USH2A SNPs
USH2A
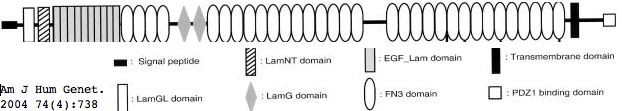
Usherin (USH2A), a 71-exon coding gene located on human chromosome 1q41], encodes a 5202 residue multi-domain protein comprised of a signal peptide, a PDZ1 binding domain (for USH1C and WHRN), 1 laminin NT-terminal domain, 10 laminin EGF-like domains, 4 fibronectin type-III domains (for collagen IV and fibronectin), and 2 laminin G-like domains followed by 31 additional fibronectin type-III domains all tethered to the cytoplasmic exterior by a single transmembrane domain.
The usherin gene is expressed in the basement membrane of many (but not all) cell types, notably in ear interstereocilia ankles and below retinal pigment epithelial cells (Bruch's layer). When normal function is disrupted by mutations in both copies, non-vestibular sensorineural deafness and degeneration of retinal photoreceptor cells called Usher syndrome type IIA results.
Initially, only the first 21 exons were studied but later it emerged that the gene was much longer and mutations along the entire length of the protein all led to the same disease: 125, 163, 230, 268, 303, 334, 346, 352, 478, 536, 595, 644, 713, 759, 1212, 1349, 1486, 1572, 1665, 1757, 2080, 2086, 2106, 2169, 2238, 2265, 2266, 2292, 2562, 2875, 2886, 3088, 3099, 3115, 3124, 3144, 3199, 3411, 3504, 3521, 3590, 3835, 3868, 3893, 4054, 4115, 4232, 4433, 4439, 4487, 4592, 4624, 4795, 5031.
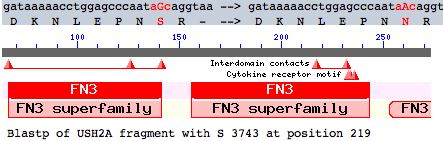
This note evaluates a tentative new SNP in USH2A with comparative genomics. The mutation occurs as a non-hotspot G-->A transition causing a seemingly innoculous S-->N amino acid change at postion 3743. This is just downstream from a glycosylation motif and very near known FN3 interdomain contact residues and a cytokine receptor motif (according to its annotation at SwissProt). This residue lies in the 22nd fibronectin domain which is split across exon 56 and 57.
This change will be shown significant (not plausibly neutral). It could represent an adaptive innovation but is more likely deleterious. The gene is single-copy so there are no prospects for compensation by a second gene. Consequently the mutation, if present on both alleles, could well result in a new form of Usher syndrome type IIA.
Background
Fibronectin FN3 domains are an ancient and exceedingly common domain in bilaterans with 2% of the human proteome containing them (400 genes), often in multiple tandem copies having a role in cell adhesion. However they are not particularly well conserved in primary sequence, though the tertiary structure likely holds up well enough for the structure at 3743 to be determined with both serine and asparagine present.
Here the best blastp match within the human proteome to the FN3 domain containg residue 3743 is to a fibronectin domain of PTPRQ, a dimly related protein tyrosine phosphatase with merely 28% of the fibronectin domains. The best match internally to the other 30 FN3 domains of USH2A is not noticably better, suggesting very substantial divergence since these domains duplicated from a common source.
As can be seen below, the internal fibronectin repeats are most often T at the position corresponding to S3743 though other residues, not including the asparagine of S3743N, also occur. Here the numbering of better matches within the full length protein indicates they do not always correspond in quality to the linear ordering of the FN3 repeat within the protein.
