File list
From genomewiki
Jump to navigationJump to search
This special page shows all uploaded files.
| Date | Name | Thumbnail | Size | User | Description | Versions |
|---|---|---|---|---|---|---|
| 21:19, 12 May 2011 | 29mammalsConstrainedElements html.txt (file) | 1 KB | Brianraney | gwUploadFile upload | 2 | |
| 22:01, 16 September 2008 | 2008-08-archive (file) | 1 KB | Kuhn | 3 | ||
| 20:26, 12 May 2011 | 29mammalsConstrainedElements track.txt (file) | 2 KB | Brianraney | gwUploadFile upload | 2 | |
| 05:07, 20 March 2010 | ABRF2010 chr21 export grayQual.png (file) | 2 KB | AngieHinrichs | checkpoint image for demo/tutorial at ABRF 2010 workshop | 1 | |
| 13:35, 24 November 2010 | FixStepToBedGraph pl v2.txt (file) | 2 KB | Max | Updated version: filters out track lines + can handle variableStep files | 1 | |
| 04:39, 20 March 2010 | ABRF2010 redbin 1.png (file) | 2 KB | AngieHinrichs | checkpoint image for demo/tutorial at ABRF 2010 workshop | 1 | |
| 00:19, 14 February 2006 | G-Wiki.png (file) |  |
2 KB | Chesman | 1 | |
| 13:39, 24 November 2010 | FixStepToBedGraph pl.txt (file) | 2 KB | Max | added one comment | 4 | |
| 23:34, 18 March 2015 | TopAll sh.txt (file) | 2 KB | Hiram | script used in lastz tuning procedure to run lastz_D on different sets of DNA sequences from query and target | 1 | |
| 17:09, 19 March 2015 | SelectedFasta sh.txt (file) | 2 KB | Hiram | generalize the chrom.size creation and noMask option on fasta creation | 2 | |
| 23:18, 18 March 2015 | MafScoreSizeScan pl.txt (file) | 2 KB | Hiram | script used to scan a MAF file to extract scores, sizes, and coverage statistics of each alignment block. Used in lastz tuning procedures. | 1 | |
| 19:00, 10 August 2006 | TestCustomTrack.data.txt.gz (file) | 2 KB | Hiram | Testing file upload feature. This is a custom track test file demonstrating all track types currently (August 2006) supported by the browser custom tracks. | 1 | |
| 16:43, 24 June 2011 | ExtractRepeats.txt (file) | 2 KB | Hiram | Small perl script used to extract lineage specific repeats from RepeatMasker .out files | 1 | |
| 05:10, 20 March 2010 | ABRF2010 chr21 export baseQual.png (file) | 2 KB | AngieHinrichs | checkpoint image for demo/tutorial at ABRF 2010 workshop | 1 | |
| 18:00, 11 June 2007 | CleanHgCentral pl.txt (file) | 2 KB | Hiram | PERL cleaner script for userDb and sessionDb tables in hgcentral | 1 | |
| 06:48, 1 May 2013 | BoG2013variationAbstract.txt (file) | 2 KB | Rhead | 3 | ||
| 05:04, 13 March 2013 | BlatJob.csh.txt (file) | 2 KB | Hiram | script to use for the same species lift over procedure run.blat/ step 1 | 1 | |
| 21:28, 10 August 2006 | DoUpdateDb sh.txt (file) | 2 KB | Acs | Shell script for updating local mysql database with data from hgdownload | 1 | |
| 03:00, 17 October 2011 | SQNewEnhancersIR3-PeakAnnotation.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 18:32, 7 January 2010 | RetrUcscGenomes.txt (file) | 3 KB | Max | python scripts to download newest version of all 2bit genome files from UCSC with rsync | 2 | |
| 21:27, 10 August 2006 | DoDownloads sh.txt (file) | 3 KB | Acs | Shell script for downloading data for selected assemblies from hgdownload | 1 | |
| 23:30, 30 September 2011 | SQ840MotifAnnotation.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 01:37, 1 October 2011 | SQ850MotifPeakAnnotation.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 01:26, 1 October 2011 | SQ854MotifPeakAnnotation.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 03:31, 1 October 2011 | SQ948MotifPeakAnnotation.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 02:59, 17 October 2011 | SQNewEnhancersIR3PeakAnnotation.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 17:10, 19 March 2015 | AdjustSizes pl.txt (file) | 3 KB | Hiram | chrom.sizes files are now expected to exist as created by selectedFasta.sh | 2 | |
| 13:32, 23 December 2007 | Opsins class tree.png (file) | 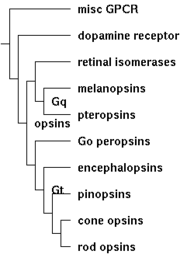 |
3 KB | Tomemerald | 1 | |
