File list
From genomewiki
Jump to navigationJump to search
This special page shows all uploaded files.
| Date | Name | Thumbnail | Size | User | Description | Versions |
|---|---|---|---|---|---|---|
| 13:01, 28 September 2008 | BadTerminol.jpg (file) | 5 KB | Tomemerald | 1 | ||
| 16:41, 18 September 2007 | Mm8 mm9.intronsTo170.png (file) |  |
5 KB | Hiram | Mouse mm8 and mm9 intron size histogram, to size 170 nucleotides | 2 |
| 16:38, 18 September 2007 | Mm8 mm9.exonSize.png (file) |  |
5 KB | Hiram | Mouse mm8 and mm9 exon size histogram, to size 50 nucleotides | 2 |
| 04:21, 6 May 2008 | 28wayPhylo.png (file) | 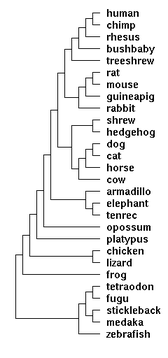 |
5 KB | Tomemerald | 1 | |
| 16:46, 18 September 2007 | Hg17 hg18.intronsTo170.png (file) |  |
5 KB | Hiram | Human hg17 and hg18 intron size to 170 nucleotides | 2 |
| 16:45, 18 September 2007 | Hg17 hg18.exonSize.png (file) |  |
5 KB | Hiram | Human hg17 and hg18 exon size histogram, to size 50 nucleotides | 2 |
| 04:57, 13 March 2013 | SameSpeciesBlatSetup.sh.txt (file) | 6 KB | Hiram | First step in the same species lift over procedure, set up for blat run, adding 11.ooc file construction for blat | 2 | |
| 17:43, 19 March 2015 | MatrixSummary pl.txt (file) | 6 KB | Hiram | used in lastz tuning procedure, examines output of the four lastz_D outputs to survey the differences | 1 | |
| 19:36, 30 November 2009 | PhyloP46wayPlacental.histogram.png (file) |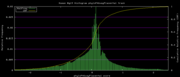  |
6 KB | Hiram | 46-way conservation track on human genome browser hg19. phyloP histogram data for all placental mammal subset | 1 |
| 19:37, 30 November 2009 | PhyloP46wayPrimates.histogram.png (file) |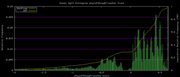  |
6 KB | Hiram | 46-way conservation track on human genome browser hg19. phyloP histogram data for all primates subset | 1 |
| 16:33, 18 September 2007 | Mm8 mm9 exonsTo300.png (file) |  |
6 KB | Hiram | Mouse mm8 and mm9 exon size histogram to size 300 nucleotides | 2 |
| 14:38, 25 March 2007 | UcscLDAPCookie.txt (file) | 6 KB | Fubar | Example of getting a local cookie set for mirror site sessions. LDAP authentication in this case. | 1 | |
| 02:56, 29 June 2012 | FireCracker10KElevationProfile.png (file) |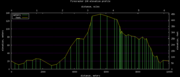  |
6 KB | Hiram | Elevation profile for the Santa Cruz FireCracker 10K run, July 4th. Corrected the GPS readings with Google Earth information and known ground truth. | 2 |
| 19:35, 30 November 2009 | PhyloP46way.histogram.png (file) |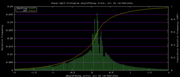  |
6 KB | Hiram | 46-way conservation track on human genome browser hg19. phyloP histogram data for all 46 vertebrates | 1 |
| 16:47, 18 September 2007 | Hg17 hg18 exonsTo300.png (file) |  |
6 KB | Hiram | Human hg17 and hg18 exon size histogram, to a size of 300 nucleotides | 2 |
| 17:36, 21 April 2008 | SELM SECIS.png (file) | 6 KB | Tomemerald | 1 | ||
| 16:28, 26 May 2008 | GPX geneTree.jpg (file) | 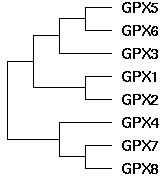 |
7 KB | Tomemerald | 1 | |
| 18:14, 4 May 2018 | OpenStackStartParaNode.sh.txt (file) | 7 KB | Chmalee | added command line argument for ssh key-pair | 2 | |
| 17:35, 26 September 2018 | OpenStackParaHubSetup.sh.txt (file) | 8 KB | Hiram | reorganize NFS setups, parameterize the installation directory | 5 | |
| 20:48, 30 September 2011 | GRIR3+Enhancers.bed (file) | 8 KB | Daria Merkurjev | 1 | ||
| 15:29, 15 June 2009 | DroPhylo.jpg (file) | 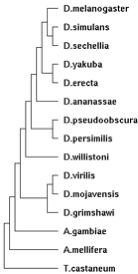 |