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
FBN.. 3702 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
FBN22 1 WSLPEKPNGLVSQYQLSRNGN-LLFLGGSEEQNFTDKNLEPNS
WS+PEK NG++ +YQ+ + G L+ ++ + T L+P +
FBN.. 3610 WSVPEKSNGVIKEYQIRQVGKGLIHTDTTDRRQHTVTGLQPYT
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLL-FLGGSEEQNFTDKNLEPNS
W PE+ NG++ Y+L RN L F N+TD+ L P S
FBN.. 4285 WIPPEQSNGIIQSYRLQRNEMLYPFSFDPVTFNYTDEELLPFS
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
W P K NG+++ Y + +G L N T +L P +
FBN.. 2553 WQHPRKSNGVITHYNIYLHGRLYLRTPGNVTNCTVMHLHPYT
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
W P PNG + Y+L R+G +++ G E + D L P
FBN.. 4464 WKPPRNPNGQIRSYELRRDGTIVYTG--LETRYRDFTLTPGV
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
W+ P+K NG+++QY L +G L++ G E+N+T +L +
FBN.. 2075 WNPPKKANGIITQYCLYMDGRLIYSG--SEENYTVTDLAVFT
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKN-LEPNS
W P + NG + Y L RNG F G S +F+DK ++P
FBN.. 3521 WRKPIQSNGPIIYYILLRNGIERFRGTS--LSFSDKEGIQPFQ
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
W+ P PNG+V++Y + N L G + +F ++L P +
FBN.. 3040 WTSPSNPNGVVTEYSIYVNNKLYKTGMNVPGSFILRDLSPFT
FBN22 1 WSLPEKPNGLVSQYQLSRN-------GNLLFLGGSEEQNFTDKN--LEPNS
W P PNGLV + + R L+ L S F DK L P +
FBN.. 2644 WQPPTHPNGLVENFTIERRVKGKEEVTTLVTLPRSHSMRFIDKTSALSPWT
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
WS P + NG++ Y + +G L + G + + F + L+P +
FBN.. 4087 WSEPMRTNGVIKTYNIFSDGFLEYSGLNRQ--FLFRRLDPFT
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEE----QNFTDKNLEPNS
WS P+ PN Y L R+G ++ + Q F D +L P +
FBN.. 1074 WSPPDSPNAHWLTYSLLRDGFEIYTTEDQYPYSIQYFLDTDLLPYT
FBN22 1 WSLPEKPNGLVSQYQLSRNG------NLLFLGGSEEQNFTDK--NLEPNS
W PEKPNG++ Y + R ++LF+ F D+ L P +
FBN.. 3887 WMPPEKPNGIIINYFIYRRPAGIEEESVLFVWSEGALEFMDEGDTLRPFT
FBN22 1 WSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNFTDKNLEPNS
WS P NG +++Y L R N L G + +L+P S
FBN.. 4376 WSPPTVQNGKITKY-LVRYDNKESLAG-QGLCLLVSHLQPYS
FBN22 1 WSLPEKPNGLVSQYQL--------SRNGNLLFLGGSEEQNFTDKNLEPNS
W+ P +PNG V Y+L R N + + +F D L P +
FBN.. 4657 WTGPLQPNGKVLYYELYRRQIATQPRKSNPVLIYNGSSTSFIDSELLPFT
FBN22 1 WSLPEKPNGLVSQYQL------SRNGNLLFLGGSEE----QNFTDKNLEPNS
W P + NG + Y L R ++ + + Q++ L+P
FBN.. 4552 WDPPVRTNGDIINYTLFIRELFERETKIIHINTTHNSFGMQSYIVNQLKPFH
FBN22 1 WSLPEKPNGLVSQYQLSRN-------GN--------LLFLGGSEEQN---FTDKNLEPNS 42
WS P PNG + +Y++ R GN ++F + E+N + D L+P +
FBN.. 4175 WSEPVNPNGKIIRYEVIRRCFEGKAWGNQTIQADEKIVFTEYNTERNTFMYNDTGLQPWT 4234
Pseudogene issues
Long isoform USH2A transcripts are over 15,000 bp in length. Consequently position 3743 is not even represented in the set of all human direct transcripts. Even should a retrogene arise from retropositioing, it is unlikely that the process would extent upstream so many exons. Unsurprisingly no processed pseudogenes are evident in any mammalian genome (tblastn of wgs division of GenBank). Thus no potential for confusion exists in locating orthologs of USH2A even in distant species with incomplete genomes.
