Uploads by Hiram
From genomewiki
Jump to navigationJump to search
This special page shows all uploaded files.
| Date | Name | Thumbnail | Size | Description | Versions |
|---|---|---|---|---|---|
| 17:47, 16 December 2008 | Metcalf.Richardson.2008-10-15.pdf (file) | 3.26 MB | Jake Metcalf and Sarah Richardson, Genecats presentation 15 Oct 2008, ethics discussion of Bruce T. Lahn controversy from U. Chicago. | 1 | |
| 16:33, 28 June 2022 | Mm10.grcIncidentDb.bb (file) | 101 KB | gwUploadFile upload | 289 | |
| 16:36, 18 September 2007 | Mm8 mm9.exonCount.png (file) |  |
5 KB | Mouse mm8 and mm9 exon count by size histogram, to size 30 nucleotides | 2 |
| 16:38, 18 September 2007 | Mm8 mm9.exonSize.png (file) |  |
5 KB | Mouse mm8 and mm9 exon size histogram, to size 50 nucleotides | 2 |
| 16:39, 18 September 2007 | Mm8 mm9.intronSize.png (file) |  |
5 KB | Mouse mm8 and mm9 intron size histogram, to size 50 nucleotides | 2 |
| 16:41, 18 September 2007 | Mm8 mm9.intronsTo170.png (file) |  |
5 KB | Mouse mm8 and mm9 intron size histogram, to size 170 nucleotides | 2 |
| 16:33, 18 September 2007 | Mm8 mm9 exonsTo300.png (file) |  |
6 KB | Mouse mm8 and mm9 exon size histogram to size 300 nucleotides | 2 |
| 16:33, 28 June 2022 | Mm9.grcIncidentDb.bb (file) | 74 KB | gwUploadFile upload | 282 | |
| 16:43, 27 April 2012 | Mm9.ncbiIncidentDb.bb (file) | 136 KB | gwUploadFile upload | 80 | |
| 22:04, 3 December 2007 | Mm9Euarchontoglires.png (file) |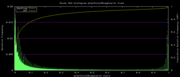  |
5 KB | Mm9 alternative phastCons data histogram, Euarchontoglires, mm9,rn4,cavPor2,oryCun1,hg18,panTro2,ponAbe2,rheMac2,calJac1,otoGar1,tupBel1 | 3 |
| 22:03, 3 December 2007 | Mm9Placental.png (file) |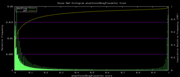  |
5 KB | Mm9 alternative phastCons data histogram, Placentals, =mm9,rn4,cavPor2,oryCun1,hg18,panTro2,ponAbe2,rheMac2,calJac1,otoGar1,tupBel1,sorAra1,eriEur1,canFam2,felCat3,equCab1,bosTau3,dasNov1,loxAfr1,echTel1 | 3 |
| 23:21, 10 October 2007 | Mm9 30way.gif (file) |  |
8 KB | Attempting to reinitialize this image to get it correct | 1 |
| 22:14, 14 October 2009 | MonsatoIntroduction Oct2009.pdf (file) | 8.37 MB | Montsanto Introduction, Genecats 14 Oct 2009 | 1 | |
| 18:55, 17 December 2008 | MouseAnnotAndCCDSVisitReport041608.ppt (file) | 2.34 MB | Rachel Harte summarizing Mouse Genome Annotation meeting and RefSeq/CCDS groups, NIH March 2008. | 1 | |
| 22:28, 17 September 2009 | MultAlign.jpg (file) |  |
14 KB | Example tool tip pop up image | 1 |
| 00:56, 19 May 2019 | NationalYoYoMuseum ChicoCA 50.jpg (file) |  |
553 KB | National YO YO Museum, Chico CA | 1 |
| 20:32, 25 September 2018 | NodeReport.sh.txt (file) | 1 KB | add usage message and directory argument for result file | 4 | |
| 17:35, 26 September 2018 | OpenStackParaHubSetup.sh.txt (file) | 8 KB | reorganize NFS setups, parameterize the installation directory | 5 | |
| 17:42, 11 July 2018 | OpenStackParaNodeSetup.sh.template.txt (file) | 4 KB | correctly fixup /mnt filesystem for broken openstack installation, and mount /mnt/data from the hub | 2 | |
| 17:00, 15 January 2023 | PAG30 2023-01-14.pptx (file) | 9.85 MB | presentation at the Plant And Animal Genomes conference, San Diego January 2023 - "The UCSC Genome Browser" - an introduction to the browser, GenArk assembly hubs, building your own assembly hub and REST API interface. | 1 | |
| 17:05, 15 January 2023 | PAG30 2023-01-15.pptx (file) | 7.55 MB | presentation at the 30th annual Plant and Animal Genomes conference - "Educational Resources in the UCSC Genome Browser" - introducing the http://genome.ucsc.edu/training/education/ resources. | 1 | |
| 21:53, 7 December 2007 | PageCount.png (file) | 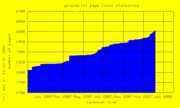 |
8 KB | Genomewiki page count graph | 1 |
| 02:53, 6 April 2018 | ParasolInstall.sh.txt (file) | 1 KB | adding git fetch of automation scripts into /data/scripts/ | 3 | |
| 18:28, 30 October 2007 | PhastCons30way.png (file) |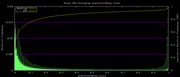  |
5 KB | Mouse Mm9 30-way conservation phastCons histogram plot for the track data phastCons30way | 1 |
| 19:24, 1 December 2009 | PhastCons46way.histogram.png (file) |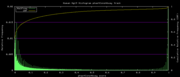  |
5 KB | 46-way conservation track on human genome browser hg19. phastCons histogram data for all 46 vertebrates | 1 |
| 19:24, 1 December 2009 | PhastCons46wayPlacental.histogram.png (file) |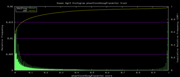  |