8 KB | Tomemerald | 1 | |
| 00:21, 25 November 2010 | UcscLinks.gif (file) | 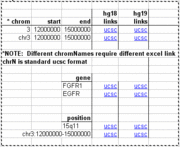 |
8 KB | Kuhn | Image showing functional portion of spreadsheet for making quick links to Genome Browser in Excel spreadsheet. | 1 |
| 18:13, 4 May 2018 | OpenStackStartParaHub.sh.txt (file) | 8 KB | Chmalee | Added command line argument for ssh key-pair name | 2 | |
| 14:21, 1 January 2010 | CilOpDecline.png (file) |  |
8 KB | Tomemerald | 1 | |
| 23:21, 10 October 2007 | Mm9 30way.gif (file) |  |
8 KB | Hiram | Attempting to reinitialize this image to get it correct | 1 |
| 21:53, 7 December 2007 | PageCount.png (file) | 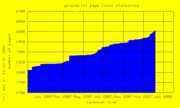 |
8 KB | Hiram | Genomewiki page count graph | 1 |
| 21:40, 7 December 2007 | UserCount.png (file) | 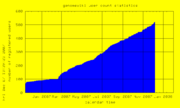 |
9 KB | Hiram | Genomewiki User Count graph | 1 |
| 15:23, 5 May 2008 | SS18L1 peg.png (file) | 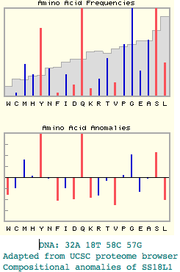 |
9 KB | Tomemerald | 1 | |
| 14:17, 1 January 2010 | TMTgeneTree.png (file) | 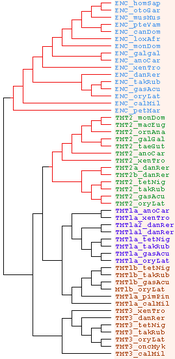 |
9 KB | Tomemerald | 1 | |
| 00:15, 21 May 2008 | SECIS GPX6.jpg (file) | 9 KB | Tomemerald | 1 | ||
| 16:57, 20 May 2008 | SECIS SEPP2.jpg (file) | 9 KB | Tomemerald | 1 | ||
| 13:24, 23 December 2009 | PolyphylOpsins.png (file) | 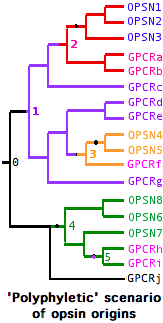 |
9 KB | Tomemerald | 5 | |
| 21:01, 2 September 2010 | CholineSulfate.png (file) | 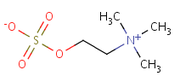 |
10 KB | Tomemerald | 1 | |
| 15:22, 6 June 2008 | SELM2 SECIS.jpg (file) |  |
10 KB | Tomemerald | 1 | |
| 15:01, 10 January 2009 | GaEvol.jpg (file) | 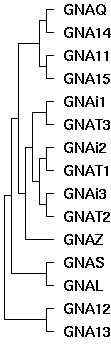 |
10 KB | Tomemerald | 1 | |
| 11:31, 13 February 2009 | MGAT5.jpg (file) |  |
11 KB | Tomemerald | 1 | |
| 15:27, 11 March 2008 | Opsin PER overall.png (file) | 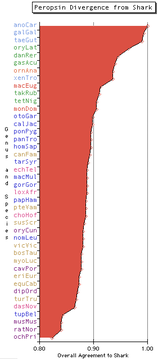 |
11 KB | Tomemerald | 1 | |
| 00:20, 14 February 2006 | G-Wiki.jpg (file) |  |
11 KB | Chesman | 2 | |
| 11:04, 17 November 2009 | PLATrates.jpg (file) |  |
11 KB | Tomemerald | 1 | |
| 15:16, 10 June 2009 | Ecdysozoa.jpg (file) |  |
11 KB | Tomemerald | 2 | |
| 15:16, 11 March 2008 | Opsin PER aveRates.png (file) |  |
11 KB | Tomemerald | 1 | |
| 15:18, 7 June 2008 | SEPHS2 SECIS.jpg (file) | 11 KB | Tomemerald | 1 | ||
| 16:30, 20 February 2008 | Opsin RGR earlySpl.png (file) |  |
11 KB | Tomemerald | 1 | |
| 21:41, 17 March 2008 | TestOOPresent.ppt (file) | 11 KB | Hiram | Testing ppt upload | 1 | |
| 16:19, 2 August 2009 | USH2A22homMus.jpg (file) | 11 KB | Tomemerald | 1 | ||
| 23:35, 9 April 2008 | D17 Subs chr2 MapUSC.png (file) |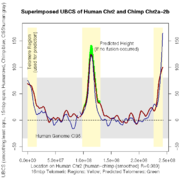  |
11 KB | Tdreszer | UBCS evidence of the mereger of Chr2a and Chr2b into Chr2. | 1 |
| 18:36, 3 June 2008 | SELH SECIS.jpg (file) | 11 KB | Tomemerald | 1 | ||
| 14:06, 16 February 2011 | Cousins.gif (file) | 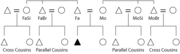 |
11 KB | Tomemerald | 1 | |
| 10:26, 15 March 2011 | LoadModencode.txt (file) | 11 KB | Max | script to load all chipseq data (~400 tracks) from modencode into a local UCSC installation automatically. Rename to .py before you use this. | 1 | |
| 07:28, 17 February 2011 | GenbankToTables.txt (file) | 11 KB | Max | script to convert genbank files to table format. Can identify the submitters of the genbank records. Please remove the .txt extension before running this. | 1 |