Paralog issues
No close paralog exists in the human proteome according to the UCSC GeneSorter track. The nearest matches are to other proteins containing laminin or fibronectin domains. No potential for confusion with other genes exists within vertebrates; however comparative genomics at and before teleost fish divergence needs more careful treatment because of whole genome and domain expansion.
Tandem domain repeat issues
In proteins with multiple copies of a given domain, both expansion and contraction can occur over evolutionary timescales resulting in different numbers of repeats in different clades. Under these circumstances it can be difficult to establish orthologs of a given domain. However here the fibronectin domains diverged early on and the 22nd domain seems to be present in all vertebrates with genome projects as a single-copy domain (meaning here no recent duplications or losses).
Known variations
There are no known issues with alternative splicing that would affect the fibronectin domain under consideration here. As noted earlier, a short version of the protein studied initially does not contain residue 3743 at all.
Structural significance
This could be readily evaluated using best-blastp to a structurally determined FN3 domain in PDB, then modelling the FN3 domain in question by submitting it to SwissModel with both S3743 and T3743. Here the percent identity is mediocre but perhaps still sufficient.
>pdb|1X5L|A Related structures Chain A, Solution Structure Of The Second Fn3 Domain Of Eph Receptor
Identities = 27/93 (29%), Positives = 44/93 (47%), Gaps = 16/93 (17%)
Query GVWVTPRHIIINSTTVELYWSLPEKPNGLVSQYQLSRNGNLLFLGGSEEQNF----------TDKNLEPNSRYTYKLEVKTGGG
V V R T+V L W PE+PNG++ +Y++ + E Q++ T L+P +RY +++ +T G
Sbjct QV-VVIRQERAGQTSVSLLWQEPEQPNGIILEYEIK-----YYEKDKEMQSYSTLKAVTTRATVSGLKPGTRYVFQVRARTSAG
Functional significance
Here human individuals homozygous for S3743T could be examined for early loss of hearing accompanied by initial loss of mid-periferal and night vision. (They need not be homozygous because compound mutations would suffice, that is, a different USH2A mutation on the other chr1 allele.
Alternatively, since mouse has an exceedingly conserved orthologous fibronectin domain, the effect of S3743N could be considered as a knockin. Here the symptoms might or might not develop within the 2-3 year lifespan of laboratory mouse.
For the immediate term, comparative genomics is best quide. Here it is clear that S3743 is immensely conserved over several billion years of evolutionary time in all clades observable. This establishes that N3743 is not part of the acceptable reduced alphabet at this residue, though T3743 at one time appears to have been acceptable in the teleost ancestor.
Consequently S3743N -- despite its innocuous appearing nature (ie high Dayhoff matrix score) is likely to have significant non-adaptive impacts on either standalone structure of USH2A protein or its interaction with other proteins in the basement membrane. If selective pressure did not exist to maintain S3743, then what could account for its constancy despite copious variation in nearby residues over the same time spans?
The large number of known loci throughout this protein that give rise to Usher Syndrome 2A suggest that not only does this protein play an exceedingly important structural role highly sensitive to mutation perturbation but also that no other gene product is able to compensate for its absence.
Comparative genomics
USH2A_homSap USH2A_panTro USH2A_gorGor USH2A_ponAbe USH2A_rheMac USH2A_calJac USH2A_micMur USH2A_otoGar USH2A_tupBel USH2A_musMus USH2A_ratNor USH2A_criGri USH2A_dipOrd USH2A_cavPor USH2A_speTri USH2A_oryCun USH2A_ochPri USH2A_vicPac USH2A_susScr USH2A_turTru USH2A_bosTau USH2A_equCab USH2A_felCat USH2A_canFam USH2A_myoLuc USH2A_eriEur USH2A_sorAra USH2A_loxAfr USH2A_proCap USH2A_echTel USH2A_monDom USH2A_macEug USH2A_sarHar1 USH2A_ornAna USH2A_galGal USH2A_taeGut USH2A_anoCar USH2A_xenTro USH2A_tetNig USH2A_takRub USH2A_gasAcu USH2A_oryLat USH2A_danRer USH2A_oncMyk USH2A_pimPro USH2A_calMil USH2A_petMar USH2A_braFlo USH2A_strPur USH2A_helRob USH2A_nemVec