Cryptochrome evolution
See also: Curated reference sequences for cryptochromes and photolyases
Introduction to Cryptochromes
Cryptochromes are large flavoproteins with a curiously complex evolutionary history, beginning billions of years ago as repair enzymes for dna damaged by ultraviolet light. An old gene duplication followed by specializing divergence gave rise to two paralogs repairing distinct types of dna damage (cyclobutane pyrimidine dimers and 6-4 pyrimidine-pyrimidone pairs). These photolyases initially used FAD activated by visible blue light to undo the damage done by UV.
Since FAD has relatively low adsorbance, photolyases evolved a second site for an antenna chromophore with better light harvesting capabilities that could transfer its excitation to the FAD at the active site. This elusive second molecule may be FMN, folate, or a 5-deazariboflavin called Fo (once thought restrict to methanogenic archaea). In the case of the much-studied Drosophila, both the photolyases utilize Fo, making it a new vitamin for this species since the Fo biosynthetic genes are absent.
A second round of gene duplication of the 6-4 photolyase gave rise to a cryptochrome which retained the conformational change induced by FAD binding of blue light but lost dna repair capacity, instead specializing in entraining the day/night circadian rhythm cycle. A third round of gene duplication gave rise to two cryptochromes CRY1 and CRY2.
These five genes were retained in various combinations in different clades during the subsequent course of evolution, causing endless comparative nomenclatural confusion. For example, Drosophila did not retain CRY2 unlike other insects while placental mammals lost all three photolyases though marsupials retained one and monotremes two. Gallinaceous birds also lost a photolyase. Rayfinned fish had a series of further duplications within the gene family. Despite this, the primary sequence, exon structure, fold and FAD, antenna and dna binding sites have largely been conserved -- along with key regulatory binding sites to other proteins -- even as antenna molecules and dna repair capacity was dispensed with.
Standard lab mouse C57BL/6J has a mutated CRY1 cryptochrome gene
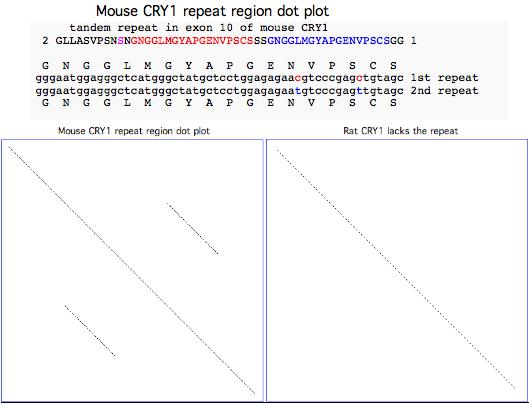
Lab mouse has an odd mutation in its 10th exon where a century of inbreeding may have inadvertently fixed a very serious 54 bp tandem stutter mutation resulting in 18 additional amino acids (the NGGLMGYAPGENVPSCSGG red and blue repeats in NM_007771 reference sequence) that would very likely disrupt the C-terminal region of the protein. The repeat is preceded by the substitution of a serine (shown in magenta in the alignment below) for a strictly invariant proline (back to chondrichthyes).
Although this region lies beyond the two main domains and has a complex evolutionary history, phylogenetic comparison to the eight available rodent and lagomorph sequences implies that this change in lab mouse will have serious functional consequences. A mutation in this critical pacemaker gene could plausibly affect lifespan, metabolic disorder and tumor progression; such a change is completely unprecedented in rodents including rat and indeed in vertebrates.
All 14 available transcripts exhibit the same anomaly -- this is not limited to one strain of mouse, not a somatic mutation, not an unfortunate heterozygous allele. The affected ESTs came from C57BL/6J, C57BL/6, C57BL/6J x DBA/2J, 129 FVB/N and embryo, eye, ventricle, thymus, mammary tumor; the affected GenBank NR entries add a keratinocyte cell line Pam. The mouse genome project used C57BL/6J, the most widely used inbred strain according to the Jackson Laboratory:
"Although C57BL/6J is refractory to many tumors, it is a permissive background for maximal expression of most mutations. C57BL/6J mice are resistant to audiogenic seizures, have a relatively low bone density, and develop age related hearing loss. They are also susceptible to diet-induced obesity, type 2 diabetes, and atherosclerosis. C57BL/6J mice are used in a wide variety of research areas including cardiovascular biology, developmental biology, diabetes and obesity, genetics, immunology, neurobiology, and sensorineural research. C57BL/6J mice are also commonly used in the production of transgenic mice. Overall, C57BL/6 mice breed well, are long-lived, and have a low susceptibility to tumors. Primitive hematopoietic stem cells from C57BL/6J mice show greatly delayed senescence relative to BALB/c and DBA/2J. This is a dominant trait. Other characteristics include: 1) a high susceptibility to diet-induced obesity, type 2 diabetes, and atherosclerosis; 2) a high incidence of microphthalmia and other associated eye abnormalities; 3) resistance to audiogenic seizures; 4) low bone density; 5) hereditary hydrocephalus (early reports indicate 1 - 4 %); 6) hairloss associated with overgrooming, 7) a preference for alcohol and morphine; 8) late-onset hearing loss; and 9) increased incidence of hydrocephalus and malocclusion."
Although this distal region is not modelled in any PDB structure as of March 2012, it has been specifically addressed in 4 of the 195 articles on mouse CRY1 or CRY2.
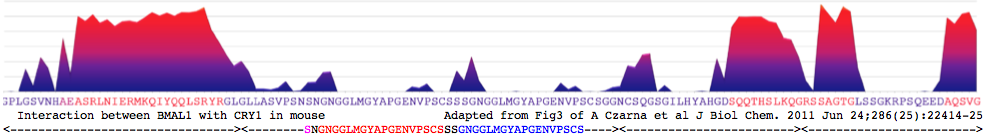
"purified mCRY1/2CCtail proteins form stable heterodimeric complexes with two C-terminal mBMAL1 fragments. The longer mBMAL1 fragment (BMAL490) includes Lys-537, which is rhythmically acetylated by mCLOCK in vivo. mCRY1 (but not mCRY2) has a lower affinity to BMAL490 than to the shorter mBMAL1 fragment (BMAL577) and a K537Q mutant version of BMAL490. Using peptide scan analysis we identify two mBMAL1 binding epitopes within the coiled coil RLNIERMKQIYQQLSRYR and tail regions of mCRY1/2 and document the importance of positively charged mCRY1 residues for mBMAL1 binding."
"mammalian CRY1 and CRY2 are integral components of the circadian oscillator. However, the function of their C terminus remains to be resolved. Here, we show that the C-terminal extension of mCRY1 harbors a nuclear localization signal and a putative coiled-coil domain that drive nuclear localization via two independent mechanisms and shift the equilibrium of shuttling mammalian CRY1 (mCRY1)/mammalian PER2 (mPER2) complexes towards the nucleus. Importantly, deletion of the complete C terminus prevents mCRY1 from repressing CLOCK/BMAL1-mediated transcription, whereas a plant photolyase gains this key clock function upon fusion to the last 100 amino acids of the mCRY1 core and its C terminus. Thus, the acquirement of different (species-specific) C termini during evolution not only functionally separated cryptochromes from photolyase but also caused diversity within the cryptochrome family."
"The mCRY1 and mCRY2 genes are located on chromosome 10C and 2E, respectively, and are expressed in all mouse organs examined. We raised antibodies specific against each gene product using its C-terminal sequence, which differs completely between the genes. Immunofluorescent staining of cultured mouse cells revealed that mCRY1 is localized in mitochondria whereas mCRY2 was found mainly in the nucleus. The subcellular distribution of CRY proteins was confirmed by immunoblot analysis of fractionated mouse liver cell extracts. Using green fluorescent protein fused peptides we showed that the C-terminal region of the mouse CRY2 protein contains a unique nuclear localization signal, which is absent in the CRY1 protein. The N-terminal region of CRY1 was shown to contain the mitochondrial transport signal. Recombinant as well as native CRY1 proteins from mouse and human cells showed a tight binding activity to DNA Sepharose, while CRY2 protein did not"
"genetic screening assay for mutant circadian clock proteins that is based on real-time circadian rhythm monitoring in cultured fibroblasts. By using this assay, we identified a domain in the extreme C terminus of BMAL1 that plays an essential role in the rhythmic control of E-box-mediated circadian transcription. Remarkably, the last 43 aa of BMAL1 are required for transcriptional activation, as well as for association with the circadian transcriptional repressor CRY1"
507 517 527 537 547 557 567 577 587 597
| | | | | | | | | |
CRY1_musMus NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNSNGNGGLMGYAPGENVPSCSSSGNGGLMGYAPGENVPSCSGG NCSQGSGILHYAHGDSQQTHSLKQ GRSSAGTGLSSGKRPSQEEDAQSVGPKVQRQSSN*
CRY1_ratNor NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMGYAPGENVPSGGSGG------------------G NCSQGSGILHYAHGDSQQTNPLKQ GRSSMGTGLSSGKRPSQEEDAQSVGPKVQRQSSN*
CRY1_criGri NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMGYTTGENLPSCSGGG------------------- SCSQGSGILHYAHGDSQQAHLLKQ GRSSMGTSLSSGKRPSQEEETRSVDPKVQRQSSN*
CRY1_spaJud NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMGYTPGENIPNCSSSG------------------- SCSQGSGILHYAHGDSQQAHLLKQ GSSSMGHGLSNGKRPSQEEDTQSIGPKVQRQSTN*
CRY1_dipOrd NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMGYAAGDNLPGSSSSG------------------- SCSQGSGILHYAHGDSQQMHLLKQ GRSSMGTGLSSGKRPSQEEDSQSIGPKVQRQSTN*
CRY1_hetGla NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMGYAPGESIPGSSGSG------------------- SCAHGSGILPCAHTDGQQAHLLKP GRNCVGPVLSSGKRPSQEEDAQSIGPKLQRQSTD*
CRY1_cavPor HHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLLGYAPGESTPGSGGG-------------------- SCVPGSSSAGVSHCAQGEAPQAPP GRDPAGPGLGGGKRPSQEEDAQSTGHKIQRQSPD*
CRY1_speTri NHEASL NIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMAYAPGENIPGCSSSG------------------- SCTQGSSILHNAHGDSQQTHLLKQ GRSSMGTGLSSGKRPSQEEDTQSIGPKVQRQSTN*
CRY1_oryCun NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNPNGNGGLMGYSPGENIPGCSSSG------------------- SCSQGSGILHYAQGDTQQTQLLKQ GRSSMGTGLSSGKRPSQEEDTQSIGPKVQRQSTN*
CRY1_musMus NHAEASRLNIERMKQIYQQLSRYRGL GLLASVPSNSNGNGGLMGYAPGENVPSCSSSGNGGLMGYAPGENVPSCSGG NCSQGSGILHYAHGDSQQTHSLKQ GRSSAGTGLSSGKRPSQEEDAQSVGPKVQRQSSN*
CRY1_ratNor .......................... .........P.................GG.G.------------------. ...................NP... ....M.............................
CRY1_criGri .......................... .........P.........TT...L....GG.------------------- S.................A.L... ....M..S...........ETR..D.........
CRY1_spaJud .......................... .........P.........T....I.N.....------------------- S.................A.L... .S..M.H...N.........T..I........T.
CRY1_dipOrd .......................... .........P..........A.D.L.GS....------------------- S.................M.L.... ...M...............S..I........T.
CRY1_hetGla .......................... .........P.............SI.GS.G..------------------- S.AH.....PC..T.G..A.L..P. .NCV.PV...............I...L....TD
CRY1_cavPor H......................... .........P......L......ST.GSGGG-------------------- S.VP..SSAGVS.CAQGEAPQAPP. .DP..P..GG............T.H.I....PD
CRY1_speTri ......--.................. .........P.......A......I.G.....------------------- S.T...S...N.........L.... ...M...............T..I........T.
CRY1_oryCun .......................... .........P.........S....I.G.....------------------- S...........Q..T...QL.... ...M...............T..I........T.
