2_danRer Danio rerio (zebrafish) 1 YCNQVLSKDVDECVSNLLKELVRFQDRLYQKDPMKARMKRRLVMGLREVLKHLKLKKVKCVIISPNCERIQSK 1 2 GGLDEALHNIIDTCRDQSVPFVFALSRKALGRCVNKAVPVSLVGIFNYDGAQ 0 0 DFYHKMIELSSEARTAYEVMLLNLEQTDAEEAQQTSPLAEKVETSSGDPQPEEPEY 1 >SECISBP2_tetNig Tetraodon nigroviridis (pufferfish) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPTKAKSKRRLVMGLREVTKHMKLQTIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 DFYHKMIELSSEARIAYEVMLSNLEQTSAEEEPQTCTLAEKINTSSEDAQPEEPEY 1 >SECISBP2_takRub Takifugu rubripes (fugu) 1 YCTQMLSKDVDECVTTLLKELVRFQDRLYQKDPIKARMKRRIVMGLREVQKHLKLRKLKCVIISPNCERIQSK 1 2 GGLDEALHTIIDTCREQAVPFVFALSRRALGRCVNKAVPVSLVGIFNYDGAQ 0 0 DFYHKMIELSSEARTAYEVMLLNLEQTDAEEAQQTSPLAEKVETSSGDPQPEEPEY 1 >SECISBP2_gasAcu Gasterosteus aculeatus (stickleback) 1 YCNQVLSKEIDESVTMLLQELVRFQERIYQKDPTKAKTKRRLVMGLREVTKHMKLNKIKCVLISPNCEKIQAK 1 2 GGLDEALYNVIAMARDQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 DFYHKMIELSSEARRAYEVMVSSLEQTGQADPESVEEKLQISSAAEEAELGRDITPPEEPEY 1 >SECISBP2_oryLap Oryzias latipes (medaka) 1 YCSQMLRKDVDECVTVLLKELVRFQDRLYHKDPIKARMKRRLVMGLREVLKHLKLRKVKCVIISPNCEQIQSK 1 2 GGLDEALHTIIQTCREQAVPFVFALSRKALGHCVNKAVPVSLVGIFNYDGAQ 0 0 DHYHKMIELSAEARKAYEVLVSSLERDQQEESHPDRGTCFGSVTAEPEKPHY 1 >SECISBP2_calMil Callorhinchus milii (elephantfish) AAVX01044988 1 YCSQVLSKDVDSCVTDLLKELVRFQDRLYQKDPIKAKKKRRIVMGLREVLKHLKLKRLKCIIISPNCEKIQSR 1 2 GGLDDALHNIISIACEQEIPFVFALNRKALGQCVNKPVPVSVLGIFSYDGAE 0 0 NQFHQMVEITEEARKAYQEMLDALQQELEADEEKGDSEEQPLISSESSTIHFNNVTSQPFSEADEPEY 1 >SECISBP2_braFlo Branchiostoma floridae (amphioxus) extra exon 1 YCNQVLDKEIDATVTMLLQDLVRFQDRQYHK 00 DPIKAKAKRRIVMGLREVTKHLKLRKLKCIIIAPNLEKIQSK 1 2 GGLDDAIETILNLCMEQDVPFVFALGRKALGRAVNKLVPVSVVGVFNYDGAE 0 0 1
Reference set of 22 deuterostome SECIS KIAA0256 L7Ae motif exons
>KIAA0256_homSap Homo sapiens (human) uc001zxd chr15 exons 14-16 75% 69% 51% 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_panTro Pan troglodytes (chimp) 2 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 0 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 1 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_macMul Macaca mulatta (rhesus) 2 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 0 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 1 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_tupBel Tupaia belangeri (tree_shrew) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_musMus Mus musculus (mouse) 0 YCNQVLSKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 1 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 2 SLFNRLVELTEEARKAYKDMVAATEQEQAEEALRSVKTVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_ratNor Rattus norvegicus (rat) 2 YCNQVLSKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 0 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 1 SLFNRLVELTEEARKAYKDMVAATEQEQAEEALRSVKAVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_canFam Canis familiaris (dog) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_equCab Equus caballus (horse) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVALTEEARRAYKDMVAALEQEQAEEASKNVKKGPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_dasNov Dasypus novemcinctus (armadillo) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSKG 1 2 GLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_monDom Monodelphis domestica (opossum) 0 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVKAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 1 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYCGAE 0 2 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_galGal Gallus gallus (chicken) 1 YCNQVLSKEIDECVTLLLQELVSFQERIYQKDPMRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 DLFNKLVSLTEEARKAYRDMVAAMEQEQAEEALKNVKKAPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_anoCar Anolis carolinensis (lizard) 1 YCNQVLSKEIDECVTLLLQELVSFQEQIYQKDPMRAKAKRRLVMGLREVTKHMKLSKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 NLFNKLVSLTEEARKAYRDMVAAMEQEQEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_xenTro Xenopus tropicalis (frog) 1 YCNQVLSKEIDECVTVLLQELVSFQERVYQKDPVKAKSKRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFSYSGAE 0 0 SLFHNLVSLTEEARKAYKDMVSSMEQEQAEEALKNIKKVHMGHSRNPSAASAISFCSVISEPISEVNEKDY 1 >KIAA0256_danRer Danio rerio (zebrafish) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKEPSKAKAKRRLVMGLREVTKHMKLHKIKCVIISPNCEKIQAK 1 2 GGLDEALHNIIDTCRDQSVPFVFALSRKALGRCVNKAVPVSLVGIFNYDGAQ 0 0 GLFNKLVSLTEEARRAYKEMVSALEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_tetNig Tetraodon nigroviridis (pufferfish) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPTKAKSKRRLVMGLREVTKHMKLQTIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 