5 KB | 46-way conservation track on human genome browser hg19. phastCons histogram data for all placental mammal subset | 1 |
| 19:25, 1 December 2009 | PhastCons46wayPrimates.histogram.png (file) |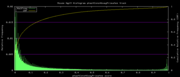  |
5 KB | 46-way conservation track on human genome browser hg19. phastCons histogram data for the primates subset | 1 |
| 18:30, 28 April 2006 | PhenSort.gif (file) | 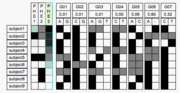 |
16 KB | Example phenotype | 1 |
| 19:35, 30 November 2009 | PhyloP46way.histogram.png (file) |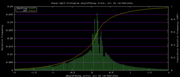  |
6 KB | 46-way conservation track on human genome browser hg19. phyloP histogram data for all 46 vertebrates | 1 |
| 19:36, 30 November 2009 | PhyloP46wayPlacental.histogram.png (file) |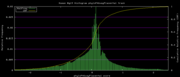  |
6 KB | 46-way conservation track on human genome browser hg19. phyloP histogram data for all placental mammal subset | 1 |
| 19:37, 30 November 2009 | PhyloP46wayPrimates.histogram.png (file) |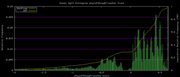  |
6 KB | 46-way conservation track on human genome browser hg19. phyloP histogram data for all primates subset | 1 |
| 04:57, 13 March 2013 | SameSpeciesBlatSetup.sh.txt (file) | 6 KB | First step in the same species lift over procedure, set up for blat run, adding 11.ooc file construction for blat | 2 | |
| 05:43, 13 March 2013 | SameSpeciesChainNet.sh.txt (file) | 5 KB | Same species lift over, step 2 of 2, chain/net the blat psl results together into the result file | 1 | |
| 22:18, 8 January 2024 | Sat Workshop2024.pptx (file) | 4.45 MB | PAG 31 - 2024 - San Diego Plant and Animal Genomes - GenArk assembly resources | 1 | |
| 17:09, 19 March 2015 | SelectedFasta sh.txt (file) | 2 KB | generalize the chrom.size creation and noMask option on fasta creation | 2 | |
| 23:13, 27 April 2018 | SizeVsTime.png (file) |  |
156 KB | lastz cluster run processing time vs. genome size blue points plot times 2 days or less processing time, left axis scale 0 - 2 days red points plot times over 2 days processing time, right axis scale 0 - 70 days | 1 |
| 17:37, 10 August 2006 | SmedleyCat.jpg (file) |  |
11 KB | Smedley the cat in the early days. Photo by Hiram Clawson, 1995. | 1 |
| 06:14, 13 July 2008 | SoftwareSermon2002 Jim Kent.ppt (file) | 113 KB | Genecats October 2002 - "Software Sermon" - software development philosophy and guidelines, Jim Kent. | 1 | |
| 18:48, 24 August 2011 | Sol GCBias GeneCats 2011-08-24.pdf (file) | 1.4 MB | Sol Katzman presentation, genecats 24 Aug 2011 | 1 | |
| 18:50, 10 August 2006 | StarTrekTOSTransporter.wav (file) | 223 KB | Testing wav file upload. Star Trek, The Original Series, Transporter sound. 10 seconds | 1 | |
| 18:46, 29 July 2009 | SvnForGb.pdf (file) | 171 KB | Mark's presentation to 2009-07-29 genecats, review of SVN subversion and how it could replace CVS | 1 | |
| 18:27, 25 March 2009 | Teal Day 05.jpg (file) |  |
1.02 MB | Trying to recover a mysterious bogus file directory link | 1 |
| 19:00, 10 August 2006 | TestCustomTrack.data.txt.gz (file) | 2 KB | Testing file upload feature. This is a custom track test file demonstrating all track types currently (August 2006) supported by the browser custom tracks. | 1 | |
| 21:41, 17 March 2008 | TestOOPresent.ppt (file) | 11 KB | Testing ppt upload | 1 | |
| 22:47, 31 May 2017 | Tmpfs filesystem readWrite performance.png (file) |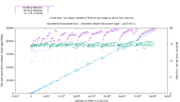  |
101 KB | testing read/write performance into a /dev/shm directory overloaded with up to 7 million files | 1 |
| 23:34, 18 March 2015 | TopAll sh.txt (file) | 2 KB | script used in lastz tuning procedure to run lastz_D on different sets of DNA sequences from query and target | 1 | |
| 21:40, 7 December 2007 | UserCount.png (file) | 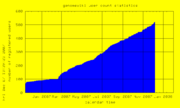 |
9 KB | Genomewiki User Count graph | 1 |
| 22:32, 20 May 2021 | VGP 2021-05-21.pptx (file) | 1.24 MB | VGP meeting 2021-05-21 presentation, VGP assembly resources in the UCSC genome browser | 1 | |
| 21:51, 5 October 2011 | XLDB 2011.pptx (file) | 1.23 MB | bigBed/bigWig 5 minute presentation at XLDB conference, Stanford, 18 October 2011. | 2 | |
| 22:48, 31 May 2017 | Xfs filesystem readWrite performance.png (file) |  |
97 KB | testing read/write performance on xfs filesystem in a directory overloaded with up to 7 million files | 1 |