Coiled coil: RLNIERMKQIYQQLSRYR for CRY1_musMus 480-493
478 R e 0.644
479 L f 0.644
480 N g 0.806
481 I a 0.806
482 E b 0.806
483 R c 0.806
484 M d 0.806
485 K e 0.806
486 Q f 0.806
487 I g 0.806
488 Y a 0.806
489 Q b 0.806
490 Q c 0.806
491 L d 0.806
492 S e 0.806
493 R f 0.806
494 Y d 0.375
495 R e 0.375
Full length CRY1 sequences are available for 10 Glires in the cryptochrome refSeq collection:
CRY1_musMus Mus musculus (mouse) NM_007771 CRY1_ratNor Rattus norvegicus (rat) NM_198750
CRY1_criGri Cricetulus griseus (hamster) XM_003505292 CRY1_spaJud Spalax judaei (blind_mole_rat) AJ606298
CRY1_dipOrd Dipodomys ordii (kangaroo_rat) ABRO01202522 CRY1_hetGla Heterocephalus glaber (blind_mole-rat)
CRY1_cavPor Cavia porcellus (guinea_pig) CRY1_speTri Spermophilus tridecemlineatus (squirrel)
CRY1_oryCun Oryctolagus cuniculus (rabbit) CRY1_ochPri Ochotona princeps (pika)
Lost distal exon in placental cryptochrome CRY1
Although cryptochromes are highly conserved in their two main domains, the C-terminal region in CRY1 has a reputation for variability. This is attributable in part to loss of an ancient exon encoding 32 amino acids in placental mammals. However this exon persists in contemporary marsupials, monotremes, birds, alligators, turtles, lizards, snakes and frogs, so its conservation implies a continuing functional role maintained by selective pressure for several hundred million years of tetrapod evolution.
In addition, some distal motifs in CRY1 are compositionally simple, predisposing not only to the replication slippage event described above for mouse but also to smaller indels in the repetitive regions, notably the 2 aa deletional synapomorphy in placentals in GLLASVPSNPNGN--GGFM (the conserved methionine is at position 514 in human) and possibly the loss of proline (P518) in post-tarsier divergence primates.
The exon loss may have preceded in stages, beginning with alternative splicing that skipped it (this conserves reading frame as the ancestral gene ends with three consecutive phase 12 exons). Later, the exon came not to be used at all and thereafter rapidly degenerated to the point it cannot be detected today by blastx of the relevant region in any placental mammal. The exon does not plausibly contribute to the core fold (photolyase and FAD domains) though it could form a better defined structure upon interacting with other proteins.
The functional consequences of exon loss are unknown; the timing matches that of overall collapse of the photolyase family in placentals. (Note the first half of placental evolution -- about 90 myr -- lacks any living representative, so events can pile up there by coincidence.) Possibly when CYT4, Cyt64, DASH and CPD were lost, the remaining two cryptochromes, especially CRY1, compensated for that loss (without however taking up catalytic roles in dna repair), with exon loss somehow contributing adaptively to that adjustment.
The loss of this exon raises certain questions about the use of marsupial model systems to understand CRY1 functionality in mouse (in turn a model system for human). For example, CRY1 of the marsupial Potorous tridactylus would still retain the exon but to date it has not been placed in a CRY1-- mouse. It would also be feasible to insert just the missing exon into an otherwise intact, ectopically expressed rat CRY1 gene, after first disentangling the effects of the mouse expansion in this same region (shown as ^^ below) as well as proline P518 removal. Note the lab mouse expansion somewhat restores length relative to marsupials, but in the wrong place.
CRY1_homSap MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYS AENIPGCSSSG <-- lost exon in placentals --> SCSQGSGILHYAHGDSQQTHLLKQ GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN CRY1_ponAbe MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYS AENVPGCSSSG SCSQGSGILHYAHGDSQQTHLLKQ GRSSMGTGLSGGKRASQEEDTQSIGPKVQRQSTN CRY1_nomLeu MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYS AENIPGCSSSG SCSQGSGILHYAHGDSQQTHLLKQ GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN CRY1_macMul MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYS TENIPGCSSSG SCSQGSGILHYTHGDSQQTHLLKQ GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN CRY1_calJac MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYS AENIPGCTSSG SCSQGSGILHCAHGDSQQTHLLKQ GRSSMSTGISGGKRPSQEEDTQSIGPKVQRQSTN CRY1_saiBol MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYS AENIPGCTSSG SCSQGSGILHCAHGDSQQTHLLKQ GRSSMSTGLGGGKRPSQEEDTQSIGPKVQRQSTN CRY1_tarSyr MKQIYQQLSRYRGL GLLASVPSNPNGN GGFMGYSPAENTPGCSSSG SCSQGSGILHYAHGDSQQTHLLKQ GRSSVGTGLSGGKRPSQEEDPQSIGPKVQRQSTN CRY1_otoGar MKQIYQQLSRYRGL GLLASVPSNPNGN GSFMEYSPPENIPGCSSSG NCSQGSGILHYAPGDGQQPHLLKQ GRSSMGTGLSGGKRPSQEEDMQSVGPKVQRQSTN CRY1_musMus MKQIYQQLSRYRGL GLLASVPSNSNGN^^GGLMGYAPGENVPSCSSSG NGGLGSGILHYAHGDSQQTHSLKQ GRSSAGTGLSSGKRPSQEEDAQSVGPKVQRQSSN CRY1_ratNor MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYAPGENVPSGGSGG GNCSQGGILHYAHGDSQQTNPLKQ GRSSMGTGLSSGKRPSQEEDAQSVGPKVQRQSSN CRY1_criGri MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYTTGENLPSCSGGG SCSQGSGILHYAHGDSQQAHLLKQ GRSSMGTSLSSGKRPSQEEETRSVDPKVQRQSSN CRY1_spaJud MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYTPGENIPNCSSSG SCSQGSGILHYAHGDSQQAHLLKQ GSSSMGHGLSNGKRPSQEEDTQSIGPKVQRQSTN CRY1_dipOrd MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYAAGDNLPGSSSSG SCSQGSGILHYAHGDSQQMHLLKQ GRSSMGTGLSSGKRPSQEEDSQSIGPKVQRQSTN CRY1_hetGla MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYAPGESIPGSSGSG SCAHGSGILPCAHTDGQQAHLLKP GRNCVGPVLSSGKRPSQEEDAQSIGPKLQRQSTD CRY1_speTri MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMAYAPGENIPGCSSSG SCTQGSSILHNAHGDSQQTHLLKQ GRSSMGTGLSSGKRPSQEEDTQSIGPKVQRQSTN CRY1_oryCun MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG SCSQGSGILHYAQGDTQQTQLLKQ GRSSMGTGLSSGKRPSQEEDTQSIGPKVQRQSTN CRY1_oviAri MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSA SCTQGSGILHYAHGDSQQTHLLKQ GRSSTAAGLGSGKRPSQEEDTQSVGPKVQRQSTN CRY1_bosTau MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSNA SCTQGSGILHYAHGDSQQTHLLKQ GRSSTGAGLGSGKRPSQEEDTQSIGPKVQRQSTN CRY1_susScr MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG SCPQGSGILHYAHGESQQNHLLKQ GRSSTGSGLSSAKRPSQEEDTQSIIGPKVQRQSTN CRY1_ailMel MKQIYQQLSRYRGL GLLASVPANPNGN GGLMGYSPGENIPGCSSSG SCSQGSGILHYAHGDSQQTHLLKQ GRSSMGSGLSSGKRPSEEEDTQSIGPKVQRQSTN CRY1_turTru MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGYSSSG SCTPGSGILHYAYGDSQQTHLLKQ GRSSTCTGLSSGKRPSQEEDTQSIGPKVQRQSTN CRY1_equCab MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG SCSQGSGILHYAHGDSQQTHLLKQ GRSSLGPGLSSGKRPGPEEDTQGIGPKVQRQSTT CRY1_canFam MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG SCSQGSGILHYAHGDSQQTHLLKQ GRSSMGTGLSSGKRPSEEEDTQTISPKVQRQSTN CRY1_myoLuc MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG SYAQGSGILHYALGDSQQTHLLKQ GRSSVGTGLSSGKRPSQEEDTQSIGRKVQRQSTN CRY1_pteVam MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG SCSQGSGSLHYAHGDCQQTHLLKQ GRSSMGTGLSSGKRPSQEEDMQSIGPKVQRQSTN CRY1_loxAfr MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENTPGCNSSG SCSQGSGILHYVHGDS....LLKQ GRSPTGTGVSSGKRPSQDEETQTLGPKVQRQSTN CRY1_triMan MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSNG SCPQGNGILHYAHRDSQQAHLLKQ GRSPTGTGVSSGKRPSQEEETQSIGPKVQRQSAN CRY1_proCap MKQIYQQLSRYRGL GLLASVPSNPNGN GGLIGYSPGESIPGCSNSG SCSQGSGILHYAHGDSQQAHLLKP GRSPMGTGISSGKRPSQEEETQTVGRKVQRQSTN CRY1_echTel MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENTTGCSSGG GCPPGNGILHYAHGDSQQAALLKQ GRSPLGTGLSSGKRPSQEEDTQSVGPKVQRQSSN CRY1_dasNov MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYAPGENILGCSSSG SCAQGSSILHYAHGDNQQTHLLKQ GRSSMGTVLSSGKRPSQEEETQSIGPKVQRQSTN CRY1_choHof MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYSPGENIPGCSSSG sCSQGSGILHYAHGDSQQTHLLKQ GRSSMGIGLSSGKRPSQEEETQGIGPKVQRQSTN CRY1_monDom MKQIYQQLSRYRGL GLLASVPSNPNGN GSLMAYTPGENIPGCSSGG GAPVGASDGQIL..QACVLPEPPTGTSGVQQP GYSQGSGISHYSHEDSQQAYMLKQ GRSSL..GVGGGKRPRQEEETQSINPKVQRQSTN CRY1_macEug MKQIYQQLSRYRGL GLLASVPSNPNGN GSLMGYTTGENIPTCSSSGG GAPAGASDGQIL..QACVLPEPPTGTSGVQQP GGYSQGGISHYSHEDSQQAYVLKQ GRNSL....GGGKRHRQEEETQSIGSKMQRQSVN CRY1_sarHar MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYTSGENGPACNSGG GAPVGASDGQIL..QSCALPEPPAGASCIQQS GYSQGSGISHYSHEDSQQAYILKQ GRSSL....SGGKRPRQEEETQSVGPKVQRQSVN CRY1_triVul MKQIYQQLSRYRGL GLLASVPSNPNGN GGLMGYAPGENIPACSSSGG GAPAGVGDGQIL..QACALPEPPTGASGVQQP GYSQGSGISHYAHEDSQQAYMLKQ GRSSL...SGGGKRHRQEEEAQSIGPKMQRQSVN CRY1_ornAna MKQIYQQLSRYRGL GLLASVPSNPNANGSGGLMAYSPGENIPGCSSGGG GVQMGASESHLL..QTCVLGESHLGPSGIQQQ GYCQGSGVLYYANGE....SHLTQ GRSSLTPGLSGGKRPCQEEESQSIGPKVQRQSTD CRY1_tacAcu MKQIYQQLSRYRGL GLLASVPSNPNANGSGGLMAYSPGENIPGCSSGG GAQIGASESHLL..QTCVLGESHLGPSGIQQQ GRSSLTPGLSGGKRHCQEEESQSIGPKVQRQSTD CRY1_galGal MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMSFSPGESISGCSSAG GAQLGTGDGQTVGVQTCALADSHTGGSGVQQQ GYCQASSILRYAHGDNQQSHLMQP GRASLGTGISAGKRPNPEEETQSVGPKVQRQSTN CRY1_melGal MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMSFSPGESISGCSSAG GAQLGTGDGQTVGVQSCALGDSHTGGNGVQQQ GYCQASSILRYAHGDNQQPHLMQP GRASLGTGISAGKRPNPEEETQSVGPKVQRQSTN CRY1_eriRub MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG GAQLGTGDGHTV.VQSCTLGDSHSGTSGIQQQ GYCQASSILHYAHGDNQQSHLLQA GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN CRY1_sylBor MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG GAQLGAGDGHSV.VQSCALGDSHTGTSGVQQQ GYCQASSILHYAHGDNQQSHLLQA GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN CRY1_taeGut MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG GAQLGTGDGHSV.VQSCALGDSHTGTSGIQQQ GYCQASSILHYAHGDNQQSHLLQA GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN CRY1_parWeb MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG GAQLGTGDGHSV.VQSCALGDSHTGTSGIQQQ GYCQASSILHYAHGDNQQSHLLQA GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN CRY1_allMis MKQIYQQLSRYRGL GLLATVPSNPNGNGNGGLMGYSPGENVSGCGSTG GAQMGSSDGHTVSVQPCALGESHGGSNGIQQQ GYFQASSILHFPHGDDQQSHLLQQ GRTSLSSGISAGKRPNPEEETQSIGPKVQRQSTN CRY1_anoCar MKQMYQQLSRYRGL GLLASVPSNGNGNGNGGLMGYSTGENIPGCTNTN GSQMGMNEGHIGNVQACTMGESHTGTSGIQQQ GYSQGSGILLYSHGDNQKTHSAQK GRISLGTGVCTGKRPSPEVETQSVGPKVQRQSSN CRY1_podSic MKQIYQQLSRYRGL GLLASVPLNGNGNGNGGLMGYSTGENIPGCTNTN GSQMGTNEAHTGSVQTCTLGESHTGTSGIQQQ GYPQGSDILHYAHGEGQKTHLIQQ GRASLVAGVCTGKRPNPEEETQSIGPKVQRQSSK CRY1_pytMol MKQIYQQLSRYRGL GAQMGTSEGHTGNVQACTLGETHTGTSGIQQQ GYSQGNSGILHYAHGDSQKTLLMQ GRTSLSVGVCTGKRPNPEEGIQSIGPKVQRQSSN CRY1_chrPic MKQIYQQLSRYRGL GLLATVPSNPNG..NGGLMGYSPGENISGCSSAS GAQMGSNDGHTVGVQTCSLEDSHAGSSGIQQH GYSQGNSIVHYAQGDHQQSHLLQQG GRTVST GISTGKRPNPEKETQSIGPKVQRQSTN CRY1_xenTro MKQIYQQLSRYRGL GLLASVPSNPNGNGNGGLMSYSPGESMSGCSNNG GGQMGVNEGSSASNPNANKGEVHPGTSGLQ.. GYWQGSSILHYSHSDSQQSY LMQ ARNPLHSVVSSGKRPNPEEETQSIGPKVQRQSSH CRY1_xenLae MKQIYQQLSRYRGL GLLASVPSNPNG..NGGLMSYSPGESMPGCSNNG GGQMGAIEGSSASNPNPNQGEVLPGTSGLQ.. GYWQGSSILHYSHSDNQQSY LMQ ARNPLHSVVSSGKRPNPEEETQSVGPKVQRQSTH CRY1_latCha MKQIYQQLSRYRGM GLLASVPSNPNGNGGLGCSLAENIPVCNSAA GAQMGGDDGHKVSVLAYTQGDSRAGEIEMQQQ CRY1_danRer MKQIYQQLSCYRGL GLLAMVPSNPNGNGENSTSLMGFQTGDMTKEVTTPS GYQMPPTSQGEWHGRTMVYSQGDQQTSSIMTSQ GFGNNGSTMCYRQDAQQIT GRGLHSSIIQTSGKRHSEESGPTTVSKVQRQCSS
When the terminal four exons of CRY1 are compared to those of its nearest homolog class CRY2, no similarity can be detected beyond the first 8 residues of the tenth exon of CRY1 (2 GLLASVPS) vs the tenth and penultimate exon of CRY2 (2 CLLASVPS). This raises the question of what the last common ancestor had for terminal exons and -- given no counterpart in CRY4, CRY64, DASH, or CPD -- where they originated. Note that last two exons of CRY2 are strongly conserved in their own right, proving a separate conserved functionality from that of CRY1. Since the tenth exons begin homologously and end after a similar length with a phase 1 splice donor, these exons could possibly be homologous their entire length, just diverged distally. The eleventh exon of CRY2 could then correspond (allowing for total sequence divergence) to any of exons 11-13 in CRY1.
