Uploads by Hiram
From genomewiki
Jump to navigationJump to search
This special page shows all uploaded files.
| Date | Name | Thumbnail | Size | Description | Versions |
|---|---|---|---|---|---|
| 16:33, 28 June 2022 | DanRer11.grcIncidentDb.bb (file) | 202 KB | gwUploadFile upload | 17 | |
| 16:33, 28 June 2022 | DanRer10.grcIncidentDb.bb (file) | 126 KB | gwUploadFile upload | 127 | |
| 16:33, 28 June 2022 | Mm9.grcIncidentDb.bb (file) | 74 KB | gwUploadFile upload | 282 | |
| 16:33, 28 June 2022 | DanRer7.grcIncidentDb.bb (file) | 111 KB | gwUploadFile upload | 336 | |
| 16:33, 28 June 2022 | Mm10.grcIncidentDb.bb (file) | 101 KB | gwUploadFile upload | 289 | |
| 16:33, 28 June 2022 | Hg19.grcIncidentDb.bb (file) | 106 KB | gwUploadFile upload | 867 | |
| 22:32, 20 May 2021 | VGP 2021-05-21.pptx (file) | 1.24 MB | VGP meeting 2021-05-21 presentation, VGP assembly resources in the UCSC genome browser | 1 | |
| 17:24, 3 September 2019 | GalGal6.grcIncidentDb.bb (file) | 19 KB | gwUploadFile upload | 1 | |
| 00:56, 19 May 2019 | NationalYoYoMuseum ChicoCA 50.jpg (file) |  |
553 KB | National YO YO Museum, Chico CA | 1 |
| 17:33, 30 January 2019 | GalGal5.grcIncidentDb.bb (file) | 27 KB | gwUploadFile upload | 7 | |
| 21:54, 1 October 2018 | AwsAttachVolume.sh.txt (file) | 4 KB | Script for AWS instance initialization to attach a data volume to a running instance. Used in the startParaHub.sh script upon starting the machine. | 1 | |
| 17:35, 26 September 2018 | OpenStackParaHubSetup.sh.txt (file) | 8 KB | reorganize NFS setups, parameterize the installation directory | 5 | |
| 17:10, 26 September 2018 | InitParasol.sh.txt (file) | 3 KB | less restrictive subnet specification and separate hub/node logs in ./logs/ directory | 3 | |
| 20:32, 25 September 2018 | NodeReport.sh.txt (file) | 1 KB | add usage message and directory argument for result file | 4 | |
| 17:42, 11 July 2018 | OpenStackParaNodeSetup.sh.template.txt (file) | 4 KB | correctly fixup /mnt filesystem for broken openstack installation, and mount /mnt/data from the hub | 2 | |
| 17:09, 28 April 2018 | LastzProcessingTimeHistogram.png (file) |  |
53 KB | lastz cluster compute time histogram 1,635 measurements from alignments performed at UCSC 459 measurements are less than 0.05 days (less than 1 hour) due to very small genome sizes 59 measurements are longer than 5 days processing time due to high c... | 1 |
| 23:13, 27 April 2018 | SizeVsTime.png (file) |  |
156 KB | lastz cluster run processing time vs. genome size blue points plot times 2 days or less processing time, left axis scale 0 - 2 days red points plot times over 2 days processing time, right axis scale 0 - 70 days | 1 |
| 02:53, 6 April 2018 | ParasolInstall.sh.txt (file) | 1 KB | adding git fetch of automation scripts into /data/scripts/ | 3 | |
| 22:10, 28 September 2017 | GenbankPrimates.jpg (file) | 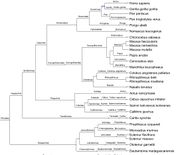 |
113 KB | primate phylogenetic tree from genbank taxIds Given a set of taxon IDs, utilizing [http://phylot.biobyte.de/ phylot.biobyte.de] to derive this phylogenetic tree. NOTE: not necessarily completely accurate. Image drawn by [http://itol.embl.de/ itol.embl. | 1 |
| 18:17, 2 June 2017 | Btrfs filesystem readWrite performance.png (file) |  |
120 KB | read/write performance test results for 'btrfs' filesystem | 1 |
| 22:48, 31 May 2017 | Xfs filesystem readWrite performance.png (file) |  |
97 KB | testing read/write performance on xfs filesystem in a directory overloaded with up to 7 million files | 1 |
| 22:47, 31 May 2017 | Tmpfs filesystem readWrite performance.png (file) |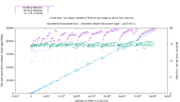  |
101 KB | testing read/write performance into a /dev/shm directory overloaded with up to 7 million files | 1 |
| 22:45, 31 May 2017 | Ext4 fs readWrite performance.png (file) |  |
106 KB | writing over 7 million files into one directory to see how performance degrades. | 1 |
| 18:26, 26 January 2017 | BME230 Winter 2017.ppt (file) | 12.98 MB | (Hiram's presentation to BME230 Winter 2016 12 January - brief browser usage introduction, assembly hub demonstration, table browser intersection, plus information on source code but not part of the discussion) | 1 | |
