TRPV3
Introduction: This article evaluates a particular non-conservative change in the temperature-sensitve loop 2 of the gene TRPV3, the issue being whether the change is adaptive or simply deleterious.
The 200-odd articles on this gene are dwarfed by the some 2,000 on another member of this small gene family, TRPV1. The two genes are adjacent on human chromosome 17, suggesting a recent duplication that TRPV3 should be sistered with it in the gene tree history rather than with TRPV2 as sometimes suggested.
This duplication has not been phylogenetically dated but if the natural signalling ligand of TRPV1 was capsaicin then too, then that also served initially for TRPV3 at the time of duplication (since the encoded proteins had not yet diverged), though it may have later diverged.
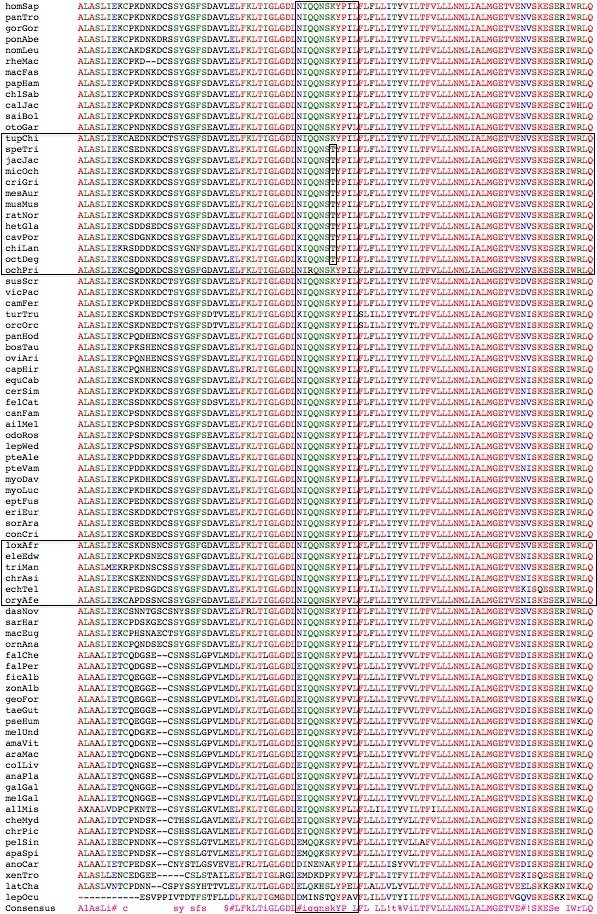
Most TRPV3 research has been conducted in mouse. However as the figure below shows, all rodents but no lagomorphs or primates (or indeed any other tetrapod), have a threonine in place of ancestral lysine two residues over from the D <->N change of interest here. This threonine must represent an adaptive change because mouse diverged from rabbit perhaps 70 mya and the total rodent branch length over which the T was conserved might total 10x that -- too long for an amino acid to persist without supportive selective pressure against, among other possibilites, a return to K.
Meanwhile lagomorphs (and other euchontoglires) continued with K suggesting supportive selective pressure against, among other possibilites, a change to T. This change on the rodent stem is a special case of a rodent synapomorphy, one that is phylogenetically coherent. (The genomic data is however weak -- only pika is available for lagomorphs as the rabbit genome is incomplete in this region.)
Along these same lines, the conserved N in first position in theran loop 2 is not ancestral; rather D/E is seen in monotremes, birds, turtle, lizard, frog, and basal fish. Again, this change is functionally signficant but this signficance is not understood.