CRY2_homSap CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA CRY2_panTro ............................... .............................................. CRY2_gorGor ...........................V... .............................................. CRY2_ponAbe ...........................V... .............................................. CRY2_rheMac ...........................VN.. ...............................K.............. CRY2_papHam ...........................VN.. ...............................K.............. CRY2_calJac ............................... .............................................V CRY2_micMur ..............................T .................................T............ CRY2_musMus ....................G......I.NT ...A.S.....................T.....T.M..Q.PA...S CRY2_ratNor ....................G......I.NT .....S...........................T.M.AQ.P....S CRY2_criGri ...........................I.NT .S...S...........................T.M.AQ.PQT... CRY2_spaJud ........................P..ITNT .....ST..........................T...A..PA.... CRY2_cavPor .....................L.....ST.T ......G.................................P..... CRY2_hetGla ....................TL.....S..T ...S..D..............................A..PT.... CRY2_speTri ....................G......I..T .....S..Q..................................... CRY2_oryCun ...........................V.G. A..................................V........AV CRY2_turTru .........M....N...........G.... ................G.................G..PS..L...V CRY2_bosTau ..............N.......I....S..V ......G.................G..........SLPS....RGV CRY2_susScr ..............N............V.A. .....................................PT...GR.V CRY2_canFam ..............N.........T...... ..........................................CR.V CRY2_ailMel ..............N.........T...... .....................................A..P..R.V CRY2_myoLuc .........M....N......L..T...... ..K..................................AT....R.V CRY2_pteVam .............NN.........T...NN. .....................................A.....R.V CRY2_loxAfr ..............S............SN...........T........................K..G.......V CRY2_proCap ..............N........P..H.....L................................K..G.....T.. CRY2_choHof ..............N....................V............................T...........V CRY2_macEug .........M....S.M..T.MG....V..T..K...CS..........T..ASR..H.....M.A..V...A.--- CRY2_monDom .........L....S.MV.A.LG...AV.GP.LK...CS..........T..A....H.......R..GS..AG..V CRY2_ornAna ..............SAA..SGLG....NI.TA...-.P.............GL.....C..PK..GR.G..P.GE.. CRY2_galGal ..............G..TDSAPG.-..ST.TAV.LPQ.DQ......H.G...LCT...Y...K.TG..A..I.G.SS CRY2_taeGut ............I.G..PDSA.G.-.CST.TAV.LSQAEQ......H.G....CS...Y...K.TG...S.ISG.SL CRY2_allMis G........A....G..TD.A.V.-.CST.TALK.SQ..Q......H.GI..MCT.D.Y...K.TG.HG..I...SL CRY2_anoCar .........M....N...DT...H-.NCIGTAS.QTHC.QT.....HDVVQ.YK-...Y...K.VASQFA.N.RQEL CRY2_xenTro .I.......M...GG.M.DS.QNISEAGKM.P.SHTSGESVLAAQYTAGI--------------------------- CRY2_ranCat .I......S.....G.M.D.A...Q..SD---.A.RLCAVD.....H.DLD----G..C.K..LQCVQEM.RAA..F
A distal alternative splice in avian cryptochrome CRY1 not used for magnetosensing
Bird CRY1 presents a further curious situation with respect to the terminal extentional exons of CRY1: an alternative splice in exon 11, more accurately a failure to recognize its splice donor with subsequent read-out to the first stop codon encountered. The vast majority of such events are misinterpreted artifacts -- the transcript terminated too soon, providing no splice acceptor and so no way for the intervening intron to be removed.
However here two types of transcripts were found in both Erithacus rubecula (Euro robin) and Sylvia borin (warbler) in targeted experiments by separate research groups. The long form, called there CRY1A, has the usual four terminal exons of vertebrates; the short form, CRY1B, provides 25 new amino acids before a stop codon.
Comparative genomics is capable of resolving artifact, coincidence, and functional. First note that GenBank chicken transcripts has a supportive entry (BU143111) that surfaced in a large transcript program not focused on particular genes. Secondly, the read-out of exon 11 in species without transcripts is highly conserved in amino acid sequence. While a certain amount of nucleotide conservation might be expected because splice sites are larger than just GT-AG and the intron may contain enhancers or conserved non-coding, and conservation can persist for a time just because of coldspots and mutational inertia, the conservation here at the protein level significantly exceeds what these factors could contribute. Gray shows species lacking conservation; blue conserved amino acids within birds.
Exon 11 read-out of CRY1 genSpp transcript support of read-out
GISKNTF* monDom Monodelphis domestica (opossum)
GISDNTFLTLTQSRGSLGIPHQS..* macEug Macropus eugenii (wallaby)
GISQNTFESVRLS* sarHar Sarcophilus harrisii (tasmanian_devil)
GISKLFSFIFKNTFN* ornAna Ornithorhynchus anatinus (platypus)
GRSSLTPGLSGGKRHCQEEESQN..* tacAcu Tachyglossus aculeatus (echidna)
GIMAVPVCRGSPNPCNYRKPDKTSK* taeGut Taeniopygia guttata (finch)
GIMAVPVCRGSPNACNYGKPDKTSK* eriRub Erithacus rubecula (robin) AY585717
GIVAVAVCRGSPNPCNYGKPDKTSE* sylBor Sylvia borin (warbler) DQ838738
GIMAVPVCRGSSNPCNCGKTDKTSK* melUnd Melopsittacus undulatus (parakeet)
GMTGVLVCRGSPGSHNYGKKDKT* anaPla Anas platyrhynchos (duck)
GIVGVPICRGSADLCN* galGal Gallus gallus (chicken) BU143111
GTVGVPICRGSANWYK* melGal Meleagris gallopavo (turkey)
GIIQQVKCLQRICKFL* allMis Alligator mississippiensis (alligator)
GAQMGSNDGHTVGVQTCSLEDSH..* chrPic Chrysemys picta (turtle)
anoCar Anolis carolinensis (lizard)
GKLAAPLISVSSIIGVFHTHEPQ..* xenTro Xenopus tropicalis (frog)
The data thus support the notion of birds having evolved a distinct function for the read-out option at exon 11 -- with nothing comparable in other reptiles (including the immediate outgroup crocodilleans) or mammals. Many more bird genomes are expected in 2012 (notably Corvus, Ficedula, Geospiza, Manacus and Paradoxornis) that can confirm if read-out conservation patterns conform to the avian phylogenetic tree. However the more common CRY1 form retaining the usual extra exons is also conserved in birds (as seen in the earlier alignment of this region).
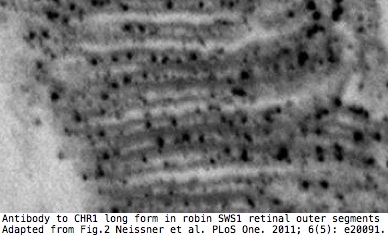
Here it has been reported that the long form only is expressed in SWS1 opsin cones of retinas of migrating birds where it detects the earth's magnetic field via electron spin pairing in tryptophan and FAD. The short form is apparently expressed in the ganglion cell layer.
Human CRY2, also strongly expressed in retina but not so specifically in cone cell outer segment membranes, can reportedly replace the invertebrate cryptochrome CRY1B in the drosophila magnetic field detection system (as can insect CRY1A). The final exon of human CRY2 bears no clear relationship to the terminal exons of CRY1 nor to the read-out exon 13 of birds and is only secondarily related homologically to invertebrate CRY1B cryptochromes.
Note the vertebrate ciliary opsin SWS1 has no counterpart in fruit flies which do however have two rhabdomeric opsins with peak sensitivity in the ultraviolet, RH5 and RH7, with characteristic lysine at position 90 and a shorter third cytoplasmic loop. RH5 is located in the larval Bolwig organ; RH7 has not been assigned an anatomical site but may be located in antenna.
Vertebrate CRY1 reference sequences
Here it is quite important to straighten out misnamed homologs, especially in zebrafish which has been studied extensively. Because chondrichthyes, lobe-finned fish, and basal ray-finned fish (gars) already have two separate genes classifying as CRY1 (ie both distinct from CRY2 and other photolyases), a gene duplication must occurred in early vertebrates and persisted almost to land animals.
Both syntentic sites can be explored in Xenopus and amniotes but only one location hosts a cryptochrome today. These two syntenic regions are not more broadly paralogous (not supportive of a large scale duplication). Zebrafish, which has four distinct CRY1 paralogs in addition to a CRY2, may have had a doubling of this ancestral pair through whole genome duplication but the sequences don't quite cluster in this way. All four CRY1 genes are actively transcribed.
The figure (taken from the Genomicus synteny tool) shows that CRY1 experienced a small local inversion in amniotes subsequent to mammalian divergence. This may have carried all upstream regulatory regions along with it or left some portion orphaned in a new downstream position with unknown functional consequences. Since the event occurred some 300 myr ago, the boundaries of the inversion cannot be precisely determined.
For a full set of 56 deuterostome CRY1 sequences, see the curated refSeqs collection.
>CRY1_homSap Homo sapiens (human) 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENIPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_xenTro Xenopus tropicalis (frog) NM_001087660 11533577 final four exons confirmed by many ESTs 0 MGVNAVHWFRKGLRLHDNPALRECIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWKITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMDPLEIPVETITAEVMEKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASTTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMDGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGKRTDPNGDYIR 2 1 RYLPILKGFPPKYIYDPWNAPETVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGNGGLMSYSPGESMSGCSNNG 1 2 GGQMGVNEGSSASNPNANKGEVHPGTSGLQ 1 2 GYWQGSSILHYSHSDSQQSYLMQ 1 2 ARNPLHSVVSSGKRPNPEEETQSIGPKVQRQSSH* 0
Vertebrate CRY2 reference sequences
These are pre-curated provisional sequences, taken from the UCSC 46-way genomic alignment relative to human except where confirmed by an accession number. This can in omission of insertions in other species and missing exons in the situation where they are too diverged or lie in isolated small contigs. The final exons, being quite variable especially in fish, are best determined from transcripts when available and then extended by blastx homology to species within the same clade lacking transcripts. After these corrections, the sequences are aligned and further anomalies are confirmed or discarded on a case-by-base basis.
