Unused files
From genomewiki
Jump to navigationJump to search
The following files exist but are not embedded in any page. Please note that other web sites may link to a file with a direct URL, and so may still be listed here despite being in active use.
Showing below up to 291 results in range #1 to #291.
View (previous 500 | next 500) (20 | 50 | 100 | 250 | 500)
- G-Wiki.png 120 × 60; 2 KB
- G-Wiki.jpg 300 × 150; 11 KB
- SmedleyCat.jpg 151 × 336; 11 KB
- StarTrekTOSTransporter.wav ; 223 KB
- TestCustomTrack.data.txt.gz ; 2 KB
- ParseBlatOutput zip.txt ; 1 KB
- TomPringle1.JPG 2,304 × 1,728; 985 KB
- TomPringle2.JPG 2,304 × 1,728; 987 KB
- TomPringle4.JPG 2,304 × 1,728; 995 KB
- TomPringle5.JPG 1,728 × 2,304; 1.38 MB
- TomPringle6.JPG 1,728 × 2,304; 1.26 MB
- TomPringle7.JPG 1,728 × 2,304; 1.33 MB
- TomPringle3.JPG 2,304 × 1,728; 981 KB
- UcscLDAPCookie.txt ; 6 KB
- CleanHgCentral pl.txt ; 2 KB
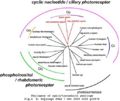
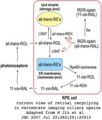
- Opsin phylo.png 577 × 323; 28 KB
- Opsin everse.png 347 × 437; 171 KB
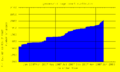
- UserCount.png 1,000 × 600; 9 KB
- PageCount.png 1,000 × 600; 8 KB
- Opsin nemato.png 213 × 310; 66 KB
- Opsin ancient cil introns.png 674 × 184; 28 KB
- Opsin RGR DRY.png 421 × 463; 47 KB
- Opsin PER aveRates.png 277 × 642; 11 KB
- TestOOPresent.ppt ; 11 KB
- SELM SECIS.png 960 × 156; 6 KB
- PhyloTime.png 695 × 603; 48 KB
- EmbedURL php.txt ; 3 KB
- Hgt genome test 5fbe 182210.pdf ; 26 KB
- LwsLCR.jpg 508 × 312; 95 KB
- CuboTax.png 487 × 336; 32 KB
- Swarm.ppt ; 310 KB
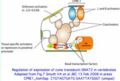
- GNAT2reg.jpg 356 × 241; 18 KB
- 2008-08-archive ; 1 KB
- PaleoPolyploidy.jpg 292 × 723; 45 KB
- RPE65 cycle.jpg 297 × 350; 27 KB
- GaEvol.jpg 111 × 348; 10 KB
- TransducinSumm.jpg 515 × 433; 41 KB
- FullPhylo.jpg 659 × 515; 55 KB
- Algae.jpg 628 × 324; 98 KB
- EnergyWorldsFlyer2009.jpg 2,550 × 3,300; 1.56 MB
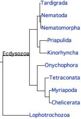
- Ecdysozoa.jpg 188 × 277; 11 KB
- InvertK90.jpg 717 × 864; 410 KB
- DroRH7.jpg 176 × 162; 12 KB
- FossilEyeLobo.pict ; 103 KB
- Ush2aPatho.jpg 854 × 238; 89 KB
- FN3align..jpg 926 × 468; 233 KB
- UsherProtDom.jpg 603 × 343; 46 KB
- WhirlUshVez.jpg 1,132 × 343; 63 KB
- 1718.chr7 maq.bed tail10000.bed ; 423 KB
- 2Hg19uc002ypa.2.pdf ; 484 KB
- 3hg19uc002ypa.2.pdf ; 441 KB
- Hg19uc002ypa.2.pdf ; 1.09 MB
- HgGeneAnnoImage.JPG 779 × 1,008; 244 KB
- TMTgeneTree.png 269 × 552; 9 KB
- CilOpDecline.png 468 × 226; 8 KB
- EncodeKent1.jpg 804 × 172; 72 KB
- Hg18.UCSC GGM BalancedCGIs.bb ; 389 KB