SLFNQLVSLTEEARKAYKDMVSALEQEQTEEALKNEKKVPHQMGHYRNHSAASAVSFCSIFSEPISEVNEKEY 1 >KIAA0256_takRub Takifugu rubripes (fugu) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPTKAKSKRRLVMGLREVTKHMKLQTIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNFSGAE 0 0 SLFNQLVSLTEEARKAYKDMVSALEQEQTEEALKNEKKVPHQMGHYRNHSAASAVSFCSIFSEPISEVNEKEY 1 >KIAA0256_gasAcu Gasterosteus aculeatus (stickleback) 1 YCNQVLSKEIDESVTMLLQELVRFQERIYQKDPTKAKTKRRLVMGLREVTKHMKLNKIKCVLISPNCEKIQAK 1 2 GGLDEALYNVIAMARDQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 GLFNRLVSLTEEARKAYKDMVSALEQEQAEEAQKNDKKLPHHMGHSRNHSAASAISFCSIFSEPISEVNEKEY 1 >KIAA0256_oryLap Oryzias latipes (medaka) 0 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPSKAKSKRRLVMGLREVTKHMKLHKIKCVIISPNCEKIQAK 1 1 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 2 GLFNQLVSLTEEARKAYKEMVSALEQEQAEEALKHDKKVPHHMGHSRNHSAASAISFCSILSEPISEVNEKEY 1 >KIAA0256_calMil Callorhinchus milii (elephantfish) AAVX01105236 1 YCNQVLSKDIDECVTLLLQELVRFQERVYQKDPIKAKMKRRLVMGLREVTKHMKLRKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 1 >KIAA0256_petMar Petromyzon marinus (lamprey) 1 LHKLRALIISPNCEKIQAK 1 2 GGLDEALQTVIALASEQSVPFVFALNRKALGHCLNKKVPVSVVGVFHYGGAE 0 0 THFQRLVALTEEARSAYRNMVSSLQRQEAAATSEPTGHTEDPLEASASPPSVPAHDPTALLHLLRPQQGPREDDPAEASGRSPGRNA 1 >KIAA0256_cioInt Ciona intestinalis (tunicate) 1 YCCQVLDKRVDEMSNQMLQRLVYFQDR 21 RLYKTDPAKAKRKRRVVLGFREVTKHLKMKKLRCVIISPNLEKIESK 1 2 GGLDDVLHEILDLCKEQNIPYVFALGKKALGRAVSKTVPVSIVGVFDYSGAE 0 0 1 >KIAA0256_strPur Strongylocentrotus purpuratus (sea_urchin) 1 YCNQVLDKDIDGCCTTLLQTLVKFQDRQYHKDPAK 00 AKMKRRLVMGLREVTKHLKLKKIKCVVVSPNLERIQSK 1 2 GGLDEAMDRISSLASEQNVPLIFALGRKALGRAVNKVVPVSVVGIFNYDGAE 0 0 DTYKQLLDLSTRARNAYADMVRKFQQELEAANAASAARMAKHRHHMGHNRNLFKG 1
Reference set of 10 deuterostome ribosomal L30 L7Ae motifs
>L30_homSap Homo sapiens (human) 4 exons numerous pseudogenes 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQTGEK* 0 >L30_tupBel Tupaia belangeri (tree_shrew) 0 MVAAKKT 0 0 KKSLESINSQLQLAMKDGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRACTLAIMDP 1 2 GDSDIIRSMPEQTGEK* 0 >L30_ratNor Rattus norvegicus (rat) Sep15 Gpx4 Gpx1 Dio1 quite weak homology 35% with BP2 exons 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQTGEK* 0 >L30_myoLuc Myotis lucifugus (microbat) 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYLLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 ISEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSD-IRSMPEQTGEK* 0 >L30_echTel Echinops telfairi (tenrec) 0 MVAAKKT 0 0 KNSLESINSRLQLVMKSGKYMLGYKQMLKMIRQGKAKLVVLANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGHNIELGTACGKSCRVCTLAITDP 1 2 GDADIIRSMPEQTGEK* 0 >L30_anoCar Anolis carolinensis (lizard) 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIQQGKAKLVILANNCPALG 2 1 KSEIEYYAMLAKTGVHHYSGNNIEMGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMQEQTAEK* 0 >L30_danRer Danio rerio (zebrafish) 94% 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQSQKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPDQQQGGEK* 0 >L30_squAca Squalus acanthias (spiny dogfish) 97% 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQISEK* 0 >L30_petMar Petromyzon marinus (lamprey) 94% 0 MSAKKT 0 0 KKAIESINSRLQLVMKSGKYCLGYRQTLKMIRQGKAKLVLLANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIEMGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQQQPQPGDK* 0 >L30_braFlo Branchiostoma floridae (amphioxus) 84% to homSap 0 MKQK 0 0 RKTMESINSRLQLVMKSGKYVLGLKETLKVLRQGKAKLIIIANNTPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYFRVCTLAITDP 1 2 GDSDIIRSMPAEDKGESK* 0
Positioning of SECIS elements in 3' UTR
Reference sets of vertebrate SECIS elements
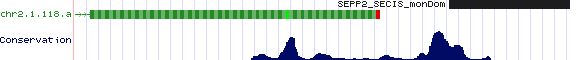
SEPP2: 5 SECIS sequences
>SEPP2_SECIS_monDom Monodelphis domestica (opossum) chr2:9455773-9456962 COVE: 30 55 bp after stop codon ACGTGCCGCTGCCCCTCCCTCCCTCCAAGAATGACGCCCACAGTGAAACCCAGAGAACTGGTCCCTGTGGGCTGATGCCCCAGAGGGGAGGAGAGGC >SEPP2_SECIS_macEug Macropus eugenii (wallaby) SECIS COVE score: 26.38 trace ti|975557399 contains terminal exon CCTCATTCCTCTTCTCCATGGCATCAGCCCACCATGGCCGGTTCCCCGGGTTTCACTGTGGGCATCATTAATTGAGGGAGGGAAGGGCAATGGGCAAG >SEPP2_SECIS_ornAna Ornithorhynchus anatinus (platypus)Ultra131:321101-348720 COVE: 34.28 no match to chicken TCCCTCGACTCCCGCTCCCGCCTCGCACTCATGACGTCCACGGTGTCAACCGGCCCGCCGGGCACCGTGGACTGACGCCGGTCGAGGCGGAGGGGT >SEPP2_SECIS_ornAna Ornithorhynchus anatinus (platypus) Ultra131:321101-348720 COVE: 22.82 no match to chicken GGAATCAGGAACCCAGTAACATGAGGTCATCTTCGGAAGCCTGTGCCTAGAGGACCAAGATAATGGAAAAAGTGACGGACAAGGGTGTGTAGCTGG >SEPP2_SECIS_tetNig Tetraodon nigroviridis (pufferfish) COVE: 17 GCTGGACCCAGGCTGCTGGTGGTCCCGTTGATGACGTCTGCGCTGGTAAACCTGCCTGCAGGAGCCTGTGGACCGACGTGTGTGGACCCACCGGCAG