It appears that the last exon in fish has lost all homology (and so functionality), in some cases simply running out into junk dna until a stop codon is encountered. Exon seven is broken up in some fish with an extra intron that might have some use in fish taxonomy as a derived characteristic.
No evidence for CRY2 currently exists in cartilaginous fish or earlier deuterostomes suggesting that ancestral CRY1 duplicated in stem bony vertebrates, giving rise to CRY2. It appears that all insect CRY2 entries at GenBank are grievously mislabelled and actually represent a CRY1 parent gene of a duplication of CRY1 in insects whose duplication was lost in drosophilids. If so, these 'CRY2' sequences cannot serve as valid model systems for vertebrate CRY2 and indeed are poorly suited as CRY1 proxies because their properties may have changed in species retaining the second copy. The drosophila cryptochrome -- being an esoteric retained sole copy -- is even more unsuitable for annotation transfer to vertebrates.
For a full set of 43 vertebrate CRY2 sequences, see: Curated reference sequences for cryptochromes and photolyases
>CRY2_homSap Homo sapiens (human) 11 exons 0 MAATVATAAAVAPAPAPGTDSASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGLVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 >CRY2_musMus Mus musculus (mouse) CF898022 0 MAAAAVVAATVPAQSMGADGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQALISRMELPKKPAVAVSSQQMESCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPGSSQAGSISNT 1 2 GPRALSSGPASPKRKLEAAEEPPGEELTKRARVTEMPTQEPASKDS* 0
Invertebrate cryptochromes
Nomenclature here is an immense source of confusion, but with the number of genomes available today, it is clear that early bilateran ancestor contained two distinct cryptochromes (in addition to three photolyases), all of which persisted into early deuterostomes, notably sea urchin. These are denoted CRY1A and CRY1B here to distinguish them from a later gene duplication of CRY1A in vertebrates yielding CRY1 and CRY2 (human gene nomenclature must be used by international agreement, excluding their use in invertebrates).
Some lophotrochozoa, notably molluscs, retained both genes. Within arthropods, generally one cryptochrome was retained but not always the same one. However some dipterans, hemipterans and lepidopterans retain both. It appears that the CRY1A/CRY1B gene duplication itself took place after divergence from cnidarians.
The two cryptochromes are intronated quite characteristically, with the first class (called CRY1A here) most similiar to that of vertebrate CRY1/2, in agreement with closer blastp clustering. The second class (called CRY1B below) bears less relevance to the cryptochromes retained in mammals but unfortunately is the one retained in Drosophila and most studied. Annotation transfer from study of CRY1B proteins to mammals is thus exceedingly problematic given that the CRY1A family retains far more sequence similarity and is not descended from CRY1B.
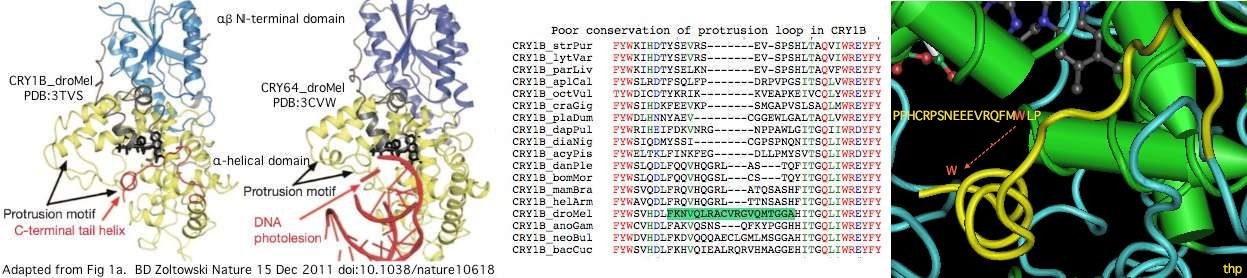
A remarkable recent crystallographic result establishes an important role for tryptophan W536 near the end of the variable region of drosophila CRY1B. This aromatic residue and its associated helix arch back to occupy the site normally occupied by a damaged dna nucleotide, spoofing the presence of a damaged dna residue for conformational change purposes.
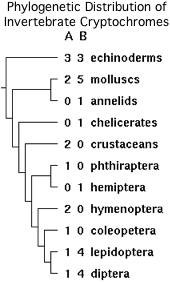
The tryptophan is part of a larger motif PPHCRPSNEEEVRQFMWLP conserved in the CRY1B orthologs from insects, crustaceans, molluscs and surprisingly three echinoderms. It is erroneously denoted the FFW motif in the fruit fly cryptochrome literature. Note the full motif is quite well conserved in amino acid sequence whereas the protrusion motif is not conserved either in residue or length.
However two substitutions should be noted, cysteine and tyrosine in daphnia and aphid respectively, suggesting that the overall motif is more critical than just a tryptophan. Further, no comparable residue or motif exists in invertebrate CRY1A proteins, vertebrate cryptochromes or other photolyase homologs. The observed phylogenetic distribution (which is unlikely to reflect convergent evolution) implies the spoofing mechanism arose in an early bilateran after to the gene duplication giving rise to CRY1A/CRY1B but before the protostome/deuterostome split.
A threonine at position 518 is reported phosphorylated in CRY1B_droMel but has no real phylogenetic support even within drosophilids, also lying outside the motif detected by WebLogo. This post-translational modification could nonetheless have regulatory signficance in the limited spectrum of species that could have it but more likely it represents an aberrant event.
For a full set of 38 invertebrate CRY1A and CRY1B sequences, see: Curated reference sequences for cryptochromes and photolyases
PPHCRPSNEEEVRQFMWLP
CRY1B_strPur KVVNKLRDTGIVHCAPSTQREVREFVWLPEKMAGGGSCRADQNCEGILGL echinoderm
CRY1B_lytVar KVINRLRDSGIVHCAPSTQKEVREFVWLPEKMAGGGSCRADQNCEGILGL echinoderm
CRY1B_parLiv KVINRLRDSGIVHCAPSTQKEVREFVWLPEKMAGGGSCRASQNCEGRTGS echinoderm
CRY1B_aplCal MEAIKKVSKDVPHIAPANEEEVLTLMWSGKQTRSELMDA----------- mollusc
CRY1B_craGig AVKDALIGKEIPHCAPSEEIEARRFSWLP--------------------- mollusc
CRY1B_octVul KVKEHLLHQDVPHCGPTNETEVWKFAWLPPIEHHDLAHNI---------- mollusc
CRY1B_rudPhi KNKLVQQGKDLEHCRPTNVEEVRMFVWMPGAHKGACGQEVPLDDKELCDG mollusc
CRY1B_plaOce MDRIKNLCKGIPHVAPTNENEVLSYMWLDKSNSEAMEESLFEACSHLSSV mollusc
CRY1B_dapPul KEFRQKFKETPAHCQPSSNSEVYKFFCLPDDSLPF--------------- crustacean
CRY1B_diaNig DEIRNRLMNPPPHCRPSSEKETRQFMWFPDDCSEHSSQ------------ orthoptera
CRY1B_acyPis LRVSMTNENRVPHCCPSDREEVQKFMYLPDECMQQLLPLENQDSKAYDIY hemiptera
CRY1B_danPle QELRRLLEKAPPHCCPSSEDEVRQFMWLGDDSQPELTTT----------- lepidoptera
CRY1B_bomMor EELRMLLEKAPPHCCPSSEDEIRQFMWLNE-------------------- lepidoptera
CRY1B_mamBra GELRHFLQKAPPHCCPSSEDEIRQFMWLNE-------------------- lepidoptera
CRY1B_helArm KELRHMLQKAPPHCCPSSEDEIRQFMWLNE-------------------- lepidoptera
CRY1B_droMel KSLRNSLITPPPHCRPSNEEEVRQFFWLADVVV----------------- diptera
CRY1B_anoGam REKLVDGGSTPPHCRPSDIEEIRQFFWLADDAATEA-------------- diptera
CRY1B_neoBul LIAEGAPDNGPPHCRPSNEEEIRNFFWLAD-------------------- diptera
CRY1B_bacCuc LIAGGAPDEGPPHCRPSNEEEVHQFFWLVE-------------------- diptera
CRY1B: C-terminal conservation in drosophilids * *
Drosophila melanogaster YECLIGVHYPERIIDLSMAVKRNMLAMKSLRNSLI T PPPHCRPSNEEEVRQFFWLAD
Drosophila simulans ................................... . .....................
Drosophila sechellia ................................... . .....................
Drosophila yakuba ............................T...... . .....................
Drosophila erecta ................T...Q.......A...... . .....................
Drosophila rhopaloa ............................A.....M . .....................
Drosophila elegans ............................A.....M . .....................
Drosophila takahashii ..................Y.........A.....M . .....................
Drosophila ficusphila .................L..........A...... . .....................
Drosophila eugracilis ............M....L..........A...... . .....................
Drosophila biarmipes .................VY.....M...A...... . .....................
Drosophila kikkawai .................K.........TA...... . .....................
Drosophila mojavensis ..........D......L.S........A...... E .....................
Drosophila persimilis ................K......M..TA...... . ....................N
Drosophila pseudoobscur .................K......M..TA...... . ....................N
Drosophila bipectinata ..........D.L...TK...G.........D... . ..............T......
Drosophila ananassae ..........D.L....K...G......T..D... . ..............T......
Drosophila willistoni ...........P.....L.L...T...TN...... . ....................E
Drosophila grimshawi ................L.S....A...A......E . T.................DE.