- Hg18.UCSC GGM CombinedEpigeneticScore.bb ; 2.02 MB
- Hg18.UCSC GGM OptimizedScore.bb ; 1.99 MB
- Hg18.UCSC GGM SensitiveCGIs.bb ; 686 KB
- Hg18.UCSC GGM SpecificCGIs.bb ; 178 KB
- Hg18.customText.UCSC GGM BalancedCGIs.txt ; 257 bytes
- Hg18.customText.UCSC GGM SpecificCGIs.txt ; 257 bytes
- Hg18.customText.UCSC GGM SensitiveCGIs.txt ; 259 bytes
- Hg17.UCSC GGM BalancedCGIs.bb ; 389 KB
- Hg17.UCSC GGM CombinedEpigeneticScore.bb ; 2.02 MB
- Hg17.UCSC GGM OptimizedScore.bb ; 1.99 MB
- Hg17.UCSC GGM SensitiveCGIs.bb ; 686 KB
- Hg17.UCSC GGM SpecificCGIs.bb ; 178 KB
- Hg17.customText.UCSC GGM BalancedCGIs.txt ; 257 bytes
- Hg17.customText.UCSC GGM SensitiveCGIs.txt ; 296 bytes
- Hg17.customText.UCSC GGM SpecificCGIs.txt ; 257 bytes
- OUTPUTMACS.bsites peaks.bed ; 1.19 MB
- Ce2.altSplice.bb ; 212 KB
- Ce4.altSplice.bb ; 213 KB
- Ce4Ws140AltSpliceCustomTrack.txt ; 194 bytes
- Ce2Ws120AltSpliceCustomTrack.txt ; 172 bytes
- Ce6.altSplice.bb ; 213 KB
- Ce6Ws190AltSpliceCustomTrack.txt ; 172 bytes
- EncodeHardison1.gif 400 × 260; 24 KB
- Cline.MuseEncode.CSB2010.ppt ; 3.92 MB
- ArskAlign.png 989 × 470; 326 KB
- ENCODE Poster CSB2010.ppt ; 6.39 MB
- GSM566847 1 251652910155.Cy5.txt.gz ; 1.68 MB
- H3K4me3 String50.bed ; 211 KB
- FixStepToBedGraph pl v2.txt ; 2 KB
- Hg19uc002ypa.2.jpg 799 × 519; 295 KB
- BisonSumVar.jpg 1,113 × 373; 63 KB
- Mari signal.bar ; 20.29 MB
- Bisyakcomp.numbers ; 95 KB
- StemtRNAcomp.png 632 × 304; 60 KB
- BisonMitochDisease7Feb.pdf ; 604 KB
- GenbankToTables.txt ; 11 KB
- Jstuart.ppt ; 12.37 MB
- LoadModencode.txt ; 11 KB
- LoadModencode.zip ; 75 KB
- PRDM9onDNA.jp2 0 × 0; 35 KB
- PRDM9setAlign.gif 1,190 × 62; 40 KB
- 29mammalsNovelExons.bb ; 411 KB
- 29mammals2xPARs.bb ; 45 KB
- 29mammals2xHARs.bb ; 45 KB
- 29mammalsExaptedElements.bb ; 3.51 MB
- 29mammalsMotifInstances.bb ; 38.25 MB
- 29mammalsConstraintStructure.bw ; 219 KB
- 29mammalsConstrainedElements omega lods.bw ; 54.74 MB
- 29mammalsConstrainedElements pi lods.bw ; 61.15 MB
- PRDM9refSeqs.pdf ; 52 KB
- 29mammalsPosSelCodonsOverall.bb ; 615 KB
- 29mammalsPosSelCodonsOverall track.txt ; 327 bytes
- 29mammalsPosSelCodonsPval.bb ; 598 KB
- 29mammalsPosSelCodonsSites.bw ; 62.76 MB
- 29mammalsExcessConstraintSCE9.bb ; 507 KB
- 29mammalsExcessConstraintSCE15.bb ; 476 KB
- 29mammalsExcessConstraintSCE30.bb ; 537 KB
- 29mammalsExcessConstraintLambda.bw ; 25.04 MB
- 29mammalsRNAStructUTR.bb ; 161 KB
- 29mammalsRNAStructParalog.bb ; 571 KB
- 29mammalsRNAStructGenome.bb ; 231 KB
- 29mammals2xARs track.txt ; 512 bytes
- 29mammalsConstraintStructure track.txt ; 397 bytes