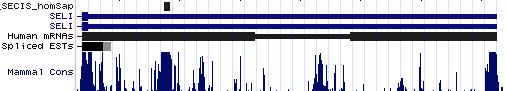
SELI: 17 SECIS sequences
>SELI_SECIS_homSap Homo sapiens (human) COVE score: 21.61 UUUCACUGAA UGAAGUU UGUGCUUGA AUGAA GAGUGUAUCUUA AA CCCCCUUUUUUUGGA CAGGCUGCACUU GGAU AAAAUA GGCACCA CUGUGUUGAU >SELI_SECIS_panTro Pan troglodytes (chimp) COVE score: 21.61 uuucacugaa ugaaguu ugugcuuga augaa gaguguaucuua aa cccccuuuuuuugga caggcugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_macMul Macaca mulatta (rhesus) COVE score: 21.25 uuucacugaa ugaaguu ugugcuuga augaa gaguguaucuua aa ccuccuuuuuuugga caggcugcacuu ggau aacaua ggcacca cuguguugau >SELI_SECIS_otoGar Otolemur garnettii (bushbaby) COVE score: 21.25 uuucacuaag ugaaguu ugugcuuga augaa gaguguaucuu aa acccuuuuuuuuggac agguugcacuu ggau aaaaua ggcacca uuguguugau >SELI_SECIS_musMus Mus musculus (mouse) COVE score: 24.62 uuccacugaa ugaaguu ugugcuuaa augaa gagugugucuu aa acccuuuuuuuuggac agguugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_ratNor Rattus norvegicus (rat) COVE score: 24.62 uuccacugaa ugaaguu ugugcuuaa augaa gagugugucuu aa acccuuuuuuuuggac agguugcacuu ggau aacaua ggcaccg cuguguugau >SELI_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 17.42 uuccagugaa ugaaguu ugugcuuga augaa gaguguaucuua aa cccuuuuuuuuuugga cagguugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_oryCun Oryctolagus cuniculus (rabbit) COVE score: 19.47 ucuacugaau gaaguuu gugcuuga augaa gaguguaucuua aa cccuuuuuuuuugga cagguugcacuu ggau aaaau aggcac cacuguugau >SELI_SECIS_sorAra Sorex araneus (shrew) COVE score: COVE score: 8.13 loose uuccacugaa ugaaguu ugugcuuga augaa gaguguaucuu aa acccuuuauuuuguuuuuuugga cggauuacacuu ggau auaaua ggcacca uucuguugau >SELI_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 14.43 loose cacggaauga aguaugu gcuuga augaa gaguguaucuu aa acccuuuuuuuuuuuugga cagguugcacuu ggau aua auaggc accacucugu >SELI_SECIS_canFam Canis familiaris (dog) COVE score: 14.04 uuccacugaa ugaaguu ugugcucga augaa gaguguaucuua aa cccuuuuuuuuuugga uggauugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_equCab Equus caballus (horse) COVE score: 19.57 uuucacugag ugaaguu ugugcucga augaa gaguguauucuu aa acccuuguuuuugga uggauugcacuu ggau aaaaua agcacca cuguguugau >SELI_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 17.76 loose cuacugaauu aaguuug ugcuuga augaa gaguguaucuu aa acccuuuguuuuuuugga cagguugcacuu ggau aaaa uaggca ccacuauguu >SELI_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 25.81 uuccacugaa ugaaguu ugugcuuga augaa gaguguaucuu aa acccuuuuuuuuggac agguugcacuu ggau aaagua ggcacca cuguguugau >SELI_SECIS_monDom Monodelphis domestica (opossum) COVE score: 26.46 uucuacugaa ugcaauu ugugcuuga augaa gagugugucuu aa auccuuuauauggac aggcugcacuu ggag aga auaagca caaccauguu >SELI_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 19.25 ucccagugaa ugaagcu ugugcuuga augaa gagugcaucuua aa cccauuuuuuuugga aaagcugcacuu ggag agaaag ggcacg acuguguuua >SELI_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 22.02 uccccuuugu guguguc cuuugugcgugu augaa gagugcggccuc aa cccaggcgucuugga agggccgcaccc ggaa gaa acggagc acagcaaaga >SELI_SECIS_galGal Gallus gallus (chicken) COVE score: 24.62 uuuuauugag uuuauuu gugcuuaa augaa gagugcgcuuc aa acccagaccaggag agggcgcacuu ggag ugagc gagucaa accuugcucc
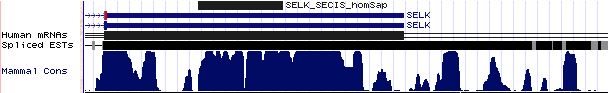
SELK: 19 SECIS sequences
>SELK_SECIS_homSap Homo sapiens (human) COVE score: 34.53 acaaggacug cucugug uccucacagauga augag gucaugcuggg aa uucccucugcaggga acuggccugac ugac augcaguuc cauaaa ugcagauguu >SELK_SECIS_panTro Pan troglodytes (chimp) COVE score: 34.53 acaaggacug cucugug uccucacagauga augag gucaugcuggg aa uucccucugcaggga acuggccugac ugac augcaguuc cauaaa ugcagauguu >SELK_SECIS_macMul Macaca mulatta (rhesus) COVE score: 36.14 acaaggacug cucugug uccucacagauga augag gucaugcuagg aa uucccucuacaggga acuggccugac ugac augcaguuc cauaaa ugcagauguu >SELK_SECIS_otoGar Otolemur garnettii (bushbaby) COVE score: 31.99 caaggauugc ucugugu cuucacagauga augag gucaggcuggg aa uucucucuucaggga acuggccugac ugac augcagu ucuauaa acgcacuuuu >SELK_SECIS_tupBel Tupaia belangeri (tree_shrew) COVE score: 36.65 acaaggauug cucugug uccccacagauga augag guuaugccggg aa uucccuccacaggga ucuggccugac ugau acgcaguuc uauaaa ugcacauguu >SELK_SECIS_musMus Mus musculus (mouse) COVE score: 31.73 acaaggauug cucugug uccccacagauga augag gucaugcuggg aa uucccucugcagga ucuagccugac ugau acgcaguuc uauaaa uguacauguu >SELK_SECIS_ratNor Rattus norvegicus (rat) COVE score: 33.02 acaaggacug cucugug uccucacagaaga augag gucaugcuggg aa