Drosophila virilis ....L.F...Q......L.S...TM...A...... E ...................TN
>CRY1B_strPur Strongylocentrotus purpuratus (sea_urchin) XM_001183029 echinoderm lacks final 2 exons 0 MPGGACIHWFRHGLRLHDNPALLEGMTLGKEFYPVFIFDNEVA 1 2 GTKTSGYNRWRFLHDCLVDLDEQLKAAGGRLFVFHGDPCLIFKEMFL 0 0 EWGVRYLTFESDPEPIWTERDRRVKALCKEMKVECIERVSHTLWNPDI 2 1 IIEKNGGTPPITYSMFMECVTEIGHPPRPMPDPILTKVNMKIPSDFEERCALPSLEVM 1 2 GVNMECTEQEKKVWKGGETRALELFRVRILHEEE 0 0 AFKGGYCLPNQYMPDLLGTPKSLSAYLRFGCLSVRRFYWKIHDTYSEVRS 0 0 EVSPSHLTAQVIWREYFYTMSVGNIHFNKMKENPICLNIEWKEDDEKLKAWTD 0 0 GRTGYPWIDACMKQLKYEGWIHQVGRHATACFLTRGDLWISWEDGLQ 0 0 VFDKYLLDADWSICAGNWMWISSSAFEKFLQCPNCFCPVRYGRRMDPTGEYVR 2 1 RYLPVLKDMPIRYLFEPWKAPRAVQERAKCIVGKDYPMPVVEHKSASAANHEQMEKVVNKLRDT 1 2 GIVHCAPSTQREVREFVWLPEKMAGGGSCRADQNCEGILGL* 0 >CRY1B_octVul Octopus vulgaris (octopus) JR450373 transcript assembly mollusc 0 MKLEKKQKIAVHWFRHGQRLHDNPALLDALKDCDEFYPVFIFDGEVA 1 2 GTKLCGFNRWRFLLENLKDLDESFSEYGGRLYTFQGKPVEVFANLQN 0 0 EWGITHITAEIDPEPIWQERDDAVKEFCQKSGIKCDFFNSHTLWDPKR 2 1 LLKKNGGTPPLTFELFQLVTSSLGPPPRPIDAPTFEGIKMPLPENHDKFSVPTLKSL 1 2 GIYPEFEEQKNPINVFIG GEKRALVLLKARLEKEAQ 0 0 SFRHGQCLPNHQEQPELLARAVSLSPYLRFGCVSIRKTYWDICDTYKR 0 0 IKKVEAPKEIVCQLYWREYFYIMSIDNINFDKIENNPYCLKINWQYNEEFLKKWEM 0 0 GQTGYPWIDAIMNQLRFEGWNHHVGRHAVSCFLTRGDLWVSWEDGLK 0 0 LFLKYQLDADWSVCAGNWMWVSSSAFEKALQCPTCYSPVMYGMRMDKNGDFVKTYVPVLKDMPL 2 1 KYLFCPWKAPLEIQEKANCIIGKDYPEPIVMHRDASKQNMAKMYKVKEHLLHQ 1 2 DVPHCGPTNETEVWKFAWLPPIEHHDLAHNI* 0 >CRY1A_dapPul Daphnia pulex (water_flea) FE418063 FE356487 ACJG01001137 crustacean 0 MYRQQKNIMSGYDSEPREKQVVHWFRKGLRLHDNPSLKDGLKGCSTYRCIFILDPWFAGSSNVDINKWR 2 1 FLLESLEDLDQNLRKLNSRLFVIRGQPAGVLPKLFK 0 0 EWETTCLTFEEDPEPFGRVRDQNIITMCKDFNIEVITRASHTLYHPQK 2 1 IIEKNGGKAPLTYRQFQNIIASVDAPPPPESDITFESIGRGYTPMDESMDDRFSVPTLEEL 1 2 GFDTDGLMPAVWHGGETEALTRLERHLERK 0 0 AWVASFGRPKMTPQSLLASQTGLSPYLRFGCLSVRLFHQQLTNLYKKIKKAQPPLSLHGQVLWREFFYCAATNNPNFDKMIGNPICVQIPWDSNAEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGTWMWLSCSSFFHQFFHCYCPVRFGREVDPNGDFIK 2 1 KYQPVLKNFPLQYIHEPWNAPESVQRAAKCVIGKDYPLPMVNHLEVSQLNIERMKQVYQRLTQYRGT 1 2 GLMSHSPQSDNGIIINVGNKNKNENSHAKQFRTDELRQNAVQRNQSNLN* 0 >CRY1_vilLie Villosa lienosa (mussel) JR505030 mollusc transcript assembly mollusc 0 MDEPPKKYVVHWFRKGLRLHDNPALCEAFKGASTFRCVYILDPWFAGVSQVGINKWR 2 1 FLLQCLEDLDSSLRKVNSRLFVIRGQPADVFPRLFK 0 0 EWQITSLSFEEDPEPFGKERDAAISAMAKEAGVEVIIRMSHTLFNLQK 2 1 IITENNGTPPLTFKRFQSILKTVGPPTKPVETVTLTTIGTARTPIENDHDDRYGVPSLEEL 1 2 GFDIDGLKPSVFQGGETEALLRLDRHLERK 0 0 AWVASFEKPKMTSQSLFPSQTTISPYLKFGCLSSRLFYWKLNDLYRR 0 0 VKKKSDPPLSLHGQLLWREFFYLAATNNPKFDRMVGNPICVQVPWDRNKEALAKWAEGKTGFPWIDAIMIQLREVGWIHHLARHSVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWLWLSCSSFFQQFLNCYCPVGFGKRADPAGDFIR 2 1 HYIPQLKGFHPKYIYEPWTAPYEVQVAAKCIIGKDYPQPMVDHNEVSRQNMERMKQVYQVLAMRASG 1 2 VITKTLTDDTISKHPHSKISYITSCSNHISGNKPSKAAILLGGDSMNKHGHTSDEDNTGNSTN* 0
Cryptochrome CRY4 sequences
This subfamily has its 8th and 9th exons split identically in position and phase to CRY1 in sea urchin and amphioxus whereas these exons are fused from lamprey to human. This suggests the split state is ancestral and that CRY4 arose as a gene duplication in stem vertebrates. Although CRY4 has persisted in most tetrapods to the present day, it has been lost in all mammals and also in crocodileans.
For the full set of 10 vertebrate CRY4 sequences, see: Curated reference sequences for cryptochromes and photolyases
>CRY4_galGal Gallus gallus (chicken) NP_001034685 CRY4 PumMed:19663499 synteny: ADIPOR1 UBE2T CRY4 LRIF1 DRAM2 CEPT1 0 MRHRTIHLFRKGLRLHDNPALLAALQSSEVVYPVYILDRAFMTSSMHIGALRWHFLLQSLEDLRSSLRQLGSCLLVIQGEYESVVRDHVQKWNITQVTLDAEMEPFYKEMEANIRGLGEELGFQVLSLMGHSLYNTQR 2 1 ILELNGGTPPLTYKRFLRILSLLGDPEVPVRNPTAEDFQ 2 1 RCSPPELGLAECYGVPLPTDLKIPPESISPWRGGESEGLQRLEQHLADQ 0 0 GWVASFTKPKTVPNSLLPSTTGLSPYFSTGCLSVRSFFYRLSNIYAQ 0 0 AKHHSLPPVSLQGQLLWREFFYTVASATPNFTKMAGNPICLQIRWYEDAERLHKWKT 0 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAAACFLTRGDLWISWEEGMK 0 0 VFEELLLDADYSINAGNWMWLSASAFFHHYTRIFCPVRFGRRTDPEGQYIR 2 1 KYLPILKNFPSKYIYEPWTASEEEQKQAGCII 12 GRDYPFPMVDHKEASDHNLQLMKQAREEQHRIAQLTR 1 2 DDADDPMEMKLKRDHSEESFTKTKAARMTEQT* 0
Cryptochrome 6-4 photolyases
For the full set of 16 metazoan CRY64 sequences, see: Curated reference sequences for cryptochromes and photolyases
>CRY64_anoCar Anolis carolinensis (lizard) XM_003225714 6-4 photolyase synteny: DCPS TIRAP CRY64 SRPR FOXRED1 0 MAHVSIHWFRKGLRLHDNPALLAAMKNSAEIYPIFILDPWFPKNMQVSINRWRFLIESLKDLDESLKKLNSR 2 1 LFVVRGRPAEVFPELFTKWKVTRLAFEVDTEPYARRDAEVVRLAAEHGVQVIQKVSHTLYDTER 2 1 IIVENSGKAPLTYTRLQTLVASLGPPKQPVPAPKLEDMK 1 2 DCCTPVKEDHDLEYGTPSYEELGQDPKTAGPHLYPGGETEALARLDLHMKRT 0 0 SWVCNFKKPETHPNSLTPSTTVLSPYVKFGCLSVRMFWWKLAEVYQG 0 0 RKHSDPPVSLHGQLLWREFFYTAGAGIPNFDRMENNPVCVQVDWDNNQEYLRAWRE 0 0 GQTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDADWSLNAANWQWLSASAFFHQFFRVYSPVTFGKKTDKNGEYIK 2 1 KYLPFLRKFSNDYIYEPWKAPRSLQERAGCIIGQDYPKPIVEHEKVYKRNLERMKAAYARRSPNLVIQAKDKVSQKKGV 1 2 NRKRPEAPTKAKVQAKKVKTKSS* 0
DASH cryptochrome CRY3 sequences
DASH is yet another member of the cryptochrome and photolyase family. It was identified only recently as active only on ssDNA repair, reportedly because of a barrier to flipping the damaged cyclobutane pyrimidine dimer dinucleotide out of dsDNA into its active repair site unless it is in a loop. In the species investigated to date, this enzyme uses folate (MTHF) and FAD, activated by blue light. It is a fairly remote outgroup to the cytochromes, with only CPD further diverged.
Its name is a peculiar acronym of Drosophila, Arabidopsis, Synechocystis and Homo -- yet the gene was never present in Drosophila or placentals. In Arabidopsis, the two copies are called CRY3 and PHR2. The many genome projects available today allow a quick determination of its rather unusual phylogenetic distribution.
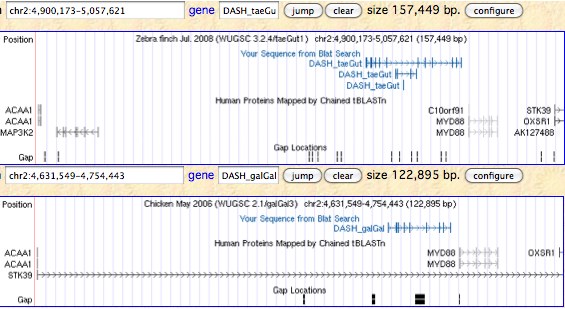
Although originally studied in plants and cyanobacteria, the DASH photolyase surprisingly extends into fish, frogs, salamanders, turtle, lizard, and birds -- duck, finch and budgerigar but not chicken or turkey -- but not any mammal. It's not clear however if the protein function has stayed the same in all these taxa or drifted in new roles like CRY1 and CRY2 or has acquired multiple functions in single species.
Using blastx on the syntenic region in gallinaceous birds (chicken and turkey) establishes that they have a fairly degenerate multi-exonic pseudogene at the expected location and orientation. It is not currently possible to date this more precisely nor determine whether pseudogenization occurred in a common ancestor or independently on account of separate domestications. Platypus does not have any pseudogene debris at the expected location but the assembly has a break here. Marsupials and placentals again appear never to have had this enzyme as it had been lost in the earliest mammals.
DASH is also missing from alligator and crocodile assemblies, the lobe-finned coelocanth assembly, and chondrichthyes genomes and transcripts. These probably represent additional, independent gene losses rather than just poor assemblies -- the overall sequence conservation of DASH is much less stringent than most cytochromes suggesting loosened constraints or even a measure of functional redundancy.
The phylogenetic loss pattern of DASH parallels the [Opsin_evolution:_trichromatic_ancestral_mammal|massive loss of opsins] that also occured early in mammalian evolution -- which GT Walls in 1942 [attributed] to mammals experiencing a sustained period of deep nocturnality where these systems did not need to function (no UV damage) and indeed could not function (insufficient blue light even with antenna) and so were lost, implying they were not sustained by a Piatigorskyian secondary functionality such as circadian rhythm, lunar calendaring, or magnetosensing.
The great oxygenation event gave rise to a stratospheric ozone protective layer at 2.4 gyr but reached an even higher lever during the early Cambrian (based indirectly on oxygen levels). This may have favored independent but simultaneous gene loss events in various clades.
However the first land plants and animals risked greatly increased levels of UV damage that may correlate with DASH retention. Here it must be clarified whether repairable damage can arise from direct oxidative damage in addition to ultraviolet light which could not plausibly be an issue for benthic marine organisms that still retain this enzyme.
The amniote DASH proteins can be modeled structurally using nearly 50% matches in Arabidopsis (cryptochrome 3: 2IJG) or equally suitable cyanobacterium Synechocystis (1NP7). The 14 exons share only one match with vertebrate CRY1 and CRY2 which is more likely coincidental than indicative of a shared ancestral protein subsequent to the era of eukaryotic intronation onset.
For a full set of metazoan DASH sequences, see: Curated reference sequences for cryptochromes and photolyases
>DASH_taeGut Taeniopygia guttata (finch) ABQF01044665 ABQF01044669 ABQF01044671 synteny: ACAA1 DASH MYD66 OXSR1 0 MSGTAGTAICLLRCDLRAHDNQ 0 0 QVLHWAQHNADFVIPLYCFDPRHYLGTHCYRLPKTGPHRLRFLLESVKDLRETLKKKGS 2 1 TLVVRKGKPEDVVCDLITQLGSVTAVVFHEE 0 0 ATQEELDVEKGLCQVCRQHGVKIQTFWGSTLYHRDDLPFRPIDR 2 1 LPDVYTHFPKGLESGAKVRPTLRMADQLKPLAPGLEEGSIPTMEDFGQK 1 2 DPVADPRTAFPCSGGETQALMRLQYYFWDT 0 0 NLVASYKETRNGLVGMDYSTKFAPW 2 1 LALGCISPRYIYEQIQKYERERTANESTYW 2 1 VLFELLWRDYFRFVALKYGRRIFSLR 1 2 GLQSKDIPWKKDLQLFSCWQ 0 0 EGKTGVPFVDANMRELSATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 0 VDYDVCSNYGNWLYSAGIGNDPRDNRKFNMIKQGLDYDGN 0 0 GDYVRLWVPELQGIKGADIHTPWALSSAALSQAGVTLGETYPQPVVTAPEWSRHIHRRP 0 0 GGSPHPRGRRGPAQRKDRGIDFYFSRKKDAC* 0
Cryptochrome CPD photolyases
This dna repair enzyme (cyclobutane pyrimidine dimers for CPD) was studied in marsupials during the pre-genomic era (1994), with two groups concluding even that that no ortholog existed in placentals. Today we are certain of that because the gene is not present in any complete placental mammal genome; no pseudogene debris exists in the partly conserved syntenic location in any species. This strongly suggests that the gene was lost once in stem placental rather than many times in later subclades (as happened with encephalopsin). The gene remains very strongly conserved in species such as opossum with no indication of impending loss.
The loss in placentals is somewhat peculiar given that CPD is a very ancient (pre-eukaryotal) member of the photolyase family, with highly conserved orthologs readily recoverable in other commonly studied marsupials, monotremes, birds, alligators, turtle, lizard, snakes, frog, fish, agnathan, amphioxus, sea urchin, many invertebrates, cnidarians, plants and so forth. However it also appears to be lost in tunicate -- indeed Ciona has lost all its photolyases leaving it a bit mysterious how it repairs these types of dna damage. Hemichordates have also lost all members of this gene family including CPD.