- 29mammalsExaptedElements track.txt ; 288 bytes
- 29mammalsMotifInstances track.txt ; 283 bytes
- 29mammalsNovelExons track.txt ; 307 bytes
- 29mammalsRNAStruct track.txt ; 920 bytes
- 29mammals track.txt ; 650 bytes
- 29mammals2xARs html.txt ; 470 bytes
- 29mammalsConstraintStructure html.txt ; 185 bytes
- 29mammalsExaptedElements html.txt ; 364 bytes
- 29mammalsExcessConstraint html.txt ; 516 bytes
- 29mammalsMotifInstances html.txt ; 708 bytes
- 29mammalsNovelExons html.txt ; 875 bytes
- 29mammalsRNAStruct html.txt ; 1 KB
- ExtractRepeats.txt ; 2 KB
- PRDMdifAlign.pdf ; 30 KB
- KnucklePhylo.pdf ; 23 KB
- PRDMseq.pdf ; 29 KB
- ZNFvariation.pdf ; 37 KB
- ChimpPRDM9anomaly.gif 375 × 300; 19 KB
- Hg19.geneReviews.bb ; 171 KB
- Hg18.geneReviews.bb ; 166 KB
- GREnhancersMotifEnrichment1.bed ; 21 KB
- GRIR3+Enhancers.bed ; 8 KB
- GRIR3-Enhancers.bed ; 13 KB
- HOMERH3K4meth1DEX-Peaks.bed ; 2.09 MB
- HOMERH3K4meth1DEX+Peaks.bed ; 2.42 MB
- HOMERGRDEX+Peaks.bed ; 180 KB
- HOMERGRDEX-Peaks.bed ; 3 KB
- SQ850Peaks.bed ; 752 KB
- SQ850PeakAnnotation.bed ; 752 KB
- SQ840MotifAnnotation.bed ; 3 KB
- SQ854MotifPeakAnnotation.bed ; 3 KB
- SW854PeakAnnotation.bed ; 487 KB
- SQ854Peaks.bed ; 487 KB
- SQ850MotifPeakAnnotation.bed ; 3 KB
- SQ948MotifPeakAnnotation.bed ; 3 KB
- SQ849Peaks.bed ; 262 KB
- SQ854PeakAnnotation.bed ; 487 KB
- SQ851PeakAnnotation.bed ; 262 KB
- SQ851tagInfo.txt ; 1,002 bytes
- SQ854Info.txt ; 971 bytes
- SQ849tagInfo.txt ; 1,002 bytes
- SQ850tagInfo.txt ; 972 bytes
- XLDB 2011.pptx ; 1.23 MB
- SQ849MotifPeakAnnotation.bed ; 262 KB
- SQ851MotifPeakAnnotation.bed ; 14 KB
- SQ854851850849NewPeakAnnotation.bed ; 1.52 MB
- SQ854851850849NewPeaksIR3-.txt ; 226 KB
- GRIR3-Enhancers.txt ; 79 KB
- SQFOXA1MotifPeakAnnotation.bed ; 15 KB
- SQFOXA1Peaks.bed ; 34 KB
- HOMERFOXA1Peaks.bed ; 113 KB
- SQNewEnhancersIR3+PeakAnnotation.bed ; 441 bytes
- CanidlstEx.jpg 0 × 0; 135 KB
- CanidlstEx2.jp2 0 × 0; 135 KB
- CanidExon.jp2 0 × 0; 91 KB
- WT2.bed ; 7.95 MB
- MetTree.gif 156 × 933; 13 KB
- KPRDM11cons%.gif 468 × 248; 31 KB
- QaReleaseProcess.pptx ; 3.65 MB
- QaReleaseProcess.pdf ; 3.7 MB
- SoftwareEngTesting.pptx ; 1.05 MB
- BME230 Winter 2012.ppt ; 7.31 MB
- DHFR2Comp.gif 1,085 × 333; 182 KB
- DHFRL2Comp.png 1,129 × 344; 317 KB
- DHFRL2Comp.jpg 1,130 × 347; 401 KB
- SeleniumTesting.pptx ; 1.8 MB
- DSC00135.JPG 4,608 × 3,456; 5.58 MB
- CPDragged.gif 618 × 257; 81 KB
- TrimmedGroomedCPD.rtf ; 135 KB
- TrimmedGroomedCPD.doc ; 723 KB
- TrimmedGroomedCRY7.doc ; 304 KB
- UCSC talk.graffle.pdf ; 7.28 MB