cucccucugcagga ucuggccugac ugau gugcaguuc uauaaa uguacaugug >SELK_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 33.72 cucuguguuc ucacaga uaa augag gucacgccagg aa uucucucagcaggga ucuggcuugac ugau acgcagu ucucua aaugcauaug >SELK_SECIS_oryCun Oryctolagus cuniculus (rabbit) COVE score: 38.80 accaggauug cucugug uccccacagauga augau gucaggcuggg aa uucccuccacaggga ucuggccugau ugag augcaguuc uauaaa ugcguauguu >SELK_SECIS_sorAra Sorex araneus (shrew) COVE score: 40.91 auaaggacug uucugug uccacacagauga augag gucaugcuggg aa uucccucuacgggga ucuggcaugac ugau augcag uucgaua aaugcacaug >SELK_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 37.30 acaaggauug cucugug uucucacagguga augag guuaugcuggg aa uucccuccaugggga ucuggcaugac ugau augcaguuc uauaaa ugcacauguu >SELK_SECIS_canFam Canis familiaris (dog) COVE score: 36.30 acaaggauug cucugug ccuucacagacgg augag guugugcuagg aa uucccuccccaggga ucuggcaugac ugac augcaguuc uauaaa ugcacauguu >SELK_SECIS_felCat Felis catus (cat) COVE score: 37.35 acgagaauug cucugug uccucacagacag augag gucgugcuggg aa uucccuccccaggga ucuggcaugac ugac augcaguuc uauaaa ugcacauguu >SELK_SECIS_bosTau Bos taurus (cow) COVE score: 39.16 cucugugucc ucacaga cga augag gucaugcuggg aa uucccuccgcaggga ucuggcaugac ugac augcagu ucuaua aaugcacguu >SELK_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 29.00 acaaggauug cucugug uccucacagauga augag gucauguuuggg aa uucccucugcaggg aucuggcaugac ugac uugcaguuc cauaaa ugcacauguu >SELK_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 34.85 acaaagacug cucugug uccccacagacgg augag gcugugcuggg aa uucccucugcaggga ucuggcauggc ugac augcaguuc cauaaa ugcacauguu >SELK_SECIS_monDom Monodelphis domestica (opossum) COVE score: 21.75 aagaauugcu cugucua cacagauua augau guugugcuggg aa cucccaucuuacagga uccaguguaac ugau ugcaa uuguaua aaugcacaug >SELK_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 33.68 aauaauugug cugugaa caagcagauua augau guuuugcuggg aa uuccuucaggga uccaguauaac ugau aaagc aauuaua uaaaggcaca >SELK_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 33.87 aauucccugc ucugcca auuggcgggacc augau guuguccuggg aa uuccuuauucuggga uccagggcaac ugaa aagcaguuc uguuaa auuaaaugca >SELK_SECIS_galGal Gallus gallus (chicken) COVE score: 30.90 ugagaacugu ucugcaa uauaagcagauga augaa guuguacuggg aa cuccuucaagga uccaguguaac ugaa gugcagug uuauuaa auacauguuu
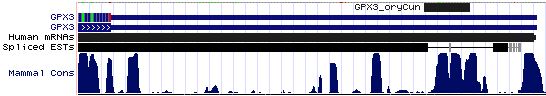
GPX3: 15 SECIS sequences
>GPX3_SECIS_homSap Homo sapiens (human) COVE score: 27.80 ACGGACCCCA UGGCAGG GGUGGCGUCUUC AUGAG GGAGGGGCCCA AA GCCCUUGUGGGC GGACCUCCCC UGAG CCUG UCUGAG GGGCCAGCCC >GPX3_SECIS_panTro Pan troglodytes (chimp) COVE score: 27.80 cacguacccc augucgg ggguggugucuuc augag ggaggggccca aa gcccuugugggc ggaccucccc ugag ccugu cugagg ggccagcccu >GPX3_SECIS_macMul Macaca mulatta (rhesus) COVE score: 27.80 cacguacccc augucag ggguggcgucuuc augag ggaggggccca aa gcccuugugggc ggaccucccc ugag ccugu cugagg ggccagcucu >GPX3_SECIS_tupBel Tupaia belangeri (tree_shrew) COVE score: 29.38 ccacaucccu gugucag ggguggcaucucc augag ggaggggcccg aa gcccuuguggg cggaccucccc ugag ccugu cugagg ggccggcccu >GPX3_SECIS_musMus Mus musculus (mouse) COVE score: 33.37 cguguacccc aggucag ggguggugucucu augaa ggaggggcccg aa gcccuuguggg cgggccucccc ugag cccgu cuguggu gccagcccuu >GPX3_SECIS_ratNor Rattus norvegicus (rat) COVE score: 33.37 caugugcucc aagucag ggguggugucucc augaa ggaggggcccg aa gcccuuguggg cgggccucccc ugag cccgu cuguggu gccagcccuu >GPX3_SECIS_canFam Canis familiaris (dog) COVE score: 29.96 cauguucccc gugucag gaauggcaucucc augaa ggaggggccc ga agcccucaugggc ggaccucccc ugag ccugu cugaag ggccggcccu >GPX3_SECIS_felCat Felis catus (cat) COVE score: 30.08 gccacguccc gugucag ggauggcgucucc augaa ggaggggccc ga agcccuugugggc ggaccucccc ugag ccugu cugaag ggccagcccu >GPX3_SECIS_equCab Equus caballus (horse) COVE score: 26.20 uacauucccc augucag ggguggcaucucc augau ggaggggcccg aa gcccugguggg cggaccucccc agag ccugu cugaag ggccagcccu >GPX3_SECIS_bosTau Bos taurus (cow) COVE score: 30.87 acguuccccg ugucagg gggcggcaucgcc augaa ggaggggcccg aa gcccgcguggg cgggccucccu ugag ccu gucugag gggccagccu >GPX3_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 16.32 cuacgucccc gugucaa gagcggcaucucc augau gguggggcccg aa gccccugugg cggaccucccc agag ccugucc caugggc cagcccuuag >GPX3_SECIS_echTel Echinops telfairi (tenrec) COVE score: 12.78 ggcugugucc cgugcag agggagcaucucc augag ggugaggcccg aa gcccccgugg cggaccucgac ccgag ucug cucugg gccugccuuc >GPX3_SECIS_monDom Monodelphis domestica (opossum) COVE score: 22.94 ugaacauaag ggauggc aucucu augau ggugggaucca aa gccucuucaggg cggguuccauc agag ccugcaaaa gguguc aggacccuua >GPX3_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 31.33 ugucccagaa uggggag guggcaucauc augac agcggggucug aa agccccuccugga uggaccccgcc cgaa ccug cucggcg guggcaugac >GPX3_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 28.20 GAGAUUUUGG GCCAAGG AAGUUGCAUCACU AUGAG GGUUAGGUCUG AA AGCUCCCAAAAAGAG CGGACCUAGCC UGAG GCUGCAAA GCUCUGGU GUAGCCCUUU >GPX3_SECIS_galGal Gallus gallus (chicken) COVE score: 30.65 GCCCUGGGGA GCAGAGG AUGACAUCUCC AUGAA GGCCUGGCCUG AA AGCCCCCACCAUGGGG UGGGCUCGGCC CGAU CCCG CCCAGGC GCGGUGCAGC