It is very unlikely that placentals displaced CPD with something better. More likely, CPD was lost during a dark phase of placental evolution when UV damage to dna was a non-issue and its photo-repair infeasible. Genes cannot be retained without selection (use it or lose it). Coming back out into the light millions of years later (having also lost DASH, CRY64 and [[Opsin_evolution:_update_blog|13 of 21 opsin genes]), they evidently made do with a less efficient excision repair that overlaps repair photolyase functionality.
The CPD gene product is very diverged from other photolyases though still retains the photolyase and FAD binding domain folds. The antenna moiety is usually reported as MTHF (folate). The best available structures are from rice (3UMV: 53% identity to marsupial) and an archaeal methanogen (Methanosarcina mazei 2XRZ) which likely uses 5-deazariboflavin Fo as antenna (which it can synthesize de novo). The latter enzyme repairs cyclobutane pyrimidine dimers in duplex DNA using blue or near-UV light.
Despite great divergence in primary sequence from other members of the gene family, fold conservation may explain in part the unexpected circadian compensatory capacity of marsupial CPD expressed in double CRY1/2 knockout mouse, seemingly driven by interaction of CPD with CLOCK of the CLOCK/BMAL1 system. CPD lacks any counterpart to the distal exons of placental CRY1.
CPD presents no special problems in classification as it clearly originated early in the history of prokaryotes and today serves as the outgroup to the overall metazoan photolyase gene family (though not as usefully as the less diverged DASH). It has never undergone gene duplication and divergence, at least none that stuck, and has been retained as single copy in the vast majority of species from choanflagellate to mammal. There are no noteworthy C-terminal expansions or supplemental exons within metazoan -- CPD is the exception among photolyases and cryptochromes for its lack of overt innovation. However as the knock-in experiment in mouse shows, CPD has unexpected properties.
The N-terminus has various extensions -- indeed the initial methionine is problematic -- but these are poorly conserved even within closely related taxa. Conservation sets in some 38 residues upstream of the first conserved methionine. While these 114 bp could represent conserved 5'UTR nucleotides rather than conserved amino acids, the two relevent crystallographic structures include this region (Methanosarcina 2XRY and rice 3UMV) as do many transcripts. Two in Xenopus (ES684787 BX851972) seem to rule out a cryptic short first exon splicing into the conserved region.
Some 32 curated CPD sequences spanning the whole of metazoan evolution are provided at the reference sequences. Many more could be extracted from GenBank should some research issue warrant more intensive surveying.
4Fe-4S photolyases and their relation to primases
An intriguing new subfamily of photolyases (1,2) recently surfaced that contains a 4Fe-4S cluster in the catalytic domain in addition to FAD. This meshes with the equally surprising finding of unmistakable fold similiarity between photolyases and the large subunit of archaeal-eukaryotic primase (eg the PRIM2 gene product in human), an ancient enzyme critical to the de novo synthesis of RNA primers neede DNA replication that also contains a 4Fe-4S cluster (as do various DNA repair enzymes such as helicases and endonucleases).
The photolyase antenna molecule, at least in Rhodobacter, is also new: the final intermediate in riboflavin biosynthesis, 6,7-dimethyl-8-ribityl-lumazine (which serves a similar role in biolumininescence). This illustrates again the flexibility of the antenna site -- the antenna molecule is unpredictable from primary sequence, indeed tertiary structure, even whether there is one. Given the list of possible antenna molecules might even now be incomplete, reconsitution experiments that don't find a suitable antenna molecule may simply have tested an insufficient range of molecules.
The new class of photolyase conflicts with the notion of a universal tryptophan triad chain in photolyases, agreeing with reports in other photolyases suggesting that the whole concept was wrong from the get-go. While there is no question that aromatic residues play a special role in this gene family and are often deeply conserved, so are many other amino acids.
Various inappropriate gene names -- PhrB already in use at GenBank for a different photolyase class, FeS-BCP inconsistent with observed phylogenetic distribution and hyphenated/capitalized in violation of nomenclature standards, CRYB for an non-cryptochrome enzyme -- won't be used here but instead PFES (photolyase iron sulfide). Reference sequences are provided below for two bacteria and two archaeal FeS photolyases, as well as yeast and human FeS primases; these suffice as GenBank probes.
Using the 4 conserved cysteines and aromatic residues as guide, representatives of the new photolyase class are readily located in bacteria (150 genera) but are more narrowly distributed in Archaea (8 genera of Euryarchaeota, no Crenarchaeota, Korarchaeota, Thaumarchaeota) suggesting horizontal gene transfer to them or gene loss. No eukaryotic photolyase yet has a 4Fe-4S domain (ignoring blast match XM_002537565 in castor bean that represents Agrobacterium contamination). Since eukaryotes arose from a relatively late symbiosis of an archaea and alphaproteobacter, one or even two homologs would initially have been present.
The 4Fe-4S cluster, being an ancient feature of primase and thus of the whole fold family, implies that FeS-photolyases are a relic, retaining a feature lost in subsequent gene duplications that gave rise first to CPD and then to the overall photolyase/cryptochrome gene family. The alternative scenario, that the 4Fe-4S cluster represents convergent evolution in photolyases (later independent acquisition) is wholly implausible given the complex requirements of geometry and the lack of utility of intermediate states.
Although in most of biochemistry, 4Fe-4S clusters serve a clear redox function, such a role has not been established for primases, helicases, other DNA repair enzymes, much less PFES photolyases. Conceivably the redox state of the 4Fe-4S cluster can sense the status of a DNA helix and facilitate rapid scanning for the odd damaged base among billions of normal ones. Since the FeS-photolyase role is not understood, the functional consequences of the loss of the cluster, presumably first in CPD, then CRY64 and its downstream duplications including cryptochromes, is difficult to evaluate.
Primase may be among the very oldest of enzymes since it is essential for DNA replication (ie, perhaps for exiting the hypothetical earlier RNA world). However UV damage is also a very old problem. Priming is not needed for RNA replication or transcription nor in DNA replication in mitochondria; bacteria generally use a non-homologous system.
One very intriguing idea starts with the observation that FAD mimics two free RNA bases with its flavin and adenine rings which are are stacked like bases (U-folded) in all studied photolyases. In primase -- which has no FAD -- two purine ribonucleotides at the FAD site, recognizing two bases of template DNA by conventional hydrogen bonding that perhaps resemble the flipped out cyclobutane pair needing repair by a photolyase.
Indeed, the template dinucleotide could even be stabilized temporarily as a cyclobutane pair, reversing the normal sense of the reaction, borrowing reductive units from the 4Fe-4S cluster (UV/blue light is not a known primase requirement). This would explain primase preference for a pyrimidine template. Photolyases then arose by replacing the two mononucleotides with FAD and adding a Rossmann fold for the antenna, with the utilization of light displacing the need for the 4Fe-4S cluster except in the PFES class of photolyases
Human primase also undergoes a profound conformational change from a three-helix binding site for DNA to a helix-sheet site as it counts primer size and passes it along to the catalytic subunit and other protein parteners. That's not so clear for not-so-large subunits archael primases which seem to lack an internal domain duplication. A large conformational change -- not just internal changes in FAD redox status -- is also needed in cryptochrome signalling, possibly this same one.
>PFES_agrTum Agrobacterium tumefaciens (bacteria) NP_355900 aka: PhrB MSQLVLILGDQLSPSIAALDGVDKKQDTIVLCEVMAEASYVGHHKKKIAFIFSAMRHFAEELRGEGYRVRYTRIDDADNAGSFTGEVKRAIDDLTPSRIC VTEPGEWRVRSEMDGFAGAFGIQVDIRSDRRFLSSHGEFRNWAAGRKSLTMEYFYREMRRKTGLLMNGEQPVGGRWNFDAENRQPARPDLLRPKHPVFAP DKITKEVIDTVERLFPDNFGKLENFGFAVTRTDAERALSAFIDDFLCNFGATQDAMLQDDPNLNHSLLSFYINCGLLDALDVCKAAERAYHEGGAPLNAV EGFIRQIIGWREYMRGIYWLAGPDYVDSNFFENDRSLPVFYWTGKTHMNCMAKVITETIENAYAHHIQRLMITGNFALLAGIDPKAVHRWYLEVYADAYE WVELPNVIGMSQFADGGFLGTKPYAASGNYINRMSDYCDCRYDPKERLGDNACPFNALYWDFLARNREKLKSNHRLAQPYATWARMSEDVRHDLRAKAAAFLRKLD* >PFES_rhoSph Rhodobacter sphaeroides (bacteria) CP000144 PDB|3ZXS PMID:22290493 6,7-dimethyl-8-ribityl-lumazine antenna aka CryPro 4Fe-4S photolyase MRGSHHHHHHGIRMLTRLILVLGDQLSDDLPALRAADPAADLVVMAEVMEEGTYVPHHPQKIALILAAMRKFARRLQERGFRVAYSRLDDPDTGPSIGAE LLRRAAETGAREAVATRPGDWRLIEALEAMPLPVRFLPDDRFLCPADEFARWTEGRKQLRMEWFYREMRRRTGLLMEGDEPAGGKWNFDTENRKPAAPDL LRPRPLRFEPDAEVRAVLDLVEARFPRHFGRLRPFHWATDRAEALRALDHFIRESLPRFGDEQDAMLADDPFLSHALLSSSMNLGLLGPMEVCRRAETEW REGRAPLNAVEGFIRQILGWREYVRGIWTLSGPDYIRSNGLGHSAALPPLYWGKPTRMACLSAAVAQTRDLAYAHHIQRLMVTGNFALLAGVDPAEVHEW YLSVYIDALEWVEAPNTIGMSQFADHGLLGSKPYVSSGAYIDRMSDYCRGCAYAVKDRTGPRACPFNLLYWHFLNRHRARFERNPRMVQMYRTWDRMEET HRARVLTEAEAFLGRLHAGEPV* >PFES_metMah Methanohalophilus mahii (Euryarchaeota) CP001994 4Fe-4S photolyase MRHYAEKLRNRGADITYIKTAELEKSLSRWIKKKGIDELNIAEPANITLKEYLGKLNIDCKIVFVDNKQFIWSIPEFNTWASSRKNLIMEDFYRTGRKNSEI LLEKDGKPSGGKWNLDRENRKLPPKNGFQKKPPQHIKFSPDKITKEIIAEVERSEYPTYGKGKDFNLAVTHEDAQKALDFFIEEKLSNFGPYQDIMLTGDNVLWHSILSPYLNLGL LHPLNVIKKAELAYYQKNLPLNSIEGFIRQILGWREYMHCIYKYTGDKYLKSNWFDHERELPDIYWYPERTSMNCMASVIEEVLNTGYAHHIQRLMILSNFALLAEVNPAKVKNWF HAAFIDAYDWVMQPNVIGMGQFADGGILATKPYISSANYINKMSDYCQNCTYNHNHRTGEDACPFNYLYWAFLHKNNEKLRDIGRMKLILKNLDRINKKELKQIMTHADDFLKSLK* >PFES_natPha Natronomonas pharaonis (Euryarchaeota) CR936257 4Fe-4S photolyase MTVLVLGDCLTEFGPLASDARSTDERVLCIEARAFARRKPYHPHKLTLVFSAMRHFRDRLREAGYTVDYRRVETFAEGLDAHFAAHPEDHIVTVRRTAHGAT DRLQRLVANRGGTVEFVADPRFHCSREEFDAWADGDPPYRHESFYRHMRRETGYLMDGDEPVGGEWNFDDENREFPGPEYVPPEPPQFEPDETTREVREWVDATFGEDGYDDAPYG GAWADPEPFSWPVTREGALQALEAFIEERLPTFGPYQDAMLGDEWAMNHALLSSSLNLGLLSPSEVIEAALAAFEEGSVSIASVEGFLRQVLGWREFVRHAYRRTPGMAAANQLGA AEPLPEFFWTGDTDMACVADAVDGVRTRGYAHHIERLMVLSNFATLYGVEPSRLNEWFHAAFVDAYHWVTTPNVVGMGTFGTDTLSTKPYVASANYIDRMSDHCSGCPYYKTKTTG DGACPFNALYWDFLGRNESQLRSNHRMGLVYSHYDDKSDGEREAIADRAETLRQRARNGTL* >PRIM2_homSap Homo sapiens (human) primase large subunit 4Fe-4S pdb|3L9Q,3Q36 MEFSGRKWRKLRLAGDQRNASYPHCLQFYLQPPSENISLIEFENLAIDRVKLLKSVENLGVSYVKGTEQYQSKLESELRKLKFSYRENLEDEYEPRRRDHISHFILRLAYCQSEELRRWF IQQEMDLLRFRFSILPKDKIQDFLKDSQLQFEAISDEEKTLREQEIVASSPSLSGLKLGFESIYKIPFADALDLFRGRKVYLEDGFAYVPLKDIVAIILNEFRAKLSKALALTARSLPAV QSDERLQPLLNHLSHSYTGQDYSTQGNVGKISLDQIDLLSTKSFPPCMRQLHKALRENHHLRHGGRMQYGLFLKGIGLTLEQALQFWKQEFIKGKMDPDKFDKGYSYNIRHSFGKEGKRT DYTPFSCLKIILSNPPSQGDYHGCPFRHSDPELLKQKLQSYKISPGGISQILDLVKGTHYQVACQKYFEMIHNVDDCGFSLNHPNQFFCESQRILNGGKDIKKEPIQPETPQPKPSVQKT* KDASSALASLNSSLEMDMEGLEDYFSEDS* >PRIM2_sacCer Saccharomyces cerevisiae (yeast) P20457 aka: PRI2_YEAST primase large subunit PDB|3LGB MFRQSKRRIASRKNFSSYDDIVKSELDVGNTNAANQIILSSSSSEEEKKLYARLYESKLSFYDLPPQGEITLEQFEIWAIDRLKILLEIESCLSRNKSIK EIETIIKPQFQKLLPFNTESLEDRKKDYYSHFILRLCFCRSKELREKFVRAETFLFKIRFNMLTSTDQTKFVQSLDLPLLQFISNEEKAELSHQLYQTVS ASLQFQLNLNEEHQRKQYFQQEKFIKLPFENVIELVGNRLVFLKDGYAYLPQFQQLNLLSNEFASKLNQELIKTYQYLPRLNEDDRLLPILNHLSSGYTI ADFNQQKANQFSENVDDEINAQSVWSEEISSNYPLCIKNLMEGLKKNHHLRYYGRQQLSLFLKGIGLSADEALKFWSEAFTRNGNMTMEKFNKEYRYSFR HNYGLEGNRINYKPWDCHTILSKPRPGRGDYHGCPFRDWSHERLSAELRSMKLTQAQIISVLDSCQKGEYTIACTKVFEMTHNSASADLEIGEQTHIAHP NLYFERSRQLQKKQQKLEKEKLFNNGNH*
247 curated refSeqs for metazoan cryptochromes and photolyases
The full length sequences have been moved to a separate page; only header lines are shown below. They are in a modern implementation of fasta format (broken into exons, with codon phase ie bp overhang shown), grouped by ortholgous clusters, and presented in phylogenetic order relative to mammals, with subtree symmetry state fixed by assembly quality. The fasta headers themselves are little databases showing gene name (following HUGO symbol use rules), genus, species, common name, accession number if not a simple genomic blat or whole genome alignment output, pubMed id if specifically studied in a journal article, followed by an unstructured comment field.