- TrimmedgroomedCRY64.rtf ; 88 KB
- TrimmedgroomedCRY64.doc ; 322 KB
- CRY6490%.png 1,101 × 139; 37 KB
- Hg19.ncbiIncidentDb.bb ; 179 KB
- Mm9.ncbiIncidentDb.bb ; 136 KB
- DanRer7.ncbiIncidentDb.bb ; 168 KB
- TrimmedgroomedDASH.doc ; 252 KB
- NansenWarmAir.gif 446 × 367; 3.06 MB
- BigJaxa4NP3big396.gif 396 × 147; 5.69 MB
- BigJaxa4NP3L204F120.gif 396 × 351; 7.94 MB
- Day74ascat3d.gif 415 × 374; 3.62 MB
- UcscGenes.pdf ; 2.97 MB
- UcscGenes.pptx ; 3.19 MB
- HgApi.ppt ; 732 KB
- TrackHub presentation.ppt ; 1.9 MB
- TrackHub presentation.pdf ; 2.33 MB
- BME230 Winter 2014.ppt ; 13.79 MB
- KarenMigaCentromere.pdf ; 43.49 MB
- File-GTEX2015Poster.pdf ; 765 KB
- BME230 Winter 2016.ppt ; 13.8 MB
- BejeranoLab 2016-06-09.pptx ; 2.92 MB
- CIRMposter2016.pdf ; 2.51 MB
- ACMGposter2016.pdf ; 618 KB
- PosterIchg2016.pdf ; 32.84 MB
- GtexAtUcsc Dec2016.pptx ; 2.46 MB
- AssemblyHubsUCSC.pdf ; 3.37 MB
- BME230 Winter 2017.ppt ; 12.98 MB
- AssemblyHubsPAG2018.pdf ; 917 KB
- CyVerseAssemblyHubsPAG2018.pdf ; 1.49 MB
- 2018 PAG Wed 1 17.pdf ; 1.05 MB
- UcscPostDoc 2018.pdf ; 954 KB
- UcscPostDoc 2018.pptx ; 1.51 MB
- UCSC BoG 2018 GtexPoster.pdf ; 732 KB
- UCSC BoG 2018 GtexPoster.pptx ; 749 KB
- OpenStackStartParaHub.sh.txt ; 8 KB
- InteractTrack.pdf ; 776 KB
- NodeReport.sh.txt ; 1 KB
- InitParasol.sh.txt ; 3 KB
- OpenStackParaHubSetup.sh.txt ; 8 KB
- AwsAttachVolume.sh.txt ; 4 KB
- UcscLinks.xls ; 13 KB
- InteractTrackEnhanced.pdf ; 978 KB
- InteractTrackEnhanced.pptx ; 1.8 MB
- Browsermap2018.pdf ; 682 KB
- UCSC PAG 2019 AssemblyHub.pdf ; 668 KB
- GalGal5.grcIncidentDb.bb ; 27 KB
- PAG Hubs.pdf ; 1.07 MB
- UO UCSC.pdf ; 1.55 MB
- UO UCSC CustomTracks.pdf ; 1.07 MB
- UO Hubs.pdf ; 1.59 MB
- NationalYoYoMuseum ChicoCA 50.jpg 2,016 × 1,512; 553 KB
- GalGal6.grcIncidentDb.bb ; 19 KB
- ASHG 2019 Interact.pdf ; 351 KB
- PAG Hubs CyVerse.pdf ; 1.27 MB
- PAG Poster API.pdf ; 1.2 MB
- 2020 PAG UCSC part1.pdf ; 1.45 MB
- 2020 PAG UCSC part2.pdf ; 1.28 MB
- 2020 PAG UCSC part3.pdf ; 666 KB
- UcscLinks.xlsx ; 13 KB
- Browsermap2020.pdf ; 708 KB
- VGP 2021-05-21.pptx ; 1.24 MB
- CartVersion and cartRewrite.c.pdf ; 580 KB
- CSHL 2021.pdf ; 747 KB
- PAG GenArk Poster.pdf ; 1.3 MB
- Hg19.grcIncidentDb.bb ; 106 KB
- Mm10.grcIncidentDb.bb ; 101 KB
- DanRer7.grcIncidentDb.bb ; 111 KB
- Mm9.grcIncidentDb.bb ; 74 KB
- DanRer10.grcIncidentDb.bb ; 126 KB
- DanRer11.grcIncidentDb.bb ; 202 KB
- Hg38.grcIncidentDb.bb ; 282 KB
- Mac background Catalina.png 2,560 × 1,440; 6.2 MB
- BME-110.2023-10-06.pptx ; 9.49 MB
- BME-110.2024-04-25.pptx ; 12.97 MB