GPX5: 0 SECIS sequences (cysteine all species)
GPX6: 10 SECIS sequences (most species have selenocysteine
>GPX6_SECISa_homSap Homo sapiens (human) chr6:28,578,901-28,580,158 Cove score: 18.10 cccaccucac augaagg gaagggcaucucc augau gguggauccc aa aaccccucuggguc gcacccugcc agag ccu uccuug gugccugucc >GPX6_SECIS_panTro Pan troglodytes (chimp) COVE score: 23.45 cccaccucac augaagg gaggggcaucucc augac gguggguccc aa aaccucucgggguc ggacccugcc agag ccu uccuug gugccugucc >GPX6_SECIS_macMul Macaca mulatta (rhesus) COVE score: 21.81 accucauuca cagaagg gaggggcaucuuu augau ggugggucuc aa aaccucucuggguc agacccuacc agag ccu uccuug gugccugucc >GPX6_SECIS_cavPor Cavia porcellus (guinea_pig) AAKN02051715 Cove score: 19.91 AGAAGCCUCC CUAGAAG GAAGGGUGUCUCC AUGAU GGUGGGUCCC AA AAGCCCUGGAUC GGACCCUACC AGAA CCU UCCCUGG UGCCUGUCCU >GPX6_SECIS_canFam Canis familiaris (dog) Cove score: 24.33 but not found in 28way CCUGGCCUCA UAUGAAA GGAGGGCAUCUCC AUGAU GGUGGGUUCC CA AGCCCCUGCGGUC GGACCCUACC AGAG CCU UCUUGG GUGCCUGUCC >GPX6_SECIS_equCab Equus caballus (horse) COVE score: 25.00 uucaucucac augagcu gaggggcaucucc augau gguggguccc aa agccccugugggc aggacccaac cagaa cucug ugccugu cccuuagugc >GPX6_SECIS_bosTau Bos taurus (cow) COVE score: 23.33 cucacuucac augagga aaagggcaucucc augau gguggguccc aa agccucucuggguc ggaccccacc agag ccu uccuugg ugccuguccc >GPX6_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 24.33 ccuggccuca uaugaaa ggagggcaucucc augau gguggguucc ca agccccugcgguc ggacccuacc agag ccu uccuugg ugucuguccu >GPX6_SECIS_sorAra Sorex araneus (shrew) frag COVE score: 22.95 uuucauauga aaggagg gggcaucucc augau ggugggucuc aa agccccucuggguc ggacccuacc agag cccugcugg guguccu gucccuuugu >GPX6_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 19.82 cucaccuuac augaagg gagggacauaucc augau gguggguccu aa agcccuucuggguc agacccuacc agaa ccu ucucug gugccugucc >GPX6_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 19.28 cucaccucac augaaug gaggggcauaucc augau gguggguccc aa agccucucuagguc ggacccuauc agaa ucuucccugg ugccugu cccuuagugc >GPX6_SECIS_echTel Echinops telfairi (tenrec) frag COVE score: 31.33 ugugucccag aaugggg agguggcaucauc augac agcggggucu ga aagccccuccuggau ggaccccgcc cgaa ccu ucccug gugccugucc
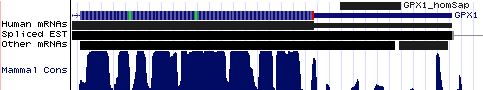
GPX1: 17 SECIS insertion sequences
>GPX1_homSap Homo sapiens (human) COVE score: 20.45 UGCUGUCUCG GGGGGGU UUUCAUCU AUGAG GGUGUUUCCUCU AA ACCUACGA GGGAGGAACACC UGAU CUUACAGAAA AUACCAC CUCGAGAUGG >GPX1_panTro Pan troglodytes (chimp) COVE score: 20.45 gugcugcucu guggggg gguuuucaucu augag gguguuuccucu aa accuacga gggaggaacacc ugau cuuacagaaa auacccc cucgagaugg >GPX1_macMul Macaca mulatta (rhesus) COVE score: 26.45 gcagcgcugc ucucugg gggguuuucaucu augag gguguuuccucu aa accuaca aggaggaacacc ugau aau acagaa aauacccccu >GPX1_otoGar Otolemur garnettii (bushbaby) COVE score: 25.30 auagcgcugc ucucugg ggggguuucauuc augau aguguuaccucu aa acuugcau gggggaacacc ugau gcc ccagaaa auccccugag >GPX1_tupBel Tupaia belangeri (tree_shrew) COVE score: 21.14 cagcgcugcg ucugggg gguuucaucc augac ggugucuccucu aa accccga aggaggaacgcc ugau guccggaaaa cccccca ggugggcgcc >GPX1_musMus Mus musculus (mouse) COVE score: 29.78 ggcugcacuc ugggggg cgguucuucc augau gguguuuccucu aa auuugca cggagaaacacc ugau uuccaggaa aaucccc ucagaugggc >GPX1_ratNor Rattus norvegicus (rat) COVE score: 25.72 ggcugcccuc cgggggg agguuuuucc augac gguguuuccucu aa auuuaca uggagaaacacc ugau uuccagaaa aaucccc ucagaugggc >GPX1_cavPor Cavia porcellus (guinea_pig) COVE score: 15.34 uugcccagau gcucucc ugaggguucuucc augaa gguguuuccucuc aa ccugua uagaggaacauc cgau uccca ggaauu ucccagagag >GPX1_oryCun Oryctolagus cuniculus (rabbit) COVE score: 24.38 guggccugcu gcucucu ggggguuucaucc augag ggcguucccccg aa aacaaa uggaggaacgcc ugau gucc gggaaac ccccaggugg >GPX1_eriEur Erinaceus europaeus (hedgehog) COVE score: 19.31 auagccgcga gccgcug ggcaucucaucc augac ggcgccgccuuc aa accugcgagcag gaaggagcgcc cgau agc cgcgaga gcccccagcg >GPX1_canFam Canis familiaris (dog) COVE score: 22.40 ucacagcgcu gcccucu ggggauuucaucc augau gguguuuccuug aa aucugca uggaggaacgcc ugau uucca ggaaagu ccccugagcu >GPX1_felCat Felis catus (cat) COVE score: 27.75 cacagcacug cccucug gggauuucaucc augau ggcguuuccucg aa auuugca uagaggaacgcc ugau uuc cagaag aauccccuga >GPX1_equCab Equus caballus (horse) COVE score: 27.08 ucacagugcu uuucucu ggggauuucaucc augau ggcguuuccucu aa acaugc augaggaacgcc ugau guu aaggaga aucccccgag >GPX1_bosTau Bos taurus (cow) COVE score: 29.83 agucagcgcu gcucucc agggauuuugccc augaa gguguucccucu aa accuacg uggaggaaugcc ugau gucca ggaaaau ccccugaggu >GPX1_dasNov Dasypus novemcinctus (armadillo) COVE score: 28.92 uguccuccac acggggu uuucaucc augac gguguuuccucu aa accugca gggaggaacacc ugaa guccggcaaa aucccc cgagaugggu >GPX1_loxAfr Loxodonta africana (elephant) COVE score: 25.48 acuaggcggc uuuccgu ggggguuucauua augag gguguuuucucu aa accugaa uggaggaacacc ugau guc ugggaa gauacccccc >GPX1_monDom Monodelphis domestica (opossum) COVE score: 26.38 gugcuaaggg uccguga ggguuuuaucu augau gguguuguuuu aa accauuaagga gaaagaacacu ugau aaugc uuguaa aaucccauga >GPX1_ornAna Ornithorhynchus anatinus (platypus) COVE score: 25.55 ugcuauuugc ccauggg ggagaucucauuu augau ggugcuccuucu aa accuuua cggaagagcacu ugau gga uccugag gaacuccaug
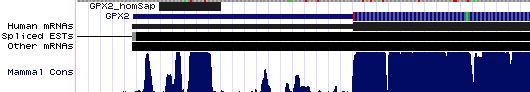
GPX2: 17 SECIS insertion sequences
>GPX2_homSap Homo sapiens (human) COVE score: 29.54 AGACUUGGGU AAGCUCU GGGCCUUCACAGA AUGAU GGCACCUUCCU AA ACCCUCA UGGGUGGUGUC UGAG AGGCGUGA AGGGCCU GGAGCCACUC >GPX2_panTro Pan troglodytes (chimp) COVE score: 29.54 agacuugggu aagcucu gggccuucacaga augau ggcaccuuccu aa acccuca ugggugguguc ugag aggcguga agggccu ggagccacuc >GPX2_macMul Macaca mulatta (rhesus) COVE score: 29.54 agacuugggu aagcucu gggccuucacaga augau ggcaccuuccu aa acccuca ugggugguguc ugag aggcguga agggccu gaagccaccc >GPX2_otoGar Otolemur garnettii (bushbaby) COVE score: 27.86 agacuugggu gggcucu gggccuucacaga augau ggcaccauccu aa acgccuc ugggugguguc ugag aagagcg gaaggcc uggagccacc >GPX2_tupBel Tupaia belangeri (tree_shrew) COVE score: 28.69 agacuuaggu gggcucu gagccuucacaga augau agcaccuuccu aa acccccc cgggagguguc ugag aagugug acaggcc cggagccagc >GPX2_musMus Mus musculus (mouse) COVE score: 33.65 agucuggggu agguucu gggccuucacaga augau ggcaucuuccu aa acccuuc ugggagauguc ugag aaguugug aaggguc cagagccagu >GPX2_ratNor Rattus norvegicus (rat) COVE score: 30.80 cugggguagg ugcuagg ccuucuucacaga augau ggcaucuuccu aa acccuuc ugggggauguc ugag acg uugugaa gggcccagag >GPX2_cavPor Cavia porcellus (guinea_pig) COVE score: 20.08 cuaggacagg uggcucu gggccuucacaga augac ggcaccguccu aa acgcua ugggugguauc ugag aagugug aauggcug gagccagccu >GPX2_sorAra Sorex araneus (shrew) COVE score: 29.00 aagcuggggc aaacucu gggccuucgcaga augau ggcaccucccu aa auccau ggggugguguc ugag gcgugcg agggccu ggaaacagcc >GPX2_eriEur Erinaceus europaeus (hedgehog) COVE score: 25.06 aaagcugggu gguuucu ggaccuucacaga augau agcaccuuccu aa accuaua gggaugguguc ugag aaaugu gaagggc cugaaguaaa >GPX2_canFam Canis familiaris (dog) COVE score: 22.57 aagacuuggg uggcucu gggccuucacaga augau ggcaccuuccu aa auagua ugggcgguguc ugag aagugug aagggcu cagagccagc >GPX2_felCat Felis catus (cat) COVE score: 22.45 aagacuuggg uggaucu gggccuucacaga augau ggcaucuuccu aa acugua ugggcgguguc ugag gaguguga agggcu cggagccagc >GPX2_equCab Equus caballus (horse) COVE score: 28.45 agacuucggu aggcucu gggccuucacaga augau ggcaccuuccu aa accugua uggacgguguc ugaa aagcgug aagggcc ccgagucagc >GPX2_bosTau Bos taurus (cow) COVE score: 26.14 agacaugggu gggcucu gugucuucacaga augau ggcaccuuccu aa aucugua ugggcgguguu ugag aagagu gaaggcc uggagccagc >GPX2_loxAfr Loxodonta africana (elephant) COVE score: 27.08 agacuugggu aggcucu ggaccuucgcaga augau ggcaccuuccu aa acucag ugggugguguc ugag aaaugug aagggccu agggccagcc >GPX2_echTel Echinops telfairi (tenrec) COVE score: 29.95 agugcuggug uggcucu gggccuucacaga augau ggcaccuuccu aa acccuc cggaagguguc ugag aaaugug aagggcc uggggccggc >GPX2_monDom Monodelphis domestica (opossum) COVE score: 26.38 gugcuaaggg uccguga ggguuuuaucu augau gguguuguuuu aa accauuaagga gaaagaacacu ugau aaugc uuguaa aaucccauga >GPX2_ornAna Ornithorhynchus anatinus (platypus) COVE score: 27.55 gggcucgagc agccucc agaccuucacaga augac ggugucuccuu aa acccuaac cgggaggcacc cgag agccggu gaagggc cuggugacug
GPX4: 14 SECIS sequences