The availability of some orthology classes is quite limited due to recent origin in conjunction with restricted phylogenetic persistance but the sequencing effort itself is spread very unevenly across the phylogenetic tree of metazoans. For species with good assemblies, the entire repertoire of cryptochromes and photolyases is provided. For large gene like these with numerous exons, absence from the assembly usually means genuine absence from the genome. Even when only an exon or two gene fragment is available, the classifier can almost always assign the correct orthology class to it. However it is risky to assemble an entire gene from many unlinked contigs and that was not done here; certain important clades such as cartilaginous fish unfortunately lack coherent assemblies and applicable transcripts and provisional gene assemblies may be provided later.
A remarkable amount of the data has surfaced at GenBank only in the last six months, implying much weaker results had the project been done in 2011 but also that much better phylogenetic coverage will surface this year.
CRY1_homSap Homo sapiens (human) CRY1_panTro Pan troglodytes (chimpanzee) XM_509339 CRY1_ponAbe Pongo abelii (orangutan) XM_002823690 CRY1_nomLeu Nomascus leucogenys (gibbon) XM_003269977 CRY1_macMul Macaca mulatta (rhesus) NM_001194159 CRY1_calJac Callithrix jacchus (marmoset) XM_002752946 CRY1_saiBol Saimiri boliviensis (squirrel_monkey) nearly identical to marmoset CRY1_tarSyr Tarsius syrichta (tarsier) ABRT010205577 unsure if exon 2 is CRY1 or CRY2 CRY1_micMur Microcebus murinus (mouse_lemur) CRY1_otoGar Otolemur garnettii (bushbaby) AAQR03016495 CRY1_tupBel Tupaia belangeri (treeshrew) CRY1_musMus Mus musculus (mouse) NM_007771 all transcripts support longer exon 10 lost splice donor CRY1_ratNor Rattus norvegicus (rat) NM_198750 CRY1_criGri Cricetulus griseus (hamster) XM_003505292 CRY1_spaJud Spalax judaei (blind_mole_rat) AJ606298 CRY1_dipOrd Dipodomys ordii (kangaroo_rat) ABRO01202522 ABRO01202521 CRY1_hetGla Heterocephalus glaber (mole-rat) stop codon in place of conserved W8, CRY1_cavPor Cavia porcellus (guinea pig) last two exons diverged 69 bp separation CRY1_speTri Spermophilus tridecemlineatus (squirrel) Ictidomys CRY1_oryCun Oryctolagus cuniculus (rabbit) CRY1_oviAri Ovis aries (sheep) NM_001129735 19341811 19150926 CRY1_bosTau Bos taurus (cow) NM_001105415 XM_616063 CRY1_susScr Sus scrofa (pig) XM_003126079 CRY1_ailMel Ailuropoda melanoleuca (panda) XM_002927658 CRY1_loxAfr Loxodonta africana (elephant) XM_003405313 CRY1_triMan Trichechus manatus (manatee) AHIN01036366 AHIN01036362 very similar to elephant CRY1_monDom Monodelphis domestica (opossum) XM_003341966 CRY1_macEug Macropus eugenii (wallaby) assembly frameshift CRY1_sarHar Sarcophilus harrisii (tasmanian_devil) nearly identical to oppossum CRY1_triVul Trichosurus vulpecula (possum) EC362500 terminal transcript fragment CRY1_ornAna Ornithorhynchus anatinus (platypus) XM_001508563 = rubbish CRY1_tacAcu Tachyglossus aculeatus (echidna) SRR000649.130490 CRY1_galGal Gallus gallus (chicken) 11684328 17324421 15459395 CRY1_melGal Meleagris gallopavo (turkey) XM_003202363 CRY1_anaPla Anas platyrhynchos (duck) scaffold157 altSplExon11: GMTGVLVCRGSPGSHNYGKKDKT* CRY1_eriRub Erithacus rubecula (robin) AY585716 CRY1_sylBor Sylvia borin (warbler) AJ632120 15381765 CRY1_taeGut Taeniopygia guttata (finch) XM_002196518 CRY1_melUnd Melopsittacus undulatus (parakeet) AGAI01062111 CRY1_parWeb Paradoxornis webbianus (parrotbill) JR867166 TSA transcript CRY1_allMis Alligator mississippiensis (alligator) genome/blat CRY1_anoCar Anolis carolinensis (lizard) XM_003220923 CRY1_podSic Podarcis siculus (wall_lizard) DQ376040 16809482 CRY1_pytMol Python molurus (python) AEQU010547455 CRY1_chrPic Chrysemys picta (turtle) AHGY01469963 AHGY01469969 CRY1_xenTro Xenopus tropicalis (frog) NM_001087660 11533577 final four exons confirmed by many ESTs CRY1A_latCha Latimeria chalumnae (coelocanth) CRY1B_latCha Latimeria chalumnae (coelocanth) CRY1A_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01025403 CRY1B_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01016727 AHAT01016728 CRY1A_danRer Danio rerio (zebrafish) NM_001077297 CRY1A2_danRer Danio rerio (zebrafish) BC044558 AW184635 olfactory CRY1B_danRer Danio rerio (zebrafish) BC095305 EB921055 aka CRY2A CRY1C_danRer Danio rerio (zebrafish) BC164795 EE210836 aka CRY2B CRY1_petMar Petromyzon marinus (lamprey) Contig24766 CRY1_braFlo Branchiostoma floridae (amphioxus) XM_002609455 end uncertain CRY1A_strPur Strongylocentrotus purpuratus (urchin) XM_001194752 CRY2_homSap Homo sapiens (human) 11 exons CRY2_panTro Pan troglodytes (chimp) CRY2_gorGor Gorilla gorilla (gorilla) CRY2_ponAbe Pongo pygmaeus (orangutan) CRY2_rheMac Macaca mulatta (rhesus) CJ488220 testis CRY2_papHam Papio hamadryas (baboon) CRY2_calJac Callithrix jacchus (marmoset) CRY2_micMur Microcebus murinus (mouse_lemur) CRY2_musMus Mus musculus (mouse) CF898022 CRY2_ratNor Rattus norvegicus (rat) DN948283 prostate CRY2_criGri Cricetulus griseus (hamster) XR_135830 CRY2_spaJud Spalax judaei (blind_mole_rat) AJ606300 CRY2_dipOrd Dipodomys ordii (kangaroo_rat) CRY2_cavPor Cavia porcellus (guinea_pig) CRY2_hetGla Heterocephalus glaber (blind_mole_rat) EHA99865 CRY2_speTri Spermophilus tridecemlineatus (squirrel) CRY2_oryCun Oryctolagus cuniculus (rabbit) CRY2_turTru Tursiops truncatus (dolphin) CRY2_bosTau Bos taurus (cow) EG706191 lens CRY2_oviAri Ovis aries (sheep) NM_001129736 PubMed:19341811 CRY2_susScr Sus scrofa (pig) XM_003122835 CRY2_equCab Equus caballus (horse) CRY2_canFam Canis familiaris (dog) XM_540761 CRY2_ailMel Ailuropoda melanoleuca (panda) XM_002922310 iMet lost to assembly gap CRY2_myoLuc Myotis lucifugus (microbat) CRY2_pteVam Pteropus vampyrus (macrobat) CRY2_loxAfr Loxodonta africana (elephant) CRY2_triMan Trichechus manatus (manatee) AHIN01126950 AHIN01126951 CRY2_choHof Choloepus hoffmanni (sloth) CRY2_macEug Macropus eugenii (wallaby) FY652314 testis CRY2_monDom Monodelphis domestica (opossum) CRY2_ornAna Ornithorhynchus anatinus (platypus) CRY2_galGal Gallus gallus (chicken) AJ396745 bursa 19456395 15459395 CRY2_taeGut Taeniopygia guttata (finch) FE716439 brain CRY2_allMis Alligator mississippiensis (alligator) genome/blat CRY2_anoCar Anolis carolinensis (lizard) XM_003214641 CRY2_xenTro Xenopus tropicalis (frog) NM_001088670 AY049035 CX389867 11533577 discrepancies CRY2_ranCat Rana catesbeiana (bullfrog) GO458565 AY256684 extra SS removed CRY2_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01038797 CRY2_danRer Danio rerio (zebrafish) aka CRY3 NM_131786 CRY2_oreNil Oreochromis niloticus (tilapia) XM_003449249 split exon 7 also in gasAcu, oryLat, tetNig not danRef or lepOcu CRY2_tetNig Tetraodon nigroviridis (fugu) CAAE01010345 CRY2_takRub Takifugu rubripes (fugu) HE592015 CRY1B_strPur Strongylocentrotus purpuratus (sea_urchin) XM_001183029 echinoderm lacks final 2 exons CRY1B_lytVar Lytechinus variegatus (sea_urchin) AGCV01081039 echinoderm many small contigs CRY1B_parLiv Paracentrotus lividus (sea_urchin) AM599080 echinoderm many transcripts CRY1B_aplCal Aplysia californica (sea_hare) FF067636 AASC02010117 scaffold_151 mollusc CRY1B_octVul Octopus vulgaris (octopus) JR450373 transcript assembly mollusc CRY1B_craGig Crassostrea gigas (oyster) GQ415324 HS189569 mollusc CRY1B_rudPhi Ruditapes philippinarum (clam) JO113369 mollusc CRY1B_vilLie Villosa lienosa (mussel) JR510441 transcript assembly mollusc fragment CRY1B_lymSta Lymnaea stagnalis (snail) ES576734 mollusc CRY1B_plaDum Platynereis dumerilii (clam_worm) GU322429 annelid mRNA fragment CRY1B_dapPul Daphnia pulex (water_flea) ACJG01002273 FE370447 FE356368 crustacean CRY1B_diaNig Dianemobius nigrofasciatus (cricket) AB291231 orthoptera CRY1B_acyPis Acyrthosiphon pisum (aphid) NM_001171061 ABLF02032292 HP303737 hemiptera CRY1B_danPle Danaus plexippus (butterfly) AY860425 AGBW01012954 lepidoptera CRY1B_bomMor Bombyx mori (silkworm) NM_001195699 wrong BABH01015108 moth lepidoptera CRY1B_mamBra Mamestra brassicae (moth) AY947639 Glossata lepidoptera CRY1B_helArm Helicoverpa armigera (cotton_bollworm) JN997418 moth lepidoptera CRY1B_droMel Drosophila melanogaster (fruit_fly) AB019389 diptera PubMed:22080955 PDB:3TVS CRY1B_anoGam Anopheles gambiae (mosquito) DQ219482 diptera PubMed:16332522 CRY1B_neoBul Neobellieria bullata (fleshfly) FJ373353 diptera CRY1B_bacCuc Bactrocera cucurbitae (melon_fly) AB517608 diptera CRY1A_dapPul Daphnia pulex (water_flea) FE418063 FE356487 ACJG01001137 crustacean CRY1A_eupSup Euphausia superba (krill) FM200054 contig crustacean CRY1A_acyPis Acyrthosiphon pisum (aphid) NM_001171102 ABLF02035823 hemiptera cry2-2 PubMed:20482645 end uncertain CRY1A_ripPed Riptortus pedestris (bean_bug) AB379863 hemiptera PubMed:18547745 CRY1A_triCas Tribolium castaneum (flour_beetle) AAJJ01000096 coleopetera CRY1A_bomImp Bombus impatiens (bumble_bee) EF110521 AEQM02008194 hymenoptera PubMed:17244599 CRY1A_apiMel Apis mellifera (bee) NM_001083630 AADG06001305 hymenoptera CRY1A_attCep Atta cephalotes (ant) ADTU01021771 hymenoptera CRY1A_exoRob Exoneura robusta (bee) HP928681 hymenoptera fragment CRY1A_nylPub Nylanderia pubens (crazy_ant) JP792144 hymenoptera fragment CRY1A_Nasonia vitripennis (wasp) XM_001606355 AAZX01001169 hymenoptera N-term shortened CRY1A_antPer Antheraea pernyi (silkmoth) EF117812 lepidoptera PubMed:17244599 dropped long C-terminus CRY1A_anoGam Anopheles gambiae (mosquito) DQ219483 diptera dropped long C-terminus CRY1A_aedAeg Aedes aegypti (mosquito) XM_001655728 diptera dropped long C-terminus CRY1_vilLie Villosa lienosa (mussel) JR505030 mollusc transcript assembly mollusc CRY1_tetUrt Tetranychus urticae (spider-mite) CAEY01002034 chelicerate N-terminus uncertain CRY1_aplCal Aplysia californica (sea_hare) scaffold_2275 mollusc small fragment CRY4_galGal Gallus gallus (chicken) NP_001034685 CRY4 PumMed:19663499 synteny: ADIPOR1 UBE2T CRY4 LRIF1 DRAM2 CEPT1 CRY4_taeGut Taeniopygia guttata (finch) XM_002198497 CRY4_melGal Meleagris gallopavo (turkey) XM_003212851 CRY4_pasDom Passer domesticus (sparrow) AY494987 16687285 fragment CRY4_anoCar Anolis carolinensis (lizard) synteny: UBE2T CRY4 LRIF1 DRAM2 CRY4_xenTro Xenopus tropicalis (frog) NP_001123706 CRY4_latCha Latimeria chalumnae (coelocanth) AFYH01009222 CRY4_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01016726 CRY4_danRer Danio rerio (zebrafish) BC164413 CRY4_braFlo Branchiostoma floridae (amphioxus) XM_002609457 wrong CRY64_anoCar Anolis carolinensis (lizard) XM_003225714 6-4 photolyase synteny: DCPS TIRAP CRY64 SRPR FOXRED1 CRY64_chrPic Chrysemys picta (turtle) AHGY01135270 AHGY01135271 no synteny CRY64_allMis Alligator mississippiensis (alligator) blat CRY64_croPor Crocodylus porosus (crocodile) blat/genome CRY64_xenTro Xenopus tropicalis (frog) synteny: STS1 RPL27A CRY64 FOXRED1 SRPR PubMed:19715341 19345672 9016626 CRY64_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01024141 CRY64_danRer Danio rerio (zebrafish) BC044204 6-4 photolyase aka CRY5 CRY64_braFlo Branchiostoma floridae (amphioxus) XM_002595028 fused exons 2-3 BW780666 odd splice phases exon 5-6, no split 8-9, no last exon CRY64_strPur Strongylocentrotus purpuratus (urchin) XM_001189626 extra 1st exon unwarranted MCGAPRSYVEIRDSEEHSRRHVARLQFQFQSDLP 12 K CRY64_aplCal Aplysia californica (sea_hare) scaffold_427 CRY64_vilLie Villosa lienosa JR505030 transcript assembly mollusc CRY64_droMel Drosophila melanogaster (fruitfly) 6-4 photolyase 3CVW CG2488 uses 5-deazariboflavin CRY64_danPle Danaus plexippus (butterfly) EF117813 PubMed:17244599 two novel exons CRY64_acyPis Acyrthosiphon pisum (aphid) XM_001945977 single exon CRY64_anoGam Anopheles gambiae (mosquito) XM_314748 CRY64_bomMor Bombyx mori (silkworm) AK381942 frameshift DASH_taeGut Taeniopygia guttata (finch) ABQF01044665 ABQF01044669 ABQF01044671 synteny: ACAA1 DASH MYD66 OXSR1 DASH_anaPla Anas platyrhynchos (duck) scaffold1769 DASH_melUnd Melopsittacus undulatus (budgerigar) AGAI01061648 DASH_galGal Gallus gallus (chicken) syntentic pseudogene, numerous indels, frameshifts, internal stops DASH_melGal Meleagris gallopavo (turkey) ADDD01036185 syntenic pseudogene DASH_anoCar Anolis carolinensis (lizard) XM_003221869 14 exons DASH_chrPic Chrysemys picta (turtle) AHGY01416294 first exon off contig DASH_xenTro Xenopus tropicalis (frog) XM_002938001 PubMed:15147276 synteny: ACAA1 DASH MYD66 transcripts AL790297 CR419606 etc DASH_hymCut Hymenochirus curtipes (frog) fragment DASH_ambMex Ambystoma mexicanum (axolotl) CO785483 fragment DASH_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01010414 DASH_danRer Danio rerio (zebrafish) NM_205686 DASH_oreNil Oreochromis niloticus (tilapa) XM_003439198 DASH_patPec Patiria pectinifera (starfish) HP101597 DASH_strPur Strongylocentrotus purpuratus (urchin) DASH_aplCal Aplysia californica (sea_hare) scaffold_151:75,790-145,485 DASH_vilLie Villosa lienosa (mussel) JR504188 transcript assembly mollusc DASH_nemVec Nematostella vectensis (sea_anemone) XP_001623243 ABAV01026885 DASH_hydMag Hydra magnipapillata (cnidarian) XM_002166508 single exon ABRM01055505 DASH_monBre Monosiga brevicollis (choanoflagellate) XP_001745157 ABFJ01000402 DASH_phaTri Phaeodactylum tricornutum (diatom) XM_002178853 CPF2 DASH_thaPse Thalassiosira pseudonana (diatom) XM_002291289 CPD_monDom Monodelphis domestica (opossum) NP_001028149:wrong OPC1 PubMed:7937136 synteny: TNK1 MUC4 CPD KIAA0226 FYTTD1 CPD_sarHar Sarcophilus harrisii (tasmanian_devil) AEFK01107967 CPD_potTri Potorous tridactylus (rat kangaroo) D26020 PubMed:7813451 CPD_ornAna Ornithorhynchus anatinus (platypus) CPD_taeGut Taeniopygia guttata (finch) XM_002190577 CPD_melUnd Melopsittacus undulatus (budgerigar) AGAI01046895 CPD_galGal Gallus gallus (chicken) XM_422729 CPD_melGal Meleagris gallopavo (turkey) XM_003209143 CPD_allMis Alligator mississippiensis (alligator) genome/blat CPD_chrPic Chrysemys picta (turtle) AHGY01112360 incomplete CPD_anoCar Anolis carolinensis (lizard) XM_003226963 CPD_pytMol Python molurus (python) CPD_xenTro Xenopus tropicalis (frog) CPD_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01034265 CPD_danRer Danio rerio (zebrafish) NM_201064 CPD_petMar Petromyzon marinus (lamprey) rough but revised sequence CPD_braFlo Branchiostoma floridae (amphioxus) XP_002586934 FE570347 CPD_strPur Strongylocentrotus purpuratus (urchin) JT122393 JT102939 FJ812411 CPD_aplCal Aplysia californica (sea_hare) scaffold_446:238,174 CPD_vilLie Villosa lienosa (mussel) JR505029 transcript assembly mollusc CPD_droMel Drosophila melanogaster (fruitfly) thymidine dimer photolyase CG11205 uses 5-deazariboflavin CPD_nasVit Nasonia vitripennis (wasp) XM_001603235 trimmed N-terminal CPD_bomImp Bombus impatiens (bumble_bee) XM_003488984 CPD_apiMel Apis mellifera (bee) XM_003250426 CPD_anoGam Anopheles gambiae (mosquito) XM_313925 trimmed N-terminal CPD_aedAeg Aedes aegypti (mosquito) XM_001653905 trimmed N-terminal CPD_acyPis Acyrthosiphon pisum (aphid) XM_001949116 trimmed N-terminal CPD_nemVec Nematostella vectensis (anemone) ABAV01006764 XM_001636204 bad CPD_ampQue Amphimedon queenslandica (sponge) ACUQ01006132 XM_003388698 bad CPD_acrDig Acropora digitifera (coral) BACK01030119 one intron missing CPD_monBre Monosiga brevicollis (choanflagellate) ABFJ01000652 related intronation but numerous differences CPD_salSpp Salpingoeca species (choanflagellate) ACSY01000967 different intronation still CRY1A_acrMil Acropora millepora (coral) EF202589 CRY1B_acrMil Acropora millepora (coral) EF202590 CRY1A_nemVec Nematostella vectensis (anemone) XM_001623096 CRY1B_nemVec Nematostella vectensis (anemone) XM_001623096 CRY1C_nemVec Nematostella vectensis (anemone) XM_001630979 CRY1D_nemVec Nematostella vectensis (anemone) XM_001632799 CRY1E_nemVec Nematostella vectensis (anemone) XM_001632800 CRY4_nemVec Nematostella vectensis (anemone) XP_001636303 ABAV01006592 last exon uncertain CRY2_ampQue Amphimedon queenslandica (sponge) XM_003386521 CRY_ampQue Amphimedon queenslandica (sponge) XM_003386534 CRY_subDom Suberites domuncula (sponge) FN421335 CRY4_craMey Crateromorpha meyeri (sponge) PubMed:20121950 CRY_aphVas Aphrocallistes vastus (sponge) PubMed:14499587 CRY64A_triAdh Trichoplax adhaerens (placozoa) XM_002108524 CRY64B_triAdh Trichoplax adhaerens (placozoa) XM_002107723 CRY1A_araTha Arabidopsis thaliana (cress) CRY1 HY4 NM_116961 AFNC01018176 CRY1B_araTha Arabidopsis thaliana (cress) CRY2 PHH1 NM_100320 AFNB01000167 no antennal chromophore CRY1C_araTha Arabidopsis thaliana (cress) UVR3 NM_001035626 AFNC01013058 DASH1_araTha Arabidopsis thaliana (cress) PHR2 NM_130327 AFNA01010806 DASH2_araTha Arabidopsis thaliana (cress) CRY3 NM_122394 AFMZ01019177 CPD_araTha Arabidopsis thaliana (cress) PHR1 NM_179320 AFMZ01000529 GC-AG splice exon 6-7 CRY_phaTri Phaeodactylum tricornutum (diatom) XM_002180059 PMID:19424294 CRY_thaPse Thalassiosira pseudonana (diatom) XM_002291108 PFES_agrTum Agrobacterium tumefaciens (bacteria) NP_355900 aka: PhrB PFES_rhoSph Rhodobacter sphaeroides (bacteria) CP000144 PDB|3ZXS PMID:22290493 6,7-dimethyl-8-ribityl-lumazine antenna aka CryPro 4Fe-4S photolyase PFES_metMah Methanohalophilus mahii (Euryarchaeota) CP001994 4Fe-4S photolyase PFES_natPha Natronomonas pharaonis (Euryarchaeota) CR936257 4Fe-4S photolyase PRIM2_homSap Homo sapiens (human) primase large subunit 4Fe-4S pdb|3L9Q,3Q36 PRIM2_sacCer Saccharomyces cerevisiae (yeast) primase large subunit 4Fe-4S
For full sequences, see: Curated reference sequences for cryptochromes and photolyases