| 05:39, 9 June 2016 | BejeranoLab 2016-06-09.pptx (file) | 2.92 MB | 2 | ||
| 18:22, 12 January 2016 | BME230 Winter 2016.ppt (file) | 13.8 MB | (Hiram's presentation to BME230 Winter 2016 12 January - brief browser usage introduction, assembly hub demonstration, table browser intersection, plus information on source code but not part of the discussion)) | 1 | |
| 17:43, 19 March 2015 | MatrixSummary pl.txt (file) | 6 KB | used in lastz tuning procedure, examines output of the four lastz_D outputs to survey the differences | 1 | |
| 17:17, 19 March 2015 | Create scores file control.txt (file) | 653 bytes | Parameters from Bob Harris for lastz tuning procedure | 1 | |
| 17:14, 19 March 2015 | Expand scores file py.txt (file) | 4 KB | Script created by Bob Harris for lastz tuning operations | 1 | |
| 17:10, 19 March 2015 | AdjustSizes pl.txt (file) | 3 KB | chrom.sizes files are now expected to exist as created by selectedFasta.sh | 2 | |
| 17:09, 19 March 2015 | SelectedFasta sh.txt (file) | 2 KB | generalize the chrom.size creation and noMask option on fasta creation | 2 | |
| 23:34, 18 March 2015 | TopAll sh.txt (file) | 2 KB | script used in lastz tuning procedure to run lastz_D on different sets of DNA sequences from query and target | 1 | |
| 23:18, 18 March 2015 | MafScoreSizeScan pl.txt (file) | 2 KB | script used to scan a MAF file to extract scores, sizes, and coverage statistics of each alignment block. Used in lastz tuning procedures. | 1 | |
| 20:59, 4 February 2014 | BME230 Winter 2014.ppt (file) | 13.79 MB | Hiram's presentation to BME230 Winter 2014 11 February - brief browser usage introduction, assembly hub demonstration, table browser intersection, plus information on source code but not part of the discussion) | 1 | |
| 22:36, 6 June 2013 | AssemblyHubs BoG 2013.pptx (file) | 2.72 MB | Assembly Hubs, Biology of Genomes meeting 2013 CSHL | 1 | |
| 22:35, 6 June 2013 | AssemblyHubs BoG 2013.pdf (file) | 3.06 MB | Assembly Hubs, Biology of Genomes meeting 2013 CSHL | 1 | |
| 05:43, 13 March 2013 | SameSpeciesChainNet.sh.txt (file) | 5 KB | Same species lift over, step 2 of 2, chain/net the blat psl results together into the result file | 1 | |
| 05:04, 13 March 2013 | BlatJob.csh.txt (file) | 2 KB | script to use for the same species lift over procedure run.blat/ step 1 | 1 | |
| 04:57, 13 March 2013 | SameSpeciesBlatSetup.sh.txt (file) | 6 KB | First step in the same species lift over procedure, set up for blat run, adding 11.ooc file construction for blat | 2 | |
| 02:56, 29 June 2012 | FireCracker10KElevationProfile.png (file) |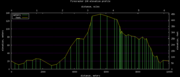  |
6 KB | Elevation profile for the Santa Cruz FireCracker 10K run, July 4th. Corrected the GPS readings with Google Earth information and known ground truth. | 2 |
| 16:44, 27 April 2012 | DanRer7.ncbiIncidentDb.bb (file) | 168 KB | gwUploadFile upload | 27 | |
| 16:43, 27 April 2012 | Mm9.ncbiIncidentDb.bb (file) | 136 KB | gwUploadFile upload | 80 | |
| 16:43, 27 April 2012 | Hg19.ncbiIncidentDb.bb (file) | 179 KB | gwUploadFile upload | 67 | |
| 21:27, 25 January 2012 | BME230 Winter 2012.ppt (file) | 7.31 MB | Hiram's presentation to BME230 Winter 2012 26 January 2012 - brief browser usage introduction, table browser intersection, plus information on source code but not part of the discussion | 1 | |
| 21:51, 5 October 2011 | XLDB 2011.pptx (file) | 1.23 MB | bigBed/bigWig 5 minute presentation at XLDB conference, Stanford, 18 October 2011. | 2 | |
| 18:48, 24 August 2011 | Sol GCBias GeneCats 2011-08-24.pdf (file) | 1.4 MB | Sol Katzman presentation, genecats 24 Aug 2011 | 1 | |
| 16:43, 24 June 2011 | ExtractRepeats.txt (file) | 2 KB | Small perl script used to extract lineage specific repeats from RepeatMasker .out files | 1 | |
| 20:14, 22 March 2011 | 29mammalsConstrainedElements pi lods.bw (file) | 61.15 MB | gwUploadFile upload | 1 | |
| 20:13, 22 March 2011 | 29mammalsConstrainedElements pi branchLength.bw (file) | 61.3 MB | gwUploadFile upload | 1 | |
| 20:13, 22 March 2011 | 29mammalsConstrainedElements omega lods.bw (file) | 54.74 MB | gwUploadFile upload | 1 |