>GPX4_SECIS_homSap Homo sapiens (human) chr19:1,057,512-1,057,793 COVE score: 29.19 CCACGCCCUU GGAGCCU UCCACCGGCACUC AUGAC GGCCUGCCUG CA AACCUGCUGGU GGGGCAGACC CGAA AAUCC AGCGUGC ACCCCGCCGG >GPX4_SECIS_panTro Pan troglodytes (chimp) COVE score: 29.19 CCACGCCCUC GGAGCCU UCCACCAGCACCC AUGAc ggccugccug ca aaccugcuggu ggggcagacc cgaa aaucc agcgugc acccugccgg >GPX4_SECIS_macMul Macaca mulatta (rhesus) COVE score: 33.09 CCACGCCCUC GGAGCCU UCCACCGGCACCC AUGAc ggccugcuugc aa accagcuccuggu gaggcagacc cgaa aaucc agcgugc accccgcugg >GPX4_SECIS_tupBel Tupaia belangeri (tree_shrew) COVE score: 31.05 CCGCGCCCCG GAACCUU CCACCCGGCACCU UUGAc ggucugccuau aa accugccacug gugaggcagacc cgag aaccu ggcgugc acccugccgg >GPX4_SECIS_musMus Mus musculus (mouse) COVE score: 35.62 UGACCCCUGG AGCCUUC CACCCCGGCACUC AUGAa ggucugccug aa aaccagccugcuggu ggggcagucc ugag gaccu ggcgugc acccugccgg >GPX4_SECIS_ratNor Rattus norvegicus (rat) COVE score: 34.33 UGACCCCUGG AGCCUUC CACCCCGGCACUC AUGAc ggucugccug aa aaccagcccgcuggu ggggcagucc cgag gaccu ggcgugc accccgccgg >GPX4_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 34.46 UGGCCGCCCC GAGUCUC CUACCCGGGUGCC AUGAc ggccugccug ca aaccagcgug cugguggggc agac ccg aggaugc gugcacugcc >GPX4_SECIS_canFam Canis familiaris (dog) COVE score: 31.12 AUGCCCCUCG GAGCCUU CCACCCGGCACCC AUGAc agucugucuaa aa accagcccgcugg uggggcagacc cgag aaccc ggcgugc acccugccgg >GPX4_SECIS_sorAra Sorex araneus (shrew) COVE score: 36.85 CUGCCCCUCG GAGCCUU CCACCCGGCGCCC AUGAc ggucugccugc aa accagccc gcugguggggc agac ccgagaaccc ggcgugc accuugccgg >GPX4_SECIS_bosTau Bos taurus (cow) COVE score: ? tgttccgagaAGCCTTCCCCCGGtgCctATGACGGTCTGtCTGAAAACCAGCCCCTGGTGGGGCAGaCCtGAGaACCTGGCGTGCACtccttccgg >GPX4_SECIS_echTel Echinops telfairi (tenrec) COVE score: 40.88 UUGAUCCCCA GAGUCUU CCACCCGGCACCA AUGAc ggucugccuuc aa accaggccucugg ugaggcagacc cgau gaccc ggcgugc acucagccgg >GPX4_SECIS_monDom Monodelphis domestica (opossum) COVE score: 40.88 UUGAUCCCCA GAGUCUU CCACCCGGCACCA AUGAC GGUCUGCCUUC AA ACCAGGCCUCUGG UGAGGCAGACC CGAU GACCC GGCGUGC ACCUCAGCCG >GPX4_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 33.89 ACCUCCGCUU UCCCGGG ACGCUCUGCCUCC AUGAc ggccggccuu ca agccaaaaccaguugg uggggccggcc ugaa caa accggca cgggucccgg >GPX4_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 38.77 UGCUCCCCGU GGGCCCC CUCCUCCAGCACC AUGAc ggccugccuug aa gccagcuugcugg ugaggcagacc cgaa gauuc ggcgugc acugcuggag >GPX4_SECIS_galGal Gallus gallus (chicken) COVE score: 38.77 UGCUCCCCGU GGGCCCC CUCCUCCAGCACC AUGAc ggccugccuug aa gccagcuugcugg ugaggcagacc cgaa gauuc ggcgugc acugcuggag
GPX7: 0 SECIS sequences (all species have cysteine)
GPX8: 0 SECIS sequences (all species have cysteine or serine)
SEPW: 13 SECIS sequences
>SEPW_SECIS_homSap Homo sapiens (human) COVE score: 36.01 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAGUCUUGUGGACACC UGGUCUUUCCC UGAU GUU CUCGUGG CUGCUGUUGG >SEPW_SECIS_panTro Pan troglodytes (chimp) COVE score: 36.01 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAGUCUUGUGGACACC UGGUCUUUCCC UGAU GUU CUuGUGG CUGCUGUUGG >SEPW_SECIS_ponPyg Pongo pygmaeus (orang_sumatran)C OVE score: 30.55 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAAUCUUGUGGACACC UGGUCUUUCCC UGAU GUU CUCGUGG CUGCUGUUGG >SEPW_SECIS_macMul Macaca mulatta (rhesus) COVE score: 33.55 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAGUCUUGUGGACgCC UGGUCUUUCCC UGAU GUU CUCGUGG CUGCUGUUGG >SEPW_SECIS_musMus Mus musculus (mouse) COVE score: 34.04 UUGGCCCAGC CCCUCGU GGCAGACGCUUC AUGAU GGGAAGAACUG AA AUGUCUCGUGGACGC CUGGUCUUUCC CUGAU GUCCCU GCGACUG CCACGUAGGG >SEPW_SECIS_ ratNor Rattus norvegicus (rat) COVE score: 26.55 UGGCCGGCCU UUCUUGG CAGCCGCUUC AUGAC AGGAAGGACUG AA AUGUCUCAAAGACCUG UGGUCUUUCUU CGAU GUU CCUGCGG CCACCAAGUC >SEPW_SECIS_oryCun Oryctolagus cuniculus (rabbit) no SECIS element found AGTAACCTTGACCCAGCCCCTTTCATGCCTCAGCCTCGTCTCCATAGGCTAAGACTGGAGAAATGAGTCCCCTGAAGAACTGAAACTGGGGGTAGAGGGTTGGTGTTTTAAGATGTGGATGAGCTGGTCTTTAC >SEPW_SECIS_canFam Canis familiaris (dog) COVE score: 34.04 UUGGCCCAGC CCCUCGU GGCAGACGCUUC AUGAU GGGAAGAACU GA AAUGUCUCGUGGACGCC UGGUCUUUCC CUGAU GUCCCU GCGACUG CCACGUAGGG >SEPW_SECIS_susScr Sus scrofa (pig) no SECIS element found AGTAACCTTGACCCAGCCCCTTTCATGCCTCAGCCTCGTCTCCATAGGCTAAGACTGGAGAAGTCTTGTGGACGCCTGGTCTTTCCCTGATGTTCTCGTGGCTGCTGTTGGGGGCAGAGATGGATGAGCTGGTCTTTAC >SEPW_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 33.54 GGCCCAGCCC CUCUCAG CAGAUGCUUC AUGAC AGGAAGGACUG AA AUGUCUUGUGGACGCC UGGUCUUUCCC UGAU GUU CUUGUGG CUGCUGGUUG >SEPW_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 28.71 CCCAGCUGCC CUUGGCA GACGCUUC AUGAG GGGAAGGACCU AA AUGCGUCGUGGAUGCC UGGUCUUUCCC UGAU GCUCCUUCAC CUGCCAG AUGGGGCAGA >SEPW_SECIS_loxAfr Loxodonta africana (elephant) no SECIS element found GGGACCTTGGCCCAGCCCCTTTCAGCAGACACTTCATGACAGGAGGACTGAAATGTCTCCCAGACGCCTGGCTCTTTCCCTGAATCTGTCGGCTGCAGGACAGGGCAGCGGTTGACTCTCTCGTTTTTTGCAT >SEPW_SECIS_echTel Echinops telfairi (tenrec) no SECIS element found AGGCCAGAGACCTTGGCCCAGTCCCTCCATGACAGGCAGAACTGAAATGTCCTCTGGACAAGTGGTCTTTTCCAGAAACCCCAGGGCTGCTGGGCCGGAGCCGAGGCTGACAACCCTGGTCTTTGCCT
DIO1: 29 SECIS sequences
>DIO1_SECIS_homSap Homo sapiens (human) COVE score: 29 ttttaactctgtgtctttacatatttgtttatgatggccacagcctaaagtacacacggctgtgacttgattcaaaagaaaatgttataag >DIO1_SECIS_ponPyg Pongo pygmaeus (orang_sumatran) COVE score: 29 ttttaactctgtgtctttacatatttgtttatgatggccacagcctaaagtacacacggctgtgacttgattcaaaagaaaatgttataag >DIO1_SECIS_macMul Macaca mulatta (rhesus) COVE score: 29 tttcaactctgtgtctttacatatttgtttatgatggccacagcctaaagtacacacggctgtgacttgattcaaaagaaaatgttataag >DIO1_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 24 tgttaactctgcttcttttcatatttgttcatgacggtcacagtctaaagtacacacagctgtgacctgatttgaaagaaaatgttttaag >DIO1_SECIS_canFam Canis familiaris (dog) COVE score: - ttttaactctgcttcttttcatgtttgtctatgacggccacagcctaaagcacacacagctgtgacttgatttgaaagaaaatgttttaag >DIO1_SECIS_felCat Felis catus (cat) COVE score: - ttttaactctgcttcttttcacgtttgtctatgacggccacagtctaaagtgcacacagctgtgacttgacttgaacgaaaatgttttaag >DIO1_SECIS_bosTau Bos taurus (cow) COVE score: 28 ttttaactctgcctcttttcatatttgttcatgacggccacagcctaaagtacacacggctgtgacttgatttgaaagaaaatgttttaag >DIO1_SECIS_sorAra Sorex araneus (shrew) COVE score: 24 cggaaactcagcttctcttcatatttgtttatgacagccccagctgaaagtacacacagctgtggcttgattggaaagaaaatgttttaag >DIO1_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 26 tttaactctgctttcttctcatatttgcttatgatggtcacagcttaaagtatacacagctgtgacttgattggaaagaaaatattttaag >DIO1_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 29 ttttaactctgcttcttttcatatttgttcatgatggccacagcctaaagtacacacggctgtgacttgatttgaaagaaaatgttttaag >DIO1_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: - ttttaactctgcttcttttcatatttgtttatgatggccacagtttaaagtacatacagctgtgacttgatatgaaaaagaaatattttaag >DIO1_SECIS_loxAfr Loxodonta africana (elephant) COVE score: - ttttaactctgcttcttttttcatgtatttatgatgggccacagcctaaagtgcacaacagctgtgacttgatttgaaaaacatctttaag >DIO1_SECIS_monDom Monodelphis domestica (opossum) COVE score: - tttccatcctgcttctacaaatatttatttatgacaatcacagcctaaagctcagggcagctgggattcgacgggagaaaaagtttgtaag >DIO1_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 24 ccccggatccggttccgtgaatattggtttatgagggtcacagtgtaaagcgcatgcagctgtgacttgatctgagaaaatatttctgcggc >DIO2_SECIS_homSap Homo sapiens (human) 5,630 bp utr COVE score: 30 cagagatgtgcagagttgaccagtgtgcggatgataactactgacgaaagagtcatcgactcagttagtggttggatgtagtcacattagtttgcctctc >DIO2_SECIS_macMul Macaca mulatta (rhesus) COVE score: 30 cagagatgtgcagagttgaccagtgtgcggatgataactactgacgaaagagtcatcgactcagttagtggttggatgtagtcacattagtttgcctctc >DIO2_SECIS_musMus Mus musculus (mouse) COVE score: 29 cggagatgttcagagctcactggtgtgcgaatgataactactgacgaaagagctgtctgctcagtctgtggttggatgtagtcacacgagtctgcctttctgca >DIO2_SECIS_ratNor Rattus norvegicus (rat) COVE score: 28 ccgagatgttcggagctcactggtgtgcgaatgataactactgacgaaagagtcatctgctcagtctgtggttggatgtagtcacacgagtctgcctctccatc >DIO2_SECIS_canFam Canis familiaris (dog) COVE score: 27 ctgggatgtgcagaggtgaccagtgtgcgaatgataactactgatgaaagagtcactgactcagttagtggttggatacagtcacattagttttcctct >DIO2_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 30 ctgggatgtgcagaggtgaccagtgtgcaaatgataactactgatgaaagagtcattgactcagttagtggttggatgtagtcacattagtttgcctctc >DIO2_SECIS_dasNov Dasypus novemcinctus (armadillo) iodothyronine deiodinase type II ctgggaagttcagaggctaccagtgtgccaatgataactactgacgaaagaggcatcgactcagttagtggttggatgtagccacattagtttgcctctc >DIO3_SECIS_homSap Homo sapiens (human) COVE score: 31 ttgggtgcacaggagccccactgctgatgacgaactatctctaactggtcttgaccacgagctagttctgaattgcaggggcctcaaagcagca >DIO3_SECIS_macMul Macaca mulatta (rhesus) COVE score: 30 ttgggtgcacaggagccccactgctgatgacgaactgtctctaactggtcttgaccacgagctagttctgaattgcaggggcctcaaaacagca >DIO3_SECIS_musMus Mus musculus (mouse) COVE score: 26 ttgggtgcgctggagccctggctgctgatgacgaaccgcctctaactgggcttgaccacgggtcggctctgaattgcagagaggctcgaaacagc >DIO3_SECIS_ratNor Rattus norvegicus (rat) COVE score: 26 ttgggtgcgctggagccctggctgctgatgacgaaccgcctctaactgggcttgaccacgggtcggctctgaattgcagagaggctcgaaacagc >DIO3_SECIS_canFam Canis familiaris (dog) COVE score: 26 ttgggtgctggcgagccccactgctgatgacgagccgcctctaactggtcttgaccacgagctggttctgagttgcaggggggcttgcagcggc >DIO3_SECIS_bosTau Bos taurus (cow) COVE score: 30 ttgggtgctcacgagccccactgctgatgaagagctgtctctaactggcctcgaccacgagctggttctgatttgcaggaggctcgcagcagc >DIO3_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 27 ttgggtgctcaggagccccactgctgatgacgaactgtctctaactggtcttgaccacgagctggttctgaattgcagggggctcgcagcagca >DIO3_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 22 ttcggtgcgctagagccccactgctgatgacgaactgtctctaactggtcttgaccacgagctgattccgaattgcagggaactcgcagcagc >DIO3_SECIS_echTel Echinops telfairi (tenrec) COVE score: - ttcggtgctctgcagccccactgctgatgacgaactgcctctcactggtcttgaccacgagctgcttctgaaatgcaggggactcgcagccgca