| 20:51, 30 September 2011 | HOMERGRDEX-Peaks.bed (file) | 3 KB | Daria Merkurjev | 1 | ||
| 22:47, 13 May 2008 | EmbedURL php.txt (file) | 3 KB | Hiram | UCSC modified EmbedURL extension code php file. Adds scrolling and frameborder options, and defaults width and height to 100% and 500 | 1 | |
| 17:10, 26 September 2018 | InitParasol.sh.txt (file) | 3 KB | Hiram | less restrictive subnet specification and separate hub/node logs in ./logs/ directory | 3 | |
| 04:47, 20 March 2010 | ABRF2010 redbinGt15.png (file) | 4 KB | AngieHinrichs | checkpoint image for demo/tutorial at ABRF 2010 workshop | 1 | |
| 21:54, 1 October 2018 | AwsAttachVolume.sh.txt (file) | 4 KB | Hiram | Script for AWS instance initialization to attach a data volume to a running instance. Used in the startParaHub.sh script upon starting the machine. | 1 | |
| 01:19, 1 September 2006 | Encodelogo.gif (file) |  |
4 KB | Kate | 1 | |
| 17:42, 11 July 2018 | OpenStackParaNodeSetup.sh.template.txt (file) | 4 KB | Hiram | correctly fixup /mnt filesystem for broken openstack installation, and mount /mnt/data from the hub | 2 | |
| 17:43, 12 August 2015 | RunLastzChain sh.txt (file) | 4 KB | Jonathan Casper | Removed B=0 from lastz parameters - it had been included by mistake. (https://groups.google.com/a/soe.ucsc.edu/d/msg/genome/BZjUyNgRKnM/ESMVNVvJDQAJ) | 2 | |
| 17:14, 19 March 2015 | Expand scores file py.txt (file) | 4 KB | Hiram | Script created by Bob Harris for lastz tuning operations | 1 | |
| 16:39, 18 September 2007 | Mm8 mm9.intronSize.png (file) |  |
5 KB | Hiram | Mouse mm8 and mm9 intron size histogram, to size 50 nucleotides | 2 |
| 16:46, 18 September 2007 | Hg17 hg18.intronSize.png (file) |  |
5 KB | Hiram | Human hg17 and hg18 intron size histogram to size 50 nucleotides | 2 |
| 19:24, 1 December 2009 | PhastCons46way.histogram.png (file) |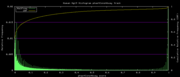  |
5 KB | Hiram | 46-way conservation track on human genome browser hg19. phastCons histogram data for all 46 vertebrates | 1 |
| 19:24, 1 December 2009 | PhastCons46wayPlacental.histogram.png (file) |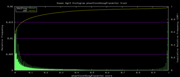  |
5 KB | Hiram | 46-way conservation track on human genome browser hg19. phastCons histogram data for all placental mammal subset | 1 |
| 05:43, 13 March 2013 | SameSpeciesChainNet.sh.txt (file) | 5 KB | Hiram | Same species lift over, step 2 of 2, chain/net the blat psl results together into the result file | 1 | |
| 12:19, 13 November 2009 | LoxDomainTree.jpg (file) | 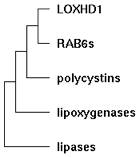 |
5 KB | Tomemerald | 1 | |
| 16:36, 18 September 2007 | Mm8 mm9.exonCount.png (file) |  |
5 KB | Hiram | Mouse mm8 and mm9 exon count by size histogram, to size 30 nucleotides | 2 |
| 16:43, 18 September 2007 | Hg17 hg18.exonCount.png (file) |  |
5 KB | Hiram | Human hg17 and hg18, exon count by size histogram, to size 30 nucleotides | 2 |
| 18:28, 30 October 2007 | PhastCons30way.png (file) |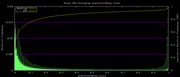  |
5 KB | Hiram | Mouse Mm9 30-way conservation phastCons histogram plot for the track data phastCons30way | 1 |
| 22:03, 3 December 2007 | Mm9Placental.png (file) |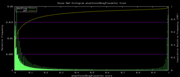  |
5 KB | Hiram | Mm9 alternative phastCons data histogram, Placentals, =mm9,rn4,cavPor2,oryCun1,hg18,panTro2,ponAbe2,rheMac2,calJac1,otoGar1,tupBel1,sorAra1,eriEur1,canFam2,felCat3,equCab1,bosTau3,dasNov1,loxAfr1,echTel1 | 3 |
| 22:04, 3 December 2007 | Mm9Euarchontoglires.png (file) |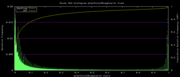  |
5 KB | Hiram | Mm9 alternative phastCons data histogram, Euarchontoglires, mm9,rn4,cavPor2,oryCun1,hg18,panTro2,ponAbe2,rheMac2,calJac1,otoGar1,tupBel1 | 3 |
| 07:41, 1 September 2018 | BlatBot pl.txt (file) | 5 KB | Galt | changed cse to soe | 4 | |
| 19:25, 1 December 2009 | PhastCons46wayPrimates.histogram.png (file) |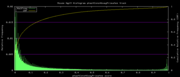  |
5 KB | Hiram | 46-way conservation track on human genome browser hg19. phastCons histogram data for the